Abstract
Solid tumors are composed not only of tumor cells but also of stromal nonneoplastic cells. In whole tumor samples, stromal cells retaining their alleles may therefore obscure detection of loss of heterozygosity (LOH) in tumor cells. An increasing number of studies have used laser-assisted tissue microdissection to improve LOH detection, but the real gain in sensitivity has been poorly quantified. We studied a group of 16 inflammatory breast carcinomas that were submitted to both standard DNA extraction from frozen whole tumor samples and laser microdissection performed on paraffin-embedded tumor samples. Using PCR with fluorescence-labeled primers, we comparatively analyzed ten polymorphic markers with both sources of DNA. With the LOH detection threshold set at −25%, we showed that 25 LOHs could not be diagnosed with whole tumor samples out of 73 LOHs positively diagnosed in microdissected samples (34%). With the LOH detection threshold set at −50%, the respective figures were 39 LOHs not diagnosed out of 55 LOHs (71%). Measuring the intensity of the allelic decrease, we showed that the mean decrease of the lost allele is −34% with whole tumor samples and −67% with microdissected samples. The increase in sensitivity of LOH detection with microdissection is associated with the density of stromal cells. This strong improvement in LOH detection in this aggressive type of breast cancer indicates that many other molecular studies performed on heterogeneous solid tumors may benefit from a first step of laser microdissection.
Similar content being viewed by others
Introduction
Loss of heterozygosity (LOH) is common in human solid tumors and allows the expressivity of recessive loss-of-function mutations in tumor suppressor genes (Lasko et al, 1991). The detection of recurrent LOH in a genomic region is considered to be critical evidence for the localization of tumor suppressor genes. Because detection of LOH is based on the comparison of tumor cells and corresponding normal cells for identification of tumor cell–specific gene deletions, it is important to obtain a collection of pure tumor cells to provide the homogeneous material required for a reliable analysis. However, most solid cancers contain, not only tumor cells, but also stromal cells (eg, fibroblasts, lymphocytes, and endothelial cells) or residual nontumor cells (eg, adipocytes and normal residual ducts). These cells usually have a normal genome and therefore may obscure losses of genetic material in tumor cells when they are too numerous in whole tumor samples. To overcome this problem, several tumor cell–enrichment protocols have been developed, such as flow cytometry based on an abnormal DNA index (Abeln et al, 1994) or tissue microdissection (TM) (Bertheau et al, 1998; Sirivatanauksorn et al, 1999). Initially performed manually (Zhuang et al, 1995) or with a micromanipulator (Going and Lamb, 1996; Küppers et al, 1994), TM has evolved to laser-assisted microdissection systems that can efficiently sample various amounts of cells (Böhm et al, 1997; Emmert-Buck et al, 1996; Fend and Raffeld, 2000; Schütze et al, 1997). These systems are much easier to handle than hydraulic micromanipulators and much more precise than manual microdissection.
Several reports have stated that the use of TM is of great benefit to the detection of LOH (Fujii et al, 1996; Giercksky et al, 1997; Shen et al, 2000; Speiser et al, 1996). Only one study (Giercksky et al, 1997) has quantitatively estimated the gain in sensitivity obtained with TM. That report compared genetic changes in frozen biopsies and in manually microdissected archival material from the same colorectal liver metastases. The authors found a 54% increase in the sensitivity of detection of genetic alterations with microdissection. Thus, we decided to make a similar comparison in a group of 16 inflammatory breast carcinomas that were submitted to both standard DNA extraction from frozen whole tumor tissue and microdissection of tumor tissue embedded in paraffin. In contrast to Giercksky et al (1997), we used a laser-based tissue microdissection system, and we detected LOHs with fluorescence-labeled primers that allow precise quantitation of allelic decrease. Our results showed that at least one-third of LOHs present in breast tumors remain undiagnosed if the tissue has not been microdissected. Furthermore, we estimated that the sensitivity of LOH detection with TM is nearly double that without TM, especially in tumors with highly cellular stroma.
Results
Sixteen tumors were studied at 10 loci. Two methods have been compared: method 1 used whole tumor tissue, whereas method 2 used microdissected tissue (Fig. 1). For both methods, two LOH detection thresholds were tested.
Figures 2 and 3 display the results. No PCR product was obtained in five tests with microdissected samples. With both methods 1 and 2, 37 tests (24%) were uninformative and 40 tests (26%) showed retention of heterozygosity. We found no cases of microsatellite instability.
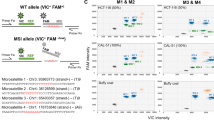
Results in all 16 patients tested for 10 loci with loss of heterozygosity (LOH) threshold at −25%. Method 1: with whole tumor tissue; method 2: with microdissected tissue. Stromal cell density is scored from 1 to 3 (see “Materials and Methods”). ▪, LOH (peak decrease more than 25%); ?, uncertain LOH (decrease between 10% and 25%); □, retained (peak decrease less than 10%); ▒, uninformative cases; npp, no PCR product.
Results in all 16 patients tested for 10 loci with LOH threshold at −50%. Method 1: with whole tumor tissue; method 2: with microdissected tissue. Stromal cell density is scored from 1 to 3 (see “Materials and Methods”). ▪, LOH (peak decrease more than 50%); ?, uncertain LOH (decrease between 10% and 50%); □, retained (peak decrease less than 10%); ▒, uninformative cases; npp, no PCR product.
Detection Threshold Set at −25%
With the detection threshold set at −25% (Fig. 2), 16 tests showed no LOH with method 1, but did show LOH with method 2 (one example is given in Fig. 4). Nine LOHs were uncertain with method 1, yet were certain using method 2. Five uncertain LOHs remained uncertain with both methods. Forty-eight tests showed LOH with both methods. Therefore 25 LOHs were not diagnosed with method 1 out of 73 LOHs diagnosed with method 2 (34% nonrecognized LOH). In the 73 tests that showed LOH with method 2, the mean decrease of the lost allele was −34% with method 1 and was −67% with method 2, indicating that method 2 is nearly twice as sensitive as method 1 to the detection of a decrease in one allele.
Patient 10, locus D8S1820. The decrease of the right allele in the tumor compared with normal is −3% with DNA extracted from whole tissue (method 1) and −76% with microdissected tissue (method 2). N, method 1, normal sample with method 1 (see “Materials and Methods”); T, method 1, tumor sample with method 1; N, method 2, normal sample with method 2 (see “Materials and Methods”); T, method 2, tumor sample with method 2.
Detection Threshold Set at −50%
With the detection threshold set at −50% (Fig. 3), six tests showed no LOH with method 1, but did show LOH with method 2. Thirty-three LOHs were uncertain with method 1, yet were certain with method 2. Ten uncertain LOHs remained uncertain with both methods. Sixteen tests showed LOH with both methods. Therefore, with the threshold at −50%, 39 LOHs were not diagnosed with method 1 out of 55 LOHs diagnosed with method 2 (71% nonrecognized LOH).
LOH Detection and Stromal Cellularity
The increase in sensitivity of LOH detection with method 2 is associated with the density of stromal cells. For the nine tumors with a poorly cellular stroma, the mean decrease of the allele was −40% with method 1 and −66% with method 2 (for 41 LOHs). For the five tumors with a highly cellular stroma, the figures were −23% and −67%, respectively.
Interestingly, five cases showed a greater decrease in alleles with method 1 than with method 2. For example, with Patient 11, the allele for p53CA decreased by 59% with method 1 and by 48% with method 2.
Discussion
Inflammatory breast cancer (IBC) is a very aggressive subtype of breast carcinoma that so far has been poorly characterized biologically. Inflammatory breast cancers are clinically defined by the presence of signs suggesting inflammation, such as breast redness, edema, and pain. Histologically however, IBC are not significantly different from noninflammatory breast cancers, except that they often contain dermal lymphatic emboli. It is crucial to find prognostic or predictive criteria for these tumors, which are treated by induction chemotherapy followed by surgery. We randomly selected 16 cases in a large population of IBC currently under investigation for a genome-wide search for specific LOHs.
Ten microsatellite markers were selected using two criteria. First, the frequency of allelic losses at each locus had to be previously described in sporadic breast cancers, and second, the PCR product length had to be less than 250 bp. PCR products longer than 250 bp are too difficult to obtain with microdissected, paraffin-embedded, formalin-fixed tissue. Despite these precautions, five tests with microdissected samples did not give any PCR product (Figs. 2 and 3) and were excluded from the analysis. Frozen tissues are not always available in routine practice, and therefore, we preferred using paraffin sections.
A stronger difference between the two methods might have been noted if the whole tumor tissue method (method 1) had been performed with frozen tissue not histologically controlled. This would have allowed us to study cases with only a few tumor cells or cases with much necrosis.
Tissue sectioning results in nuclear truncation, thus affecting any calculated DNA yield. For the microdissection method (method 2), we therefore took special care to sample enough cells (nearly 500 cell profiles for each PCR) to avoid artificial allelic losses (“allelic dropout ”).
The detection threshold for LOH assessment varies greatly among studies (Amari et al, 1999; Fujii et al, 1996; Kerangueven et al, 1997; Marsh and Varley, 1998). We decided to use two different threshold values and were able to show that the sensitivity gain with microdissection was higher with the −50% threshold. This is because most LOHs diagnosed with microdissection show peak decreases beyond −50%. However, only 55 LOHs could be diagnosed with the −50% threshold, compared with 73 LOHs with the −25% threshold.
It is likely that TM increases the sensitivity of LOH detection, not only in inflammatory breast cancer, but also in most other types of solid tumors. However, the gain in sensitivity is probably higher in breast cancer than in other cancers because breast cancer is more histologically heterogeneous, consisting of infiltrating tumor cells, noninfiltrating tumor cells (intraductal carcinoma, lobular carcinoma in situ), stromal cells, adipocytes, and residual epithelial cells.
It is interesting to ask why microdissection, especially with a laser system, does not systematically give −100% allelic decrease in cases of LOH. One possible explanation is that, even with a very careful use of a laser system, it is likely that a few stromal cells, perhaps very closely associated with tumor cells, will be sampled along with tumor cells. A second reason is intratumoral heterogeneity. It is now well known that cancer is a juxtaposition of subclones (Fey and Tobler, 1996) and that LOH distribution among tumor cells is heterogeneous (Chen et al, 1992; Hugel and Wernert, 1999; Yatabe et al, 2000). Our study supports the notion that some LOHs are not distributed homogeneously in tumor cells. This is because, in several tests, method 2 (with microdissection) showed smaller allelic decrease than method 1 (without microdissection). Also, cells microdissected in a single area of the tumor may contain proportionally fewer deletions than the whole tumor cell population. In our study, the small size of the biopsy samples (usually less than 1 cm) did not allow us to microdissect several tumor areas.
Our results clearly show that the gain in sensitivity is much greater in tumors with dense stroma because of the low sensitivity of method 1 in these tumors. However, the mean allelic decrease obtained with TM is not dependent on the cellularity of the stroma (−66% for tumors with poorly cellular stroma versus −67% for tumors with highly cellular stroma). Our results demonstrate that TM is precise enough to allow enrichment in tumor cells whatever the number of inflammatory cells.
Gains in sensitivity with TM are important for losses of genetic material, but TM also can improve the detection of mutations and amplifications (Lehmann et al, 2000; Pappalardo et al, 1998). TM also allows better assessment of gene expression (Specht et al, 2001) and better protein analysis (Emmert-Buck et al, 2000) in heterogeneous tumor tissues. Laser-assisted microdissection is likely to have a profound impact on molecular pathology and may soon be a prerequisite for many molecular studies that benefit from the pure cell populations.
Materials and Methods
Patients and Tissues
From 1993 to 1998, 72 patients were referred to Hospital Saint-Louis for inflammatory breast cancer. All these patients underwent frozen section examination at the time of initial diagnostic biopsy. Ten subsequent frozen sections (10 μm thick) were pooled in DNA extraction buffer, and the remaining tissue was fixed in AFA (Carlo Erba, Rodano, Italy), a mix of 2% formalin 40% (v/v), 5% acetic acid (v/v), 75% ethanol (v/v), and 18% water (v/v) and then embedded in paraffin for histopathological diagnosis. For our study, we randomly selected 16 patients in that population. All 16 tumors were infiltrating ductal carcinomas; 5 were Grade 2 and 11 were Grade 3 (Elston, 1987).
Stromal Cellular Density
On the tissue block from which frozen sections had been made for DNA extraction, the density of stromal cells (mostly small lymphocytes) was evaluated semiquantitatively on one H&E section according to the following criteria: 1 = less than 10% stromal cells, 2 = 10% to 50% stromal cells, and 3 = over 50% stromal cells. The stroma was scored “1” in 9 cases, “2” in 2 cases, and “ 3” in 5 cases (Figs. 2 and 3).
DNA Extraction
Frozen tumor sections were immersed in a buffer containing 8 M urea, 0.3 M NaCl, 10 mm EDTA, 2% SDS, and 10 mm Tris-HCl (pH 7.5) and then submitted to phenol chloroform DNA extraction. Control normal DNA was prepared from peripheral blood as previously described (Muniz et al, 1994). Tumor DNA and normal DNA were stored in 1 mm EDTA and 10 mm Tris-HCl (pH 7.5) at 4° C until further use.
Tissue Microdissection
Six-micrometer-thick paraffin sections were spread on nonpretreated glass slides and stained with H&E (Fig. 1). Tissue microdissection was performed with the laser microbeam microdissection system (PALM, Bernried, Germany) (Schütze et al, 1997). Briefly, a 337 nm UV-laser is used to “catapult” small tissue fragments directly into the cap of a sample tube without any mechanical contact. We used no membrane on the slide. For each tumor, at least 5000 infiltrating tumor cells microdissected in a single area of the section and 5000 nontumor cells (lymphocytes, adipocytes, and normal breast epithelial cells) were catapulted in separate vials containing 30 μl of lysis buffer (50 mm Tris-HCl [pH 7.5], 1 mm EDTA, 0.5% Tween20, 0.2 mg/ml proteinase K). Cells were then incubated overnight at 37° C and proteinase K was inactivated by heating at 95° C for ten minutes. No further DNA extraction was performed.
PCR
PCRs were performed in 25 μl final volume with either 5 ng of extracted DNA (method 1) or 3 μl of lysed microdissected cells (method 2) corresponding to nearly 500 cell profiles. Ten polymorphic microsatellite markers were used in this study. Primers for PCR amplification of the following markers were designed based on the nucleotide sequences obtained from Internet databases (http://www.gdb.org, http://www3.ncbi.nlm.nih.gov): D3S1573, D7S490, D8S261, D8S1820, D11S860, D11S1356, D13S171, D16S496, and D17S855. Another marker, p53CA, is a dinucleotide repeat at the p53 locus (Jones and Nakamura, 1992).
The PCR mix contained 1U Taq Gold (Applied Biosystems, Foster City, California), 2.5 to 4 mm MgCl2, 0.2 mm dNTP, 0.2 μm Cy5-labeled primers, and 0.2 μm nonlabeled primers. Thirty-five cycles were performed.
LOH Analysis
We used an automated DNA analysis system, ALFexpress II (Amersham Pharmacia Biotech, Uppsala, Sweden), that separates fluorescently labeled DNA fragments by electrophoresis. The detection range is 50 attomol to 45 femtomol DNA. PCR products were run on a 0.3-mm-thick UV-polymerized polyacrylamide gel (Reprogel, Amersham Pharmacia Biotech). For automated allele quantification, we used the software AlleleLocator 1.03 (Amersham Pharmacia Biotech). The intensity of fluorescence for each peak, and therefore for each allele, is directly computed by the system. We compared the intensity of fluorescence peaks between blood DNA and whole tumor DNA (method 1) and between microdissected nontumor cells and microdissected tumor cells (method 2). We used two different thresholds for the detection of LOH. With the −25% threshold, LOH was considered certain when one allele was decreased by at least 25% as compared to the normal profile. Decreases ranging from −10% to −25% were classified “uncertain LOH.” Decreases of less than 10% indicated “retention of heterozygosity.” With the −50% threshold, LOH was considered certain when one allele was decreased by at least 50% compared with the normal profile, and decreases ranging from −10% to −50% were classified “uncertain LOH.”
All PCRs with “certain LOH” or “uncertain LOH” were done twice. PCRs with “retention of heterozygosity” or with homozygosity were done only once.
References
Abeln EC, Corver WE, Kuipers-Dijkshoorn NJ, Fleuren GJ, and Cornelisse CJ (1994). Molecular genetic analysis of flow-sorted ovarian tumour cells: Improved detection of loss of heterozygosity. Br J Cancer 70: 255–262.
Amari M, Suzuki A, Moriya T, Yoshinaga K, Amano G, Sasano H, Ohuchi N, Satomi S, and Horii A (1999). LOH analyses of premalignant and malignant lesions of human breast: Frequent LOH in 8p, 16q, and 17q in atypical ductal hyperplasia. Oncol Rep 6: 1277–1280.
Bertheau P, Meignin V, and Janin A (1998). Microdissection sur préparation histologique et cytologique: Une approche de l'hétérogénéité tissulaire. Ann Pathol 18: 110–119.
Böhm M, Wieland I, Schutze K, and Rubben H (1997). Microbeam moment: Non-contact laser microdissection of membrane-mounted native tissue. Am J Pathol 151: 63–67.
Chen LC, Kurisu W, Ljung BM, Goldman ES, Moore DD, and Smith HS (1992). Heterogeneity for allelic loss in human breast cancer. J Natl Cancer Inst 84: 506–510.
Elston CW (1987). Grading of invasive carcinoma of the breast. In: Page DL and Anderson TJ, editors. Diagnostic histopathology of the breast. New York: Churchill Livingstone, 303–307.
Emmert-Buck MR, Bonner RF, Smith PD, Chuaqui RF, Zhuang ZP, Goldstein SR, Weiss RA, and Liotta LA (1996). Laser capture microdissection. Science 274: 998–1001.
Emmert-Buck MR, Gillespie JW, Paweletz CP, Ornstein DK, Basrur V, Appella E, Wang QH, Huang J, Hu N, Taylor P, and Petricoin EF 3rd (2000). An approach to proteomic analysis of human tumors. Mol Carcinog 27: 158–165.
Fend F and Raffeld M (2000). Laser capture microdissection in pathology. J Clin Pathol 53: 666–672.
Fey MF and Tobler A (1996). Tumour heterogeneity and clonality: An old theme revisited. Ann Oncol 7: 121–128.
Fujii H, Zhou WB, and Gabrielson E (1996). Detection of frequent allelic loss of 6q23-q25.2 in microdissected human breast cancer tissues. Genes Chromosomes Cancer 16: 35–39.
Giercksky HE, Thorstensen L, Qvist H, Nesland JM, and Lothe RA (1997). Comparison of genetic changes in frozen biopsies and microdissected archival material from the same colorectal liver metastases. Diagn Mol Pathol 6: 318–325.
Going JJ and Lamb RF (1996). Practical histological microdissection for PCR analysis. J Pathol 179: 121–124.
Hugel A and Wernert N (1999). Loss of heterozygosity (LOH), malignancy grade and clonality in microdissected prostate cancer. Br J Cancer 79: 551–557.
Jones MH and Nakamura Y (1992). Detection of loss of heterozygosity at the human TP53 locus using a dinucleotide repeat polymorphism. Genes Chromosomes Cancer 5: 89–90.
Kerangueven F, Noguchi T, Coulier F, Allione F, Wargniez V, Simonylafontaine J, Longy M, Jacquemier J, Sobol H, Eisinger F, and Birnbaum D (1997). Genome-wide search for loss of heterozygosity shows extensive genetic diversity of human breast carcinomas. Cancer Res 57: 5469–5474.
Küppers R, Rajewsky K, Zhao M, Simons G, Laumann R, Fischer R, and Hansmann ML (1994). Hodgkin disease: Hodgkin and Reed-Sternberg cells picked from histological sections show clonal immunoglobulin gene rearrangements and appear to be derived from B cells at various stages of development. Proc Natl Acad Sci USA 91: 10962–10966.
Lasko D, Cavenee WK, and Nordenskjöld M (1991). Loss of constitutionnal heterozygosity in human cancer. Annu Rev Genet 25: 281–314.
Lehmann U, Glockner S, Kleeberger W, von Wasielewski HF, and Kreipe H (2000). Detection of gene amplification in archival breast cancer specimens by laser-assisted microdissection and quantitative real-time polymerase chain reaction. Am J Pathol 156: 1855–1864.
Marsh KL and Varley JM (1998). Loss of heterozygosity at chromosome 9p in ductal carcinoma in situ and invasive carcinoma of the breast. Br J Cancer 77: 1439–1447.
Muniz ES, Plassa F, Amselem S, Goossens M, and Vernant JP (1994). Molecular analysis of polymorphic loci to study chimerism after allogeneic bone marrow transplantation. Heteroduplex analysis in denaturing gradient gel electrophoresis: A new approach to detecting residual host cells. Transplantation 57: 451–456.
Pappalardo PA, Bonner R, Krizman DB, Emmert-Buck MR, and Liotta LA (1998). Microdissection, microchip arrays, and molecular analysis of tumor cells (primary and metastases). Semin Radiat Oncol 8: 217–223.
Schütze K, Becker I, Becker KF, Thalhammer S, Stark R, Heckl WM, Böhm M, and Posl H (1997). Cut out or poke in—the key to the world of single genes—laser micromanipulation as a valuable tool on the look-out for the origin of disease. Genet Anal 14: 1–8.
Shen CY, Yu JC, Lo YL, Kuo CH, Yue CT, Jou YS, Huang CS, Lung JC, and Wu CW (2000). Genome-wide search for loss of heterozygosity using laser capture microdissected tissue of breast carcinoma: An implication for mutator phenotype and breast cancer pathogenesis. Cancer Res 60: 3884–3892.
Sirivatanauksorn Y, Drury R, Crnogorac-Jurcevic T, Sirivatanauksorn V, and Lemoine NR (1999). Laser-assisted microdissection: Applications in molecular pathology. J Pathol 189: 150–154.
Specht K, Richter T, Muller U, Walch A, Werner M, and Hofler H (2001). Quantitative gene expression analysis in microdissected archival formalin-fixed and paraffin-embedded tumor tissue. Am J Pathol 158: 419–429.
Speiser P, Gharehbaghischnell E, Schneeberger C, Eder S, and Zeillinger R (1996). Microdissection as a means to verify allelic imbalance in tumour biology samples. Anticancer Res 16: 461–464.
Yatabe Y, Konishi H, Mitsudomi T, Nakamura S, and Takahashi T (2000). Topographical distributions of allelic loss in individual non-small-cell lung cancers. Am J Pathol 157: 985–993.
Zhuang ZP, Bertheau P, Emmertbuck MR, Liotta LA, Gnarra J, Linehan WM, and Lubensky IA (1995). A microdissection technique for archival DNA analysis of specific cell populations in lesions less than 1 mm in size. Am J Pathol 146: 620–625.
Acknowledgements
We thank Mohamed Ismaïl and Nicole Croci, who prepared tissue sections and performed PCR, and Marcel Brusramer, who read the manuscript.
This work was supported by Grant PHRC No. AOM 97120 from the Délégation à la Recherche Clinique and by Grant No. 9726 from the Association pour la Recherche sur le Cancer.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bertheau, P., Plassa, L., Lerebours, F. et al. Allelic Loss Detection in Inflammatory Breast Cancer: Improvement with Laser Microdissection. Lab Invest 81, 1397–1402 (2001). https://doi.org/10.1038/labinvest.3780353
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3780353
This article is cited by
-
Molecular differences between ductal carcinoma in situ and adjacent invasive breast carcinoma: a multiplex ligation-dependent probe amplification study
Cellular Oncology (2011)
-
Protein pathway biomarker analysis of human cancer reveals requirement for upfront cellular-enrichment processing
Laboratory Investigation (2010)
-
Changes in allelic imbalances in locally advanced breast cancers after chemotherapy
British Journal of Cancer (2007)