Abstract
The vast majority of familial ovarian cancers harbor a germline mutation in either the breast cancer gene BRCA1 or BRCA2 tumor suppressor genes. However, mutations of these genes in sporadic ovarian cancer are rare. This suggests that in contrast to hereditary disease, BRCA1 and BRCA2 are not commonly involved in sporadic ovarian cancer and may indicate that there are two distinct pathways for the development of ovarian cancer. To characterize further differences between hereditary and sporadic cancers, the comparative genomic hybridization technique was employed to analyze changes in copy number of genetic material in a panel of 36 microdissected hereditary ovarian cancers. Gains at 8q23-qter (18 of 36, 5 cases with high-level amplifications), 3q26.3-qter (18 of 36, 2 cases with high-level amplifications), 11q22 (11 of 36) and 2q31–32 (8 of 36) were most frequent. Losses most frequently occurred (in decreasing order of frequency) on 8p21-pter (23 of 36), 16q22-pter (19 of 36), 22q13 (19 of 36), 9q31–33 (16 of 36), 12q24 (16 of 36), 15q11–15 (16 of 36), 17p12–13 (14 of 36), Xp21–22 (14 of 36), 20q13 (13 of 36), 15q24–25 (12 of 36), and 18q21 (12 of 36). Comparison with the literature revealed that the majority of these genetic alterations are also common in sporadic ovarian cancer. Deletions of 15q11–15, 15q24–25, 8p21-ter, 22q13, 12q24 and gains at 11q22, 13q22, and 17q23–25, however, appear to be specific to hereditary ovarian cancer. Aberrations at 15q11–15 and 15q24–25 have not yet been described in familial ovarian cancer. In these regions, important tumor suppressor genes, including the hRAD51 gene, are located. These and other yet unknown suppressor genes may be involved in a specific carcinogenic pathway for familial ovarian cancer and may explain the distinct clinical presentation and behavior of familial ovarian cancer.
Similar content being viewed by others
Introduction
Epithelial ovarian cancer ranks as the fourth most frequent cancer among women in the Western world. In The Netherlands, the age-standardized incidence rate is 14.9 per 100,000 woman-years and 1,029 deaths occurred in 1993 (Koper et al, 1996). Ten percent of the incidence can be attributed to an inherited autosomal dominant susceptibility factor (Boyd, 1998). It is currently estimated that more than 90% of these hereditary cases are due to germline mutations in one of two tumor suppressor genes, breast cancer gene (BRCA)1 and BRCA2. To date very few somatic mutations of BRCA1 or BRCA2 have been identified in sporadic ovarian cancer (Foster et al, 1996; Merajver et al, 1995). This implies that inactivation of these two genes is not required for the development of the more common sporadic form of the disease and suggests a difference in the molecular genetic pathway of carcinogenesis between hereditary and sporadic ovarian cancer. Further differences between the familial and sporadic forms of the disease have been described. Hereditary ovarian cancer is thought to occur at an earlier age. Histologically, serous tumors predominate in hereditary ovarian cancer cases, more so than in sporadic ovarian cancer (Zweemer et al, 1998). To learn more about the critical molecular genetic events in hereditary ovarian cancer, we studied a panel of 36 hereditary ovarian cancer cases by comparative genomic hybridization (CGH) and compared the findings with data from the literature on ovarian cancer cases unselected for genetic predisposition or family history.
Results
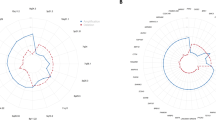
A summary of the copy number changes for all 36 hereditary tumors is displayed in Figure 1. The mean number of losses was 9.7 and the mean number of gains was 4. Frequent gains occurred at 8q23-qter (18 of 36) and 3q26.3 (18 of 36). High-level amplifications (defined as a localized chromosomal region showing a higher than 1.5 ratio of tumor to normal DNA hybridization) occurred in eight cases: five on chromosome 8q, two on 3q, and a single case on 11q. Further frequent amplifications involved 11q22 (11 of 36), 2q31–32 (8 of 36), 6cent-p23 (8 of 36), 13q22 (7 of 36), and 17q24–25 (7 of 36).
Summary of copy number changes in 36 familial ovarian carcinomas. Vertical lines to the left and right of the chromosome ideograms represent loss and gain of copy number, respectively. Thick bars indicate high-level amplification. Grey and black lines indicate tumors with and without breast cancer gene (BRCA) mutations, respectively.
The most frequently deleted regions were 8p21-pter (23 of 36), 16p22-pter (19 of 36), 22q13 (19 of 36), 9q31–33 (16 of 36), 12q24 (16 of 36), and 15q11–15 (16 of 36). Deletions also occurred at 17p12–13 (14 of 36), Xp21–22 (14 of 36), 20q13 (13 of 36), 15q24–25 (12 of 36), 18q21 (12 of 36), 11p15 (11 of 36), and 13q14 (11 of 36) (Table 1).
There were no statistically significant differences between familial tumors with and without a proven BRCA mutation. Analysis of tumor stage in relation to DNA copy number events identified that loss of 15q11–15 occurred more frequently in Stages I and II than in III and IV (p = 0.03, two-tailed Fisher exact test). Further differences, although not statistically significant, included a higher frequency of loss at 17p12–13 in Stages I and II than in III and IV, and a higher frequency of amplification of 8q23–24.1 in Stages III and IV than in I and II (p = 0.08 and 0.09, respectively). The small number of Grade 1, Grade 2, and nonserous histologic subtypes did not allow useful comparison of grade and histology with CGH results.
Discussion
We present the CGH analysis of a series of 36 cases of familial ovarian cancer based on a strong family history of ovarian (and breast) cancer in a least three first-degree relatives in two successive generations. Of these 36, mutation analysis has so far identified germline mutations in either the BRCA1 (n = 13) or BRCA2 (n = 2) genes in 15 cases. This is the largest published series so far subjected to this technique.
We detected, on average, 4 gains and 9.7 losses in copy number per tumor. This is relatively high compared with the only other comparable study published (Tapper et al, 1998). Tapper et al (1998) found approximately the same number of gains as we did, but the ratio of gains to losses was 3:1. This may be due to differences in the methodology used for DNA extraction. Using archival material and microdissection, we were able to accurately isolate pure (>80%) tumor DNA, whereas Tapper et al (1998) selected tissue containing more than 50% tumor cells without microdissection. Weiss et al (1999) demonstrated that copy losses easily become undetectable at 50% contamination of normal tissue. As a result, our technique using microdissected material is more sensitive to the detection of single copy chromosomal losses. Gains, however, are less sensitive to contamination because the gain is usually more than one copy, and a locus can only be lost once.
In our analysis we have excluded gains and losses of chromosome 16 and 19 because CGH data of these chromosomes is notoriously difficult to interpret due to the repetitive nature of the DNA surrounding the centromeres (Kallioniemi et al, 1994; Weiss et al, 1999).
Gains
A summary of the currently available data on CGH in ovarian cancer is shown in Table 2. Gains of 8q22-ter (50%) and 3q (50%) were common in this study. This is in accordance with other studies reporting CGH analysis of ovarian cancers unselected for family history (Arnold et al, 1996; Iwabuchi et al, 1995; Sonoda et al, 1997), as well as with a study on hereditary ovarian cancer (Tapper et al, 1998). The frequency of amplification of both 8q and 3q is similar in all studies and is probably explained by the localization of specific oncogenes already known to be involved in ovarian cancer. C-myc, localized on 8q24.12–13, is an oncogene the product of which is overexpressed in up to 30% of ovarian cancers (Baker et al, 1990). Gain of 3q material is a frequent event in several types of cancer, including head and neck cancer and cervical cancer. The PIK3CA gene important in signal transduction has been suggested to be the target of amplification of this region (Kapeller and Cantley, 1994). Recently this gene has been suggested to be involved in ovarian carcinogenesis (Shayesteh et al, 1999). Gains on 6cent-p23 were encountered in 22% of our familial ovarian cancers. Other researchers have found similar data on unselected ovarian cancer cases (Table 2). The amplification in this region may be due to the growth factor VEGFA localized on 6p21.3.
We found significantly more gains on chromosome 11q22, 13q22, and 17q24–25 than have studies on unselected ovarian cancer. The latter region was recently found to be frequently amplified in brain metastases of solid tumors, suggesting that this region is involved in tumor dissemination (Petersen et al, 2000).
Losses
Loss of chromosome 17p12–13 is frequent in both unselected and familial ovarian cancer (Table 2). The likely tumor suppressor gene responsible is TP53 located on 17p13.1. We recently reported accumulation of p53 protein to be an important event in both hereditary and sporadic ovarian cancer (Zweemer et al, 1999). The current study supports this notion. Loss of (part of) the X chromosome has been detected in unselected ovarian cancer cases (Table 2) and was also frequent in our tumor panel (14 of 36; 39%). Interestingly, inactivation of the X chromosome, specifically Xp22.2–3 of the active allele, has recently been suggested to encode a product that specifically interacts with BRCA1 in some hereditary ovarian cancers (Buekers et al, 2000). Chromosome 18q21 harbors three candidate tumor suppressor genes; the SMAD4 gene located on this band was recently found to be mutated in ovarian cancer (Takakura et al, 1999). We found 12 of 36 (33%) losses in this region; previously, Arnold et al (1996) also detected frequent loss of this region (11 of 47; 23%). CGH analysis of the region on chromosome 13q14, harboring the retinoblastoma (Rb), showed loss in 17% of unselected ovarian cancer cases (Arnold et al, 1996). We also found this region to be frequently lost (31%) in familial ovarian cancer. A final region of interest is 9q31–33 that showed loss in 16 of 36 (44%) familial ovarian cancer cases. Loss of 9q has been seen in unselected ovarian cancer (Table 2) and harbors a region that is commonly associated with bladder and renal cell carcinoma (Habuchi et al, 1997; Simoneau et al, 1996), although loss of heterozygosity (LOH) of this region has also been linked to metastatic breast cancer (Minobe et al, 1998; Nishizaki et al, 1997). Our analyses suggest that losses at 17p12–13, Xp22, 18q21, 13q14, and 9q31–33 in ovarian cancer are unrelated to hereditary predisposition.
When comparing the CGH copy number changes in familial cases with those in ovarian cancer cases unselected for family history described in the literature (Table 2), losses at 8p21-ter, 22q13, 15q.11–14, 15q.24–25, and 12q24 occur in high frequency in familial cases but are rare in sporadic (unselected) cases. Loss of 8p21-ter was the most frequent alteration in chromosome copy number that occurred in our panel of familial ovarian cancers (64%), as well as the most frequent region of loss in the familial cases studied by Tapper et al (1998). Losses at 8p21-ter, detected by CGH, were not common in sporadic cancers in the studies of Iwabuchi et al (1995), Arnold et al (1996), and Sonoda et al (1997). There is mounting evidence from the literature that this region harbors an important tumor suppressor gene involved in a range of malignancies (Ishii et al, 1999), including laryngeal tumors (Rizos et al, 1998), prostate cancer (Watanabe et al, 1998), and sporadic female breast cancer (Hermsen et al, 1998; Seitz et al, 1997; Wang et al, 1999), as well as male breast cancer (Tirkkonen et al, 1999). One candidate gene for this locus may be the Frizzled-related gene (Ugolini et al, 1999). Loss of chromosome region 22q13 is the second most frequent event in this study. Several LOH studies of sporadic ovarian cancer have demonstrated that chromosomal region 22q is frequently lost (Bryan et al, 1996; Cliby et al, 1993). However, this is the first report of significant chromosome arm 22q loss detected by CGH in familial ovarian cancer. 22q13 harbors several target genes in the minimally deleted region recently defined by further LOH studies; none have yet been found to show mutations in sporadic ovarian cancer (Bryan et al, 2000). In the current study, 19 of 36 ovarian cancer cases displayed evidence of loss of all or parts of chromosome arm 15q, making it (and 16q) the third most frequently deleted chromosome arm in our familial ovarian tumors.
The patterns of loss clearly defined two distinct regions, 15q11–15 and 15q24–25, with several cases showing selective loss of these regions and retention of the intervening region 15q21–22 (Fig. 2). There was a significantly higher frequency of loss at 15q11–15 in Stages I and II than in Stages III and IV (p = 0.03). This implies that ovarian cancers presenting in an early stage are cytogenetically different, and these copy number changes may have prognostic significance. This requires further investigation in larger series. Losses at 15q11–15 and 15q24–25 have not previously been reported in familial or sporadic ovarian carcinoma. However, 17% of uterine (Pere et al, 1998) and 73% of pancreatic adenocarcinoma (Mahlamaki et al, 1997) have been shown to demonstrate loss of 15q by CGH. The upper of the two regions (15q11–15) harbors three possible candidates. A tumor suppressor gene involved in advanced stage breast cancer has been identified at 15q14 (Wick et al, 1996). The region 15q15 harbors the thrombospondin-1 gene, which is thought to be an inhibitor of angiogenesis. However, the most promising candidate gene on 15q15 is the human homolog of the Escherichia coli RecA gene, hRAD51. BRCA1 and BRCA2 have been found to bind and complex with Rad51 and may together be involved in the repair of double-strand DNA-breaks (Scully et al, 1997; Sharan et al, 1997).
Loss of 12q has not been described by other CGH studies in ovarian cancer, nor could we find any cytogenetic or molecular studies describing the involvement of this chromosome arm in epithelial ovarian cancer. Its loss may therefore be specific to familial ovarian cancer. These loci of interest will be subjected to further studies by microarray CGH to narrow down the deleted regions and by microarray expression analysis to analyze target genes.
The only statistically significant difference between BRCA-related familial tumors and sporadic tumors reported by Tapper et al (1998) was amplification of 2q24–q32 (50% of familial ovarian tumors displayed amplification versus 10% of sporadic tumors). Of our 36 familial tumors only eight displayed amplification of this region (22%).
Loss of the BRCA1 locus (17q21) was rare (10 of 36) in the familial tumors studied in this analysis, which can be explained by the limited resolution of the CGH technique (approximately 10 Mb), whereas the “second hit” according to Knudsen's hypothesis is usually a small deletion or even a point mutation.
In conclusion, hereditary ovarian cancer differs from sporadic ovarian cancer not only by the involvement of germline BRCA mutations, but also by somatic chromosomal losses at 8p21-ter, 22q13, 15q11–15, 15q24–25, and 12q24. This suggests a role for tumor suppressor genes specifically associated with hereditary ovarian cancer. In addition, significantly more gains were seen at 11q22, 13q22, and 17q24–25, pointing to the involvement of specific oncogenes in the carcinogenic pathway of familial ovarian cancer. It remains to be seen whether these differences also explain clinical and histopathologic differences between hereditary and sporadic ovarian cancer.
Materials and Methods
Sample Selection
A total of 35 ovarian and 1 primary peritoneal serous adenocarcinoma cases were selected on the basis of a strong family history of ovarian and/or breast cancer in at least three first-degree relatives in two successive generations. Distribution of stage, grade histology, as well as type of BRCA-mutation are represented in Table 3. Mutation analysis had previously identified germline mutations in BRCA1 (n = 13) and BRCA2 (n = 2). In 21 cases, no mutations were detected. Screening for germline mutations in BRCA1 and BRCA2 was performed using a protein truncation test (PTT)–based assay, as previously described (Hogervorst et al, 1995). All patients were screened for Askenazi-Jewish founder mutations (185delAG, 5382insC) by direct sequencing. In addition we screened for two common Dutch founder mutations, the BRCA1 exon 13 and exon 22 genomic deletions, using a PCR-based assay (Petrij-Bosch et al, 1997).
DNA Extraction
To accurately obtain tumor-specific DNA, all of the samples were extracted from paraffin-embedded formalin-fixed archival tissue, as previously described (Weiss et al, 1999). Briefly, sections were hematoxylin stained and areas of malignant tissue were identified and marked by a gynecologic pathologist. Tumor cells were isolated by microdissecting all areas containing more than 80% tumor cells from 10 uncovered sections. The procedure was performed by hand, on an inverting preparation microscope, with a glass micropipette and/or a small surgical knife. After overnight incubation in 1 m sodium thiocyanate at 37° C, the DNA was extracted by treatment with 2 mg/ml proteinase K at 55° C for 72 hours. Finally, the DNA was purified using a QIAGEN DNA purification column (QIAGEN, Crawley, West Sussex, United Kingdom).
CGH
The CGH procedure was performed as described by Meijer et al (1998) and Weiss et al (1999). In short, tumor DNA was labeled with a green fluorochrome, mixed with red labeled normal (diploid) DNA of comparable fragment length, and hybridized to normal metaphase preparations. The green to red fluorescence ratio on the chromosomes (calculated by image analysis software; Applied Imaging, Newcastle-upon-Tyne, United Kingdom) is a measure of under- or overrepresentation of genetic material of the tumor (loss or gain, respectively). For each tumor sample, 10 to 15 metaphases were selected and the averaged fluorescence ratios and their 95% confidence intervals (CI) were plotted along ideograms of the corresponding chromosomes in a relative copy number karyotype. Deviations from normal were interpreted as gains or losses when the 95% CI of the fluorescence ratio did not include 1.0. Chromosomes 1pter, 16p, and 19 were not included in the analysis for reasons of unreliability due to repetitive sequences located within these regions (Kallioniemi et al, 1994).
References
Arnold N, Hagele L, Walz L, Schempp W, Pfisterer J, Bauknecht T, and Kiechle M (1996). Overrepresentation of 3q and 8q material and loss of 18q material are recurrent findings in advanced human ovarian cancer. Genes Chromosomes Cancer 16: 46–54.
Baker VV, Borst MP, Dixon D, Hatch KD, Shingleton HM, and Miller D (1990). c-myc amplification in ovarian cancer. Gynecol Oncol 38: 340–342.
Boyd J (1998). Molecular genetics of hereditary ovarian cancer. Oncology 12: 399–406.
Bryan EJ, Thomas NA, Palmer K, Dawson E, Englefield P, and Campbell IG (2000). Refinement of an ovarian cancer tumour suppressor gene locus on chromosome arm 22q and mutation analysis of CYP2D6, SREBP2 and NAGA. Int J Cancer 87: 798–802.
Bryan EJ, Watson RH, Davis M, Hitchcock A, Foulkes WD, and Campbell IG (1996). Localization of an ovarian cancer tumor suppressor gene to a 0.5-cM region between D22S284 and CYP2D, on chromosome 22q. Cancer Res 56: 719–721.
Buekers TE, Lallas TA, and Buller RE (2000). Xp22.2-3 loss of heterozygosity is associated with germline BRCA1 mutation in ovarian cancer. Gynecol Oncol 76: 418–420.
Cliby W, Ritland S, Hartmann L, Dodson M, Halling KC, Keeney G, Podratz KC, and Jenkins RB (1993). Human epithelial ovarian cancer allelotype. Cancer Res 53 (Suppl 10): 2393–2398.
Foster KA, Harrington P, Kerr J, Russell P, DiCioccio RA, Scott IV, Jacobs I, Chenevix-Trench G, Ponder BA, and Gayther SA (1996). Somatic and germline mutations of the BRCA2 gene in sporadic ovarian cancer. Cancer Res 56: 3622–3625.
Habuchi T, Yoshida O, and Knowles MA (1997). A novel candidate tumour suppressor locus at 9q32-33 in bladder cancer: Localization of the candidate region within a single 840 kb YAC. Hum Mol Genet 6: 913–919.
Hermsen MA, Baak JP, Meijer GA, Weiss JM, Walboomers JW, Snijders PJ, and Van Diest PJ (1998). Genetic analysis of 53 lymph node-negative breast carcinomas by CGH and relation to clinical, pathological, morphometric, and DNA cytometric prognostic factors. J Pathol 186: 356–362.
Hogervorst FBL, Cornelis RS, Bout M, van Vliet M, Oosterwijk JC, Olmer R, and Bakker B (1995). Rapid detection of BRCA1 mutations by the protein truncation test. Nat Genet 10: 208–212.
Ishii H, Baffa R, Numata SI, Murakumo Y, Rattan S, Inoue H, Mori M, Fidanza V, Alder H, and Croce CM (1999). The FEZ1 gene at chromosome 8p22 encodes a leucine-zipper protein, and its expression is altered in multiple human tumors. Proc Natl Acad Sci USA 96: 3928–3933.
Iwabuchi H, Sakamoto M, Sakunaga H, Ma YY, Carcangiu ML, Pinkel D, Yang-Feng TL, and Gray JW (1995). Genetic analysis of benign, low-grade, and high-grade ovarian tumors. Cancer Res 55: 6172–6180.
Kallioniemi OP, Kallioniemi A, Piper J, Isola J, Waldman FM, Gray JW, and Pinkel D (1994). Optimizing comparative genomic hybridization for analysis of DNA sequence copy number changes in solid tumors. Genes Chromosomes Cancer 10: 231–243.
Kapeller R and Cantley LC (1994). Phosphatidylinositol 3-kinase. Bioessays 16: 565–576.
Koper NP, Kiemeney LA, Massuger LF, Thomas CM, Schijf CP, and Verbeek AL (1996). Ovarian cancer incidence (1989–1991) and mortality (1954–1993) in The Netherlands. Obstet Gynecol 88: 387–393.
Mahlamaki EH, Hoglund M, Gorunova L, Karhu R, Dawiskiba S, Andren-Sandberg A, Kallioniemi OP, and Johansson B (1997). Comparative genomic hybridization reveals frequent gains of 20q, 8q, 11q, 12p, and 17q, and losses of 18q, 9p, and 15q in pancreatic cancer. Genes Chromosomes Cancer 20: 383–391.
Meijer GA, Hermsen MA, Baak JP, van Diest PJ, Meuwissen SG, Belien JA, Hoovers JM, Joenje H, Snijders PJ, and Walboomers JM (1998). Progression from colorectal adenoma to carcinoma is associated with nonrandom chromosomal gains as detected by comparative genomic hybridisation. J Clin Pathol 51: 901–909.
Merajver SD, Pham TM, Caduff RF, Chen M, Poy EL, Cooney KA, Weber BL, Collins FS, Johnston C, and Frank TS (1995). Somatic mutations in the BRCA1 gene in sporadic ovarian tumours. Nat Genet 9: 439–443.
Minobe K, Onda M, Iida A, Kasumi F, Sakamoto G, Nakamura Y, and Emi M (1998). Allelic loss on chromosome 9q is associated with lymph node metastasis of primary breast cancer. Jpn J Cancer Res 89: 916–922.
Nishizaki T, DeVries S, Chew K, Goodson WH 3rd, Ljung BM, Thor A, and Waldman FM (1997). Genetic alterations in primary breast cancers and their metastases: Direct comparison using modified comparative genomic hybridization. Genes Chromosomes Cancer 19: 267–272.
Pere H, Tapper J, Wahlstrom T, Knuutila S, and Butzow R (1998). Distinct chromosomal imbalances in uterine serous and endometrioid carcinomas. Cancer Res 58: 892–895.
Petersen I, Hidalgo A, Petersen S, Schluns K, Schewe C, Pacyna-Gengelbach M, Goeze A, Krebber B, Knosel T, Kaufmann O, Szymas J, and Von Deimling A (2000). Chromosomal imbalances in brain metastases of solid tumors. Brain Pathol 10: 395–401.
Petrij-Bosch A, Peelen T, van Vliet M, van Eijk R, Olmer R, Drusedau M, Hogervorst FB, Hageman S, Arts PJ, Ligtenberg MJ, Meijers-Heijboer H, Klijn JG, Vasen HF, Cornelisse CJ, van't Veer LJ, Bakker E, van Ommen GJ, and Devilee P (1997). BRCA1 genomic deletions are major founder mutations in Dutch breast cancer patients. Nat Genet 17: 341–345.
Rizos E, Sourvinos G, and Spandidos DA (1998). Loss of heterozygosity at 8p, 9p and 17q in laryngeal cytological specimens. Oral Oncol 34: 519–523.
Scully R, Chen J, Plug A, Xiao Y, Weaver D, Feunteun J, Ashley T, and Livingston DM (1997). Association of BRCA1 with Rad51 in mitotic and meiotic cells. Cell 88: 265–275.
Seitz S, Rohde K, Bender E, Nothnagel A, Pidde H, Ullrich OM, El-Zehairy A, Haensch W, Jandrig B, Kolble K, Schlag PM, and Scherneck S (1997). Deletion mapping and linkage analysis provide strong indication for the involvement of the human chromosome region 8p12-p22 in breast carcinogenesis. Br J Cancer 76: 983–991.
Sharan SK, Morimatsu M, Albrecht U, Lim DS, Regel E, Dinh C, Sands A, Eichele G, Hasty P, and Bradley A (1997). Embryonic lethality and radiation hypersensitivity mediated by Rad51 in mice lacking Brca2. Nature 386: 804–810.
Shayesteh L, Lu Y, Kuo WL, Baldocchi R, Godfrey T, Collins C, Pinkel D, Powell B, Mills GB, and Gray JW (1999). PIK3CA is implicated as an oncogene in ovarian cancer. Nat Genet 21: 99–102.
Simoneau AR, Spruck CH 3rd, Gonzalez-Zulueta M, Gonzalgo ML, Chan MF, Tsai YC, Dean M, Steven K, Horn T, and Jones PA (1996). Evidence for two tumor suppressor loci associated with proximal chromosome 9p toq and distal chromosome 9q in bladder cancer and the initial screening for GAS1and PTC mutations. Cancer Res 56: 5039–5043.
Sonoda G, Palazzo J, du Manoir S, Godwin AK, Feder M, Yakushiji M, and Testa JR (1997). Comparative genomic hybridization detects frequent overrepresentation of chromosomal material from 3q26, 8q24, and 20q13 in human ovarian carcinomas. Genes Chromosomes Cancer 20: 320–328.
Takakura S, Okamoto A, Saito M, Yasuhara T, Shinozaki H, Isonishi S, Yoshimura T, Ohtake Y, Ochiai K, and Tanaka T (1999). Allelic imbalance in chromosome band 18q21 and SMAD4 mutations in ovarian cancers. Genes Chromosomes Cancer 24: 264–271.
Tapper J, Sarantaus L, Vahteristo P, Nevanlinna H, Hemmer S, Seppala M, Knuutila S, and Butzow R (1998). Genetic changes in inherited and sporadic ovarian carcinomas by comparative genomic hybridization: Extensive similarity except for a difference at chromosome 2q24-q32. Cancer Res 58: 2715–2719.
Tirkkonen M, Kainu T, Loman N, Johannsson OT, Olsson H, Barkardottir RB, Kallioniemi OP, and Borg A (1999). Somatic genetic alterations in BRCA2-associated and sporadic male breast cancer. Genes Chromosomes Cancer 24: 56–61.
Ugolini F, Adelaide J, Charafe-Jauffret E, Nguyen C, Jacquemier J, Jordan B, Birnbaum D, and Pebusque MJ (1999). Differential expression assay of chromosome arm 8p genes identifies Frizzled-related (FRP1/FRZB) and fibroblast growth factor receptor 1 (FGFR1) as candidate breast cancer genes. Oncogene 18: 1903–1910.
Wang JC, Radford DM, Holt MS, Helms C, Goate A, Brandt W, Parik M, Phillips NJ, DeSchryver K, Schuh ME, Fair KL, Ritter JH, Marshall P, and Donis-Keller H (1999). Sequence-ready contig for the 1.4-cM ductal carcinoma in situ loss of heterozygosity region on chromosome 8p22-p23. Genomics 60: 1–11.
Watanabe M, Shiraishi T, Muneyuki T, Nagai M, Fukutome K, Murata M, Kawamura J, and Yatani R (1998). Allelic loss and microsatellite instability in prostate cancers in Japan. Oncology 55: 569–574.
Weiss MM, Hermsen MA, Meijer GA, van Grieken NC, Baak JP, Kuipers EJ, and Van Diest PJ (1999). Comparative genomic hybridisation. Mol Pathol 52: 243–251.
Wick W, Petersen I, Schmutzler RK, Wolfarth B, Lenartz D, Bierhoff E, Hummerich J, Muller DJ, Stangl AP, Schramm J, Wiestler OD, and Von Deimling A (1996). Evidence for a novel tumor suppressor gene on chromosome 15 associated with progression to a metastatic stage in breast cancer. Oncogene 12: 973–978.
Zweemer RP, Verheijen RH, Gille JJ, van Diest PJ, Pals G, and Menko FH (1998). Clinical and genetic evaluation of thirty ovarian cancer families. Am J Obstet Gynecol 178: 85–90.
Zweemer RP, Shaw PA, Verheijen RM, Ryan A, Berchuck A, Ponder BA, Risch H, McLaughlin JR, Narod SA, Menko FH, Kenemans P, and Jacobs IJ (1999). Accumulation of p53 protein is frequent in ovarian cancers associated with BRCA1and BRCA2 germline mutations. J Clin Pathol 52: 372–375.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zweemer, R., Ryan, A., Snijders, A. et al. Comparative Genomic Hybridization of Microdissected Familial Ovarian Carcinoma: Two Deleted Regions on Chromosome 15q Not Previously Identified in Sporadic Ovarian Carcinoma. Lab Invest 81, 1363–1370 (2001). https://doi.org/10.1038/labinvest.3780350
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3780350
This article is cited by
-
DNA copy number profiling reveals extensive genomic loss in hereditary BRCA1 and BRCA2 ovarian carcinomas
British Journal of Cancer (2013)
-
Molecular and immunohistochemical evidence for the origin of uterine leiomyosarcomas from associated leiomyoma and symplastic leiomyoma-like areas
Modern Pathology (2009)
-
Genome-wide-array-based comparative genomic hybridization reveals genetic homogeneity and frequent copy number increases encompassing CCNE1 in Fallopian tube carcinoma
Oncogene (2003)