Abstract
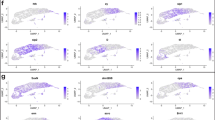
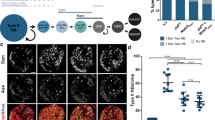
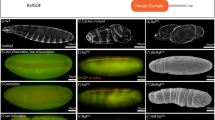
THE Drosophila central nervous system develops from a segment-ally reiterated array of 30 unique neural precursors, called neuro-blasts. Each neuroblast goes through a stereotyped cell lineage to produce an invariant clone of neural progeny. It is critical to identify the genes that specify neuroblast identity as these genes control the time of formation, gene expression profile, and cell lineage characteristics of each neuroblast. Here we show that the Pax-type gooseberry-distal gene specifies row 5 neuroblast identity. Initially, four rows of neuroblasts form per segment (1,3,5,7) and gooseberry-distal is expressed in row 5 neuroblasts1á¤-3. By using 10 molecular markers, and by following the number and orientation of neuroblast divisions, we show that lack of gooseberry-distal transforms row 5 neuroblasts into row 3 neuroblasts, whereas ubiquitous gooseberry-distal generates the reciprocal transformation. Thus, gooseberry-distal is necessary and sufficient to specify row 5 neuroblast identity autonomously. The 10 genes coordinately regulated by gooseberry-distal are prime candidates for controlling specific aspects of neuroblast identity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Doe, C. Q. Development 119, 855–863 (1992).
Gutjahr, T., Patel, N. H., Li, X., Goodman, C. S. & Noll, M. Development 119, 21–31 (1993).
Zhang, Y., Ungar, A., Fresquez, C. & Holmgren, R. Development 120, 1151–1161 (1994).
Goodman, C. S. & Doe, C. Q. in The Development of Drosophila (eds Bate, M. & Martinez-Arias, A.) 1131–1206 (Cold Spring Harbor Laboratory Press, New York, 1993).
Jimenez, F. & Campos-Ortega, J. A. Neuron 5, 81–89 (1990).
Cabrera, C. V., Martinez-Arias, A. & Bate, M. Cell 50, 425–433 (1987).
Martin-Bermudo, M. D., Gonzalez, F., Dominguez, M., Rodriguez, L. & Jimenez, F. Development 119, 1003–1012 (1991).
Skeath, J. B. & Carroll, S. B. Development 114, 939–946 (1992).
Akam, M. Development 101, 1–21 (1987).
Patel, N. H., Schafer, B., Goodman, C. S. & Holmgren, R. Genes Dev. 3, 890–904 (1989).
Chu-LaGraff, Q. & Doe, C. Q. Science 261, 1594–1597 (1993).
Bopp, D., Burri, M., Baumgartner, S., Frigerio, G. & Noll, M. Cell 47, 1033–1040 (1986).
Baumgartner, S., Bopp, D., Burri, M. & Noll, M. Genes Dev. 1, 1247–1267 (1987).
Chalepakis, G., Stoykova, A., Wijnholds, J., Tremblay, P. & Gruss, P. J. Neurobiol 24, 1367–1384 (1993).
Cote, S. et al. EMBO J. 6, 2793–2801 (1987).
Skeath, J. B., Panganiban, G. F. & Carroll, S. B. Genes Dev. 6, 2606–2619 (1992).
Doe, C. Q., Hiromi, Y., Gehring, W. J. & Goodman, C. S. Science 239, 190–195 (1988).
Ingham, P. W. & Martinez-Arias, A. Cell 68, 221–235 (1992).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Skeath, J., Zhang, Y., Holmgren, R. et al. Specification of neuroblast identity in the Drosophila embryonic central nervous system by gooseberry-distal. Nature 376, 427–430 (1995). https://doi.org/10.1038/376427a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/376427a0
This article is cited by
-
Specification of the Drosophila Orcokinin A neurons by combinatorial coding
Cell and Tissue Research (2023)
-
Early embryonic development of Johnston’s organ in the antenna of the desert locust Schistocerca gregaria
Development Genes and Evolution (2022)
-
Integration of temporal and spatial patterning generates neural diversity
Nature (2017)
-
Early development, pattern, and reorganization of the planula nervous system in Aurelia (Cnidaria, Scyphozoa)
Development Genes and Evolution (2008)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.