Abstract
We applied comparative genomic hybridization (CGH) to 46 breast carcinoma samples, collected from 1993 to 1995, in order to detect chromosome 1q gains and 16q losses and to define whether samples showing both these alterations had distinct biopathologic features and different clinical outcome. A total of 22 samples (48%) had simultaneous chromosome 1q gain and 16q loss, which was always associated with other genetic changes. In total, 23 samples had various chromosome imbalances (including chromosome 1q gain independent of chromosome 16q loss and vice versa) and one sample did not show detectable alterations. Samples having chromosome 1q gain/16q loss were compared to the other samples with regard to neoplasm size, lymph-node status, histologic and nuclear grade, estrogen and progesterone receptor presence, Ki-67, pRB, Cyclin D1, Cyclin A, p53, p21 and p27 expression as detected by immunohistochemistry. The samples showing chromosome 1q gain/16q loss had high steroid hormone receptor expression (P=0.02), low cell growth fraction (Ki-67, P=0.03) and high p27 expression (P<0.001). No statistical correlation with disease-free survival and overall survival or response to hormonal therapy was found. We conclude that simultaneous chromosome 1q gain/16q loss is a frequent event in invasive breast cancer, which occurs in a subset of both intermediate- and high-grade breast carcinomas. Although the final chromosome 1q and 16q imbalances might have originated from different chromosome alterations in low- and high-grade samples, the gene–dosage effect might be important in conferring peculiar biopathologic characteristics to this subset of samples. The cytogenetic and molecular mechanisms underlying these chromosome changes deserve further investigations.
Similar content being viewed by others
Main
Chromosome 1q gains and 16q losses have often been described to be simultaneously present in breast carcinoma samples by karyotype analysis1, 2 and comparative genomic hybridization (CGH) studies.3, 4 Gains of the long arm of chromosome 1 and losses of chromosome 16q are often the result of unbalanced translocations between these two chromosomes:1, 6 less frequently, they are the result of other rearrangements.2, 5 Chromosome 1q gains and 16q losses are considered as early changes since they have been detected as the sole chromosome abnormalities in near-diploid breast carcinoma1, 6 and in well-differentiated samples with a few alterations.3, 7 These genetic changes and the resulting chromosome imbalances have been thought to play a pathogenic role in breast carcinoma development. As simultaneous chromosome 1q gains and 16q losses have often been detected, we hypothesized that they might be related with particular biopathologic characteristics.
In the present study, we intended to define the biopathologic profile of samples having both chromosome 1q gain/16q loss. To this aim, we applied CGH since it is one of the most suitable techniques to reveal chromosome imbalances in solid neoplasm by a single experiment using a small quantity of DNA. We investigated 46 breast carcinoma samples, collected between 1993 and 1995, with 8–10 years follow-up. Samples showing simultaneous chromosome 1q gain/16q loss were compared to the other specimens with regard to biopathologic characteristics and clinical outcome.
Materials and methods
Samples
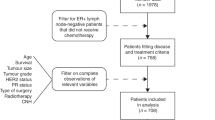
Tumor samples were obtained from 46 women (age range 28–83 years) undergoing surgery for primary breast carcinoma at the S.Orsola University Hospital of Bologna, Italy, from 1993 to 1995. Only consecutive cases that had a frozen tissue block were selected for this study.
In total, 12 patients underwent conservative breast surgery followed by radiation therapy and 34 underwent radical mastectomy. None of the patients received preoperative chemo- or hormonal chemotherapy. Adjuvant systemic chemotherapy (cisplatinum, methotrexate, 5-fluorouracil) was given to 14 patients. In all, 32 patients, presenting steroid receptor positive status, received adjuvant hormonal therapy (tamoxifen, 20 mg daily).
A part of each neoplastic tissue fragment was immediately frozen and stored at −80°C for subsequent DNA extraction. Another part of each sample was fixed in buffered 4% formaldehyde and embedded in paraffin. Sections of 4 μm thickness were cut and stained by hematoxylin and eosin for routine histopathologic classification according to WHO.8 In total, 42 samples were invasive ductal carcinoma, three were invasive lobular carcinoma, and one was a mucinous carcinoma. Histologic grading was carried out according to Elston and Ellis9 and nuclear grading (NG) as follows: mild (NG1), moderate (NG2) and high nuclear atypia (NG3). In all, 44 patients were followed for 19–99 months (median = 84 months).
DNA Extraction and Labeling
DNA was extracted from 46 frozen breast carcinoma tissue fragments and human normal female liver tissue using the Qiamp Tissue Kit (Qiagen, GmbH, Hilden, Germany) according to the instructions of the supplier. In order to reduce the quantity of false-negative results, samples showing a percentage of neoplastic cells fewer than 50% were not used for DNA extraction and CGH analysis.10, 11 The percentage of neoplastic vs stromal and inflammatory cells was defined by one pathologist (DS), scoring the control hematoxylin–eosin–stained sections of each sample.
Measures of 1 μg of normal DNA and 1 μg of tumor DNA were labeled by nick translation (Gibco BRL, Gaithersburg, MD, USA). Normal DNA was always labeled by digoxigenin-11-dUTP (Roche, Basel, Switzerland) and tumor DNA by biotin-16-dUTP (Roche, Basel, Switzerland). DNA fragment size was checked by 1% agarose gel electrophoresis and ethidium bromide staining and DNAse I concentration was adjusted in order to obtain an optimal average fragment size (500–1500 bp). Normal and tumor-labeled DNA were precipitated with 40 μl Cot-1 DNA (1 mg/ml) (Roche, Basel, Switzerland) and dissolved in 6 μl hybridization mixture (50% formamide, 10% dextran sulfate, 2 × SSC (300 mM sodium chloride plus 30 mM sodium citrate)).
CGH
Metaphase preparations from normal peripheral blood lymphocytes were obtained according to standard conventional cytogenetic techniques and used for CGH. Metaphase chromosomes were denatured in 70% formamide 2 × SSC, pH 7, at 80°C for 3 min, washed in 70, 90 and 100% ethanol and air dried. The DNA probes were denatured at 80°C for 10 min and applied onto the slides, which were coverslipped, sealed by rubber cement, put in a moist chamber and let to hybridize at 37°C for 72 h. Washings were performed at room temperature (2 × SSC, 5 min for two times) and at 45°C (0.1 × SSC, 5 min for three times). Blocking step was performed using 0.5% blocking reagent (Roche, Basel, Switzerland) dissolved into 1 × TN (100mM Tris-Hcl pH 7.5, 150mM Nacl) for 10 min at 37°C and followed by immunocytochemical detection. Digoxigenin was detected by a sheep anti-digoxigenin-tetramethylrhodamine-isothiocyanate (TRITC)-conjugated antibody (1:50 in TNB) (Roche, Basel, Switzerland) and biotin by a avidin-fluorescein-isothiocyanate (FITC)-conjugate (1:200) (Vector, Burlingame, CA, USA). The antibodies were incubated at 37°C for 1 h. The slides were washed in TNT at room temperature (5 min for three times), dehydrated in ethanol, air dried and mounted with antifade containing DAPI (0.35 μg/μl).
CGH images were recorded by a Leica DMRA fluorescence microscope and analyzed by the Cytovision CGH software package 3.5 (Applied Imaging, Sunderland, UK). In total, 9–12 metaphases were analyzed for each sample and the average green-to-red intensity ratio profiles were calculated and plotted with 95% confidence interval (CI).11 Any relatively small, distinct locus showing a green-to-red fluorescence ratio greater than 1.5 was regarded as high-level amplification. Chromosomes 19, 1p32–ter and 16p– were excluded from the analysis since it has been demonstrated that the green-to-red fluorescence ratio in these regions cannot be reliably measured.10
Immunohistochemistry
In order to define steroid hormone receptor status and cell proliferation in our samples, the following monoclonal antibodies were used: anti-estrogen receptor (ER) clone 1D5 (BioGenex Laboratories, San Ramon, CA, USA), diluted 1:120; anti-progesterone receptor (PR) clone 1A6 (BioGenex Laboratories, San Ramon, CA, USA), diluted 1:65; anti-Ki-67 antigen clone Ki-88 (BioGenex Laboratories, San Ramon, CA, USA), diluted 1:100; anti-retinoblastoma protein (pRB) clone G3-245 (BioGenex Laboratories, San Ramon, CA, USA), diluted 1:160; anti-cyclin A clone 6E6 (NovoCastra Laboratories, Newcastle upon Tyne, UK) diluted 1:300; anti-cyclin D1 clone P2D11F11 (NovoCastra Laboratories, Newcastle upon Tyne, UK) diluted 1:200; anti-p53 protein (p53) clone BP53.12.1 (BioGenex Laboratories, San Ramon, CA, USA) diluted 1:3000; anti-p21 protein (p21) clone EA10 (Oncogene, Boston, MA, USA) diluted 1:450; anti-p27 protein (p27) clone DCS-72.F6 (Neomarkers, Fremont, CA, USA) diluted 1:190.
Serial sections from neutral buffered formaldehyde-fixed, paraffin-embedded tissue blocks were collected on 3-aminopropyltriethoxysilane (Sigma Co., St Louis, MO, USA) coated slides, dried overnight at 37°C, and processed for immunohistochemistry according to a streptavidin–biotin–peroxidase preformed complex (SABC) technique. Microwave pretreatment was performed with a citrate buffer solution (pH 6.0) for anti-ER, anti-PR, anti-Ki-67, anti-cyclin A, anti-p53, and anti-p27 antibodies, or with EDTA buffer solution (pH 8.0) for anti-pRB and anti-cyclin D1 antibodies.
Image Cytometry
Nuclear immunostaining of ER, PR, cyclin D1, cyclin A, p21, p27, p53, pRB, and Ki-67, was quantified by image cytometry with Cytometrica software (C & V, Bologna, Italy), as previously detailed.12 A labeling index was obtained for all the biologic parameters evaluated by image cytometry and expressed as the percentage of the labeled nuclear area over the total neoplastic nuclear area (percentage Labeling Index area, %LIa). All the above-mentioned parameters were categorized using the following cutoff values: ER and PR %LIa <10% = negative, ≥10% positive; cyclin D1 %LIa <5% = negative, ≥5% = positive; CycA %LIa <20% = negative, ≥20% = positive; p21 %LIa <10% = negative; ≥10% = positive; p27 %LIa <50% = negative, ≥50% = positive; p53 %LIa < 4% = negative (1), ≥4% and <30% = intermediate (2), ≥30% = positive (3); pRB loss of expression was recorded as previously detailed13 and reported using two categories: 0 = loss of function, 1 = functional pRB protein; Ki-67 %LIa <20% = low proliferation (1) and ≥20% = high proliferation (2).
Statistics
To assess differences between the two groups of samples with respect to biopathologic parameters, χ2-test has been applied. Correlation between 1q gains and biopathologic parameters were investigated by χ2-test as well as correlation between chromosome 16q losses and all the investigated variables. Disease-free survival (DFS) and overall survival (OS) were evaluated by Kaplan–Meier curves and logrank test. P-values below 0.05 were regarded as significant.
Results
Genomic Alterations as Detected by CGH
Genomic alterations were found in 45 out of 46 samples. DNA copy number gains were more frequent than losses, as summarized in Figure 1. The most frequent minimal common regions (mcr) gained were: 1q23–q32 (78%), 8q21–qter (74%), 5q14–23 (35%), 5p13–pter (33%), 3q11–q21 (33%), 4q11–31 (30%), 12p11–p12 (30%), 12q14–q21 (30%), Xq11–qter (30%). Mcr lost were found at 17p12–pter (74%), 16q12–qter (59%), 8p21–pter (33%), 11q21–qter (28%) and 17q11–qter (28%).
A total number of 47 high-level amplification events was detected in 15 samples, involving chromosome 1q23–32, 2p24–25, 3q25–q26, 4q26–32, 5p14, 5q15–q22, 6q11–22, 7p21–22, 8p11.2, 8q23–q24, 9p22–p24 11p14, 11q13, 12p11.2-12, 12q15–21, 13q22, 14q12–q21, 14q24–q31, 15q26, 16p12, 17q22–23, 18p11.2–11.3, 20p11.2, 20q12–q13.1, and Xq21–26.
Relationship Between Chromosome 1q Gain, 16q Loss, Biopathologic Variables and Clinical Outcome
Chromosome 1q gain/16q loss was found in 22 out of 46 samples (48%) and was never detected as the sole genetic change. Out of 22 samples having chromosome 1q gain/16q loss, 12 also showed chromosome 12q gain (P=0.002): no other chromosomal imbalance was found to be associated with.
As simultaneous chromosome 1q gain/16q loss has been reported to occur in intermediately and well-differentiated samples, we evaluated the incidence of these alterations in our set of samples stratified by grade. Only one sample was classified as G1 and did not show either chromosome 1q gain or chromosome 16q loss. G2 and G3 samples showed chromosome 1q gain/16q loss in 56 and 40% samples, respectively.
Chromosome 1q gains and 16q losses affected large chromosome regions (Table 1). All but one sample lost a large part or the whole long arm of chromosome 16. Chromosome 1q gains only involved a part of the long arm in six cases: the common region of overlap included chromosome 1q31 band.
We divided samples showing both chromosome 1q gain/16q loss from the other samples (24 samples: 15 showing 1q gain but not 16q loss, five samples showing 16q loss but not 1q gain and four samples having neither 1q gain nor 16q loss) and compared the two groups with respect to several biopathologic parameters. As shown in Table 2, we found a positive correlation with steroid hormone receptor presence (P=0.02), low cell growth fraction as evaluated by Ki-67 expression (P=0.03), and high p27 expression (P<0.001).
In univariate analysis, chromosome 1q gain was correlated with p27 (P= 0.002) and cyclin D1 (P=0.01) expression. Chromosome 16q loss was associated with p27 (P=0.01) expression. On their own, none of these events were correlated with both ER and PgR presence and Ki-67 immunostaining.
We did not find any correlation between the presence of chromosome 1q gain/16q loss and survival. Figures 2a and b show the Kaplan–Meier curve for DFS and OS, respectively. Out of 21 patients, eight relapsed in the group of samples having chromosome 1q gain/16q loss and six out of 23 in the group having other chromosome imbalances (logrank Mantel–Cox test, P=0.771). Six patients died of disease in the group showing chromosome 1q gain/16q loss and seven in the group with other chromosome imbalances (logrank Mantel–Cox test, P=0.916). Finally, we investigated DFS and OS in the 32 patients who had steroid hormone receptors presence and received tamoxifen. Chromosome 1q gain/16q loss was not related to difference in DFS and OS, as shown in Figures 3a and b (logrank Mantel–Cox test, P=0.603 and P=0.718, respectively).
Prognostic significance of chromosome 1q gain/16q loss in 46 breast carcinoma samples. Shown here are the Kaplan–Meier-estimated (a) DFS and (b) OS, respectively, for Group A (open triangle, including samples showing chromosome 1q gain/16q loss) and B (solid triangle, including samples with other abnormalities) patients. Symbols indicate censored observations.
Prognostic significance of chromosome 1q gain/16q loss in 32 breast carcinoma patients who underwent hormonal therapy. Shown here are the Kaplan–Meier–estimated (a) DFS and (b) OS rates, respectively, for Group A (open triangle, including samples showing chromosome 1q gain/16q loss) and B (solid triangle, including samples with other abnormalities) patients. Symbols indicate censored observations.
Discussion
In the present study, we applied CGH technique to 46 breast carcinoma samples in order to detect simultaneous chromosome 1q gain/16q loss and we compared the biopathologic characteristics of samples showing these alterations with respect to other samples. As chromosome 1q gain/16q loss have often been found together, we speculated that this association might be synergistic. This hypothesis was supported by the association we found between chromosome 1q gain/16q loss and steroid hormone receptor presence, low cell growth fraction, and high p27 expression. In univariate analysis, only the correlation with p27 expression was present. p27 is a cell cycle-related protein, the expression of which is indicative of low proliferation and good prognosis. p27 is a direct inhibitor of cyclin-dependent kinase 2 (cdk2), one of the cdk's responsible for the activation of E2F1 transcription factors that promote DNA replication.14, 15 Altogether, these data suggest a functional link between genes located on chromosome 1q and 16q (dysregulated or lost), steroid hormone sensitivity and low proliferation rate.
In contrast, chromosome 1q gain/16q loss does not seem to be important either for survival or for response to tamoxifen treatment, although breast carcinoma patients having ERs and PRs represent a clinically well-defined group of patients who are expected to have a better prognosis. The present results are too preliminary to enable us to find an explanation: they only support the hypothesis that the simultaneous presence of both alterations is important to acquire these biopathologic characteristics. A larger number of patients, selected on the basis of the therapeutic treatment received, ought to be investigated to draw definitive conclusions about the impact of chromosome 1q gain/16q loss presence on prognosis.
We speculated that chromosome 1q gain/16q loss might be an event pathogenically important, as it could induce a gene–dosage effect and a consequent gene dysregulation.6 In fact, DNA copy number alterations are associated with a corresponding change in mRNA levels and, finally, in cell function.16 A large part of chromosome 1q and 16q DNA was gained and lost as a consequence of these different genetic changes. According to cytogenetic studies, these chromosome imbalances are thought to occur as a consequence of breakage and recombination of alfa and beta satellite DNA located at chromosome regions 1q12, 16cen and 16q11.2.17 These repetitive DNA sequences are prone to anomalous recombination and fusion because of a high homology degree. As they are not transcribed, it is rather difficult to speculate about specific genes affected by these chromosome changes. Indeed, we suggest that the gene–dosage effect might have a pathogenic role in conferring these biopathologic characteristics.
In several studies, chromosome 1q and 16q imbalances have been investigated separately: chromosome 1q gains have been found at any stage,18, 19, 20 whereas chromosome 16q losses have been mostly detected in low and intermediately differentiated carcinoma3, 4 or in invasive lobular carcinoma.21 Altogether, these data supported the hypothesis that low/intermediate- and high-grade breast carcinoma develop according to distinct pathways: chromosome 16q loss was proposed as a marker of a distinct genetic pathway in breast carcinoma development.3, 7 In the present study, all but one samples were classified as G2 or G3 and only three samples were invasive lobular carcinoma. The percentage of samples showing chromosome 1q gain/16q loss was lower in G3 with respect to G2 samples. The present data support the hypothesis that in G2 and G3 samples, chromosome 1q gain/16q loss originated, at least in part, by distinct chromosome changes. They might have developed according to different pathways: unbalanced translocation between chromosome 1 and 16 in G1/G2 samples vs other chromosome changes in G3 samples. Molecular studies, able to define better the chromosome changes occurring in different samples, might be important to further clarify this point. We conclude that chromosome 1q gain/16q loss is a frequent event in both low- and high-grade breast carcinoma samples. Although the final chromosome imbalance might have originated from different chromosome alterations in low- and high-grade samples, the resulting gene–dosage effect might be important in conferring peculiar biopathologic characteristic to this subset of samples.
References
Dutrillaux B, Gerbault-Seureau M, Zafrani B . Characterization of chromosomal anomalies in human breast cancer. A comparison of 30 paradiploid cases with few chromosome changes. Cancer Genet Cytogenet 1990;49:203–217.
Pandis N, Jin Y, Gorunova L, et al. Chromosome analysis of 97 primary breast carcinomas: identification of eight karyotypic subgroups. Genes Chromosomes Cancer 1995;12:173–185.
Roylance R, Gorman P, Harris W, et al. Comparative genomic hybridization of breast tumors stratified by histological grade reveals new insights into the biological progression of breast cancer. Cancer Res 1999;59:1433–1436.
Bürger H, Otterbach F, Simon R, et al. Different genetic pathways in the evolution of invasive breast cancer are associated with distinct morphological subtypes. J Pathol 1999;189:521–526.
Texeira MR, Pandis N, Heim S . Cytogenetic clues to breast carcinogenesis. Genes Chromosomes Cancer 2002;33:1–16.
Tsarouha H, Pandis N, Bardi G, et al. Karyotypic evolution in breast carcinomas with i(1)(q10) and der(1;16)(q10;p10) as the primary chromosome abnormality. Cancer Genet Cytogenet 1999;13:156–161.
Bürger H, Mommers E, Littmann R, et al. Ductal invasive G2 and G3 carcinomas of the breast are the end stage of at least two different lines of genetic evolution. J Pathol 2001;194:165–170.
WHO. The World Health Organization histologic typing of breast tumours. (2nd edition). Am J Clin Pathol 1982;78:806–816.
Elston CW, Ellis IO . Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 1991;19:403–410.
Kallioniemi OP, Kallioniemi A, Piper J, et al. Optimizing comparative genomic hybridization for analysis of DNA sequence copy number changes in solid tumors. Genes Chromosomes Cancer 1994;10:231–243.
Weiss MM, Hermsen MA, Meijer GA, et al. Comparative genomic hybridisation. Mol Pathol 1999;52:243–251.
Faccioli S, Chieco P, Gramantieri L, et al. Cytometric measurement of cell proliferation in echo-guided biopsies from focal lesions of the liver. Mod Pathol 1996;9:120–125.
Ceccarelli C, Santini D, Chieco P, et al. Retinoblastoma (RB1) gene product expression in breast carcinoma. Correlation with Ki-67 growth fraction and biopathological profile. J Clin Pathol 1998;51:818–824.
Lloyd RV, Erickson LA, Jin L, et al. p27kipi: a multifunctional cyclin-dependent kinase inhibitor with prognostic significance in human cancers. Am J Pathol 1999;154:313–323.
Sherr CJ, Roberts JM . CDK inhibitors: positive and negative regulators of G1-phase progression. Genes Dev 1999;13:1501–1512.
Pollack JR, Sorlie T, Perou CM, et al. Microarray analysis reveals a major direct role of DNA copy number alteration in the transcriptional program of human breast tumors. Proc Natl Acad Sci USA 2002;99:12963–12968.
Kokalj-Vokac N, Alemeida A, Gerbault-Seureau M, et al. Two-color FISH characterization of i(1q) and der(1;16) in human breast cancer cells. Genes Chromosomes Cancer 1993;7:8–14.
Trent J, Yang JM, Emerson J, et al. Clonal chromosome abnormalities in human breast carcinomas. II. Thirty-four cases with metastatic disease. Genes Chromosomes Cancer 1993;4:194–203.
Farabegoli F, Ceccarelli C, Santini D, et al. Chromosome 1 aneusomy with 1p36 under-representation is related to histologic grade, DNA aneuploidy, high c-erb B-2 and loss of bcl-2 expression in ductal breast carcinoma. Int J Cancer 1996;69:381–385.
Tirkkonen M, Tanner M, Karhu R, et al. Molecular cytogenetics of primary breast cancer by CGH. Genes Chromosomes Cancer 1998;3:177–184.
Nishizaki T, Chew K, Chu L, et al. Genetic alterations in lobular breast cancer by comparative genomic hybridization. Int J Cancer 1997;74:513–517.
Acknowledgements
Fulvia Farabegoli was a recipient of a short-term EMBO fellowship (ASTF 9620). This work was supported by MURST, University of Bologna (funds for selected topics) and Pallotti Legacy for Cancer Research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Farabegoli, F., Hermsen, M., Ceccarelli, C. et al. Simultaneous chromosome 1q gain and 16q loss is associated with steroid receptor presence and low proliferation in breast carcinoma. Mod Pathol 17, 449–455 (2004). https://doi.org/10.1038/modpathol.3800059
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800059
Keywords
This article is cited by
-
Role of columnar cell lesions in breast carcinogenesis: analysis of chromosome 16 copy number changes by multiplex ligation-dependent probe amplification
Modern Pathology (2018)
-
Integrative analysis of copy number and gene expression in breast cancer using formalin-fixed paraffin-embedded core biopsy tissue: a feasibility study
BMC Genomics (2017)
-
Chromosome 1q25.3 copy number alterations in primary breast cancers detected by multiplex ligation-dependent probe amplification and allelic imbalance assays and its comparison with fluorescent in situ hybridization assays
Cellular Oncology (2013)
-
BRCA1-mutated and basal-like breast cancers have similar aCGH profiles and a high incidence of protein truncating TP53 mutations
BMC Cancer (2010)
-
Microarray-based comparative genomic hybridisation of breast cancer patients receiving neoadjuvant chemotherapy
British Journal of Cancer (2007)