Abstract
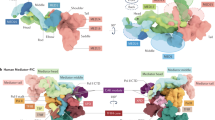
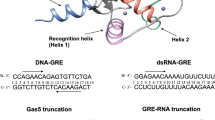
LYMPHOID enhancer-binding factor (LEF-1) and the closely related T-cell factor 1 (TCF-1) are sequence-specific and cell-type-specific DNA-binding proteins that play important regulatory roles in organogenesis and thymocyte differentiation1–5. LEF-1 participates in regulation of the enhancer associated with the T cell receptor (TCR)-α gene by inducing a sharp bend in the DNA and facilitating interactions between Ets-1, PEBP2-α, and ATF/ CREB transcription factors bound at sites flanking the LEF-1 site1,2,6,7. It seems that LEF-1 plays an architectural role in the assembly and function of this regulatory nucleoprotein complex7,8. LEF-1 recognizes a specific nucleotide sequence through a high-mobility-group (HMG) domain1,2. Proteins containing HMG domains bind DNA in the minor groove, bend the double helix6,9,10, and recognize four-way junctions and other irregular DNA structures9,11. Here we report the solution structure of a complex of the LEF-1 HMG domain and adjacent basic region with its cognate DNA. The structure reveals the HMG domain bound in the widened minor groove of a markedly distorted and bent double helix. The basic region binds across the narrowed major groove and contributes to DNA recognition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Travis, A., Amsterdam, A., Belanger, C. & Grosschedl, R. Genes Dev. 5, 880–894 (1991).
Waterman, M. L., Fischer, W. H. & Jones, K. A. Genes Dev. 5, 656–669 (1991).
Oosterwegel, M. et al. J. exp. Med. 173, 1133–1142 (1991).
Van Genderen, C. et al. Genes Dev. 8, 2691–2703 (1994).
Verbeek, S. et al. Nature 374, 70–74 (1995).
Giese, K., Cox, J. & Grosschedl, R. Cell 69, 185–195 (1992).
Giese, K., Kingsley, C., Kirshner, J. R. & Grosschedl, R. Genes Dev. 9, 995–1008 (1995).
Grosschedl, R., Giese, K. & Pagel, J. Trends Genet. 10, 94–100 (1994).
Ferrari, S. et al. EMBO J. 11, 4497–4506 (1992).
Paull, T. T., Haykinson, M. J. & Johnson, R. C. Genes Dev. 7, 1521–1534 (1993).
Pil, P. M. & Lippard, S. J. Science 256, 234–237 (1992).
Weir, H. M. et al. EMBO J. 12, 1311–1319 (1993).
Read, C. M., Cary, P. D., Crane-Robinson, C., Driscoll, P. C. & Norman, D. G. Nucleic Acids Res. 21, 3427–3436 (1993).
Jones, D. N. M. et al. Structure 2, 609–627 (1994).
Giese, K., Amsterdam, A. & Grosschedl, R. Genes Dev. 5, 2567–2578 (1991).
van de Wetering, M. & Clevers, H. EMBO J. 11, 3039–3044 (1992).
Peters, R. et al. Biochemistry 34, 4569–4576 (1995).
Carlsson, P., Waterman, M. L. & Jones, K. A. Genes Dev. 7, 2418–2430 (1993).
Read, C. M., Cary, P. D., Preston, N. S., Lnenicek-Allen, M. & Crane-Robinson, C. EMBO J. 13, 5639–5646 (1994).
Harley, V. R., Lovell-Badge, R. & Goodfellow, P. N. Nucleic Acids Res. 22, 1500–1501 (1994).
Kim, Y., Geiger, J. H., Hahn, S. & Sigler, P. B. Nature 365, 512–520 (1993).
Kim, J. L., Nikolov, D. B. & Burley, S. K. Nature 365, 520–528 (1993).
Schumacher, M. A., Choi, K. Y., Zalkin, H. & Brennan, R. G. Science 266, 763–770 (1994).
Marion, D., Kay, L. E., Sparks, S. W., Torchia, D. A. & Bax, A. J. Am. chem. Soc. 111, 1515–1517 (1989).
Güntert, P., Braun, W. & Wüthrich, K. J. molec. Biol. 217, 517–530 (1991).
Güntert, P. & Wüthrich, K. J. Biomol. NMR 1, 447–456 (1991).
Weiner, S. J., Kollman, P. A., Nguyen, D. T. & Case, D. A. J. comput. Chem. 7, 230–252 (1986).
Seip, S., Balbach, J. & Kessler, H. J. magn. Reson. B104, 172–179 (1994).
Otting, G. & Wüthrich, K. Q. Rev. Biophys. 23, 39–96 (1990).
Lavery, R. & Sklenár, V. J. biomolec. Struct. Dyn. 6, 63–91 (1988).
Werner, M. H., Huth, J. R., Gronenborn, A. M. & Clore, G. M. Cell 81, 705–714 (1995).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Love, J., Li, X., Case, D. et al. Structural basis for DNA bending by the architectural transcription factor LEF-1. Nature 376, 791–795 (1995). https://doi.org/10.1038/376791a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/376791a0
This article is cited by
-
Structural basis of the interaction between BCL9-Pygo and LDB-SSBP complexes in assembling the Wnt enhanceosome
Nature Communications (2023)
-
TCF7L2 acts as a molecular switch in midbrain to control mammal vocalization through its DNA binding domain but not transcription activation domain
Molecular Psychiatry (2023)
-
Intrinsically disordered domain of transcription factor TCF-1 is required for T cell developmental fidelity
Nature Immunology (2023)
-
Tcf1 preprograms the mobilization of glycolysis in central memory CD8+ T cells during recall responses
Nature Immunology (2022)
-
TCF-1 promotes chromatin interactions across topologically associating domains in T cell progenitors
Nature Immunology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.