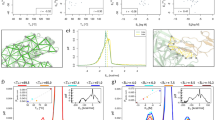
Abstract
RESIDUES in β-sheets occur in two distinct tertiary contexts: central strands, bordered on both sides by other β-strands, and edge strands, bordered on only a single side by another β-strand1. The ΔΔG values for β-sheet formation measured at an edge β-strand of the IgG-binding domain of protein G(GB1) are quite different from those obtained previously2,3 at a central position in the same protein. In particular, there is no correlation at the edge position with statistically determined β-sheet-forming preferences4. The differences between β-sheet propensities measured at central and edge β-strands, ΔΔΔG values, correlate with the values of water/octanol transfer free energies5 and side-chain non-polar surface area for the amino acids6. These results strongly suggest that, unlike α-helix formation, β-sheet formation is determined in large part by tertiary context, even at solvent-accessible sites, and not by intrinsic secondary structure preferences.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Chothia, C. J. molec. Biol. 105, 1–14 (1976).
Minor, D. L. Jr & Kim, P. S. Nature 367, 660–663 (1994).
Smith, C. K., Withka, J. M. & Regan, L. Biochemistry 33, 5510–5517 (1994).
Chou, P. Y. & Fasman, G. D. Biochemistry 13, 211–222 (1973).
Fauchere, J.-L. & Pliska, V. Eur. J. med. Chem.-Chim. Ther. 18, 369–375 (1983).
Livingstone, J. R., Spolar, R. S. & Record, M. T. Jr Biochemistry 30, 4237–4244 (1991).
Alexander, P., Fahnestock, S., Lee, T., Orban, J. & Bryan, P. Biochemistry 31, 3597–3603 (1992).
Frick, I.-M. et al. Proc. natn. Acad. Sci. U. S. A. 89, 8532–8536 (1992).
Gronenborn, A. M. & Clore, G. M. J. molec. Biol. 223, 331–335 (1993).
McGregor, M. J., Islam, S. A. & Sternberg, M. J. E. J. molec. Biol. 198, 295–310 (1987).
Lyu, P. C., Liff, M. I., Marky, L. A. & Kallenbach, N. R. Science 250, 669–673 (1990).
O'Neil, K. T. & DeGrado, W. F. Science 250, 646–651 (1990).
Padmanabhan, S., Marqusee, S., Ridgeway, T., Laue, T. M. & Baldwin, R. L. Nature 344, 268–270 (1990).
Wojcik, J., Altman, K.-H. & Scheraga, H. A. Biopolymers 30, 121–134 (1990).
Chakrabartty, A., Schellman, J. A. & Baldwin, R. L. Nature 351, 586–588 (1991).
Horovitz, A., Matthews, J. M. & Fersht, A. R. J. molec. Biol. 227, 560–568 (1992).
Serrano, I., Neira, J.-L., Sancho, J. & Fersht, A. R. Nature 356, 453–455 (1992).
Blaber, M., Zhang, X. & Matthews, B. W. Science 260, 1637–1640 (1993).
Kim, C. A. & Berg, J. M. Nature 362, 267–270 (1993).
Lifson, S. & Sander, C. J. molec. Biol. 139, 627–639 (1980).
Priestle, J. P. J. appl. Crystallogr. 21, 572–576 (1988).
Gronenborn, A. M. et al. Science 253, 657–661 (1991).
Oas, T. G. & Kim, P. S. Nature 336, 42–48 (1988).
Kunkel, T. A., Roberts, J. D. & Zakour, R. A. Meth. Enzym. 154, 367–382 (1987).
Edelhoch, H. Biochemistry 6, 1948–1954 (1967).
Grathwohl, C. & Wüthrich, K. Biopolymers 20, 2623–2633 (1981).
Mclntosh, L. P., Wand, A. J., Lowry, D. F., Redfield, A. G. & Dahlquist, F. W. Biochemistry 29, 6341–6362 (1990).
Wüthrich, K. NMR of Proteins and Nucleic Acids (John Wiley, New York, 1986).
Rosner, B. Fundamentals of Biostatistics (Duxbury, Boston, 1982).
Fahnestock, S. R., Alexander, P., Filpula, D. & Nagle, J. in Bacterial Immunoglobin-Binding Proteins (ed. Boyle, M. D. P.) (Academic, San Diego, 1990).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Minor, D., Kim, P. Context is a major determinant of β-sheet propensity. Nature 371, 264–267 (1994). https://doi.org/10.1038/371264a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/371264a0
This article is cited by
-
Global versus local mechanisms of temperature sensing in ion channels
Pflügers Archiv - European Journal of Physiology (2018)
-
Efficient molecular mechanics simulations of the folding, orientation, and assembly of peptides in lipid bilayers using an implicit atomic solvation model
Journal of Computer-Aided Molecular Design (2011)
-
Systematic assessment of accuracy of comparative model of proteins belonging to different structural fold classes
Journal of Molecular Modeling (2011)
-
Free-floating ultrathin two-dimensional crystals from sequence-specific peptoid polymers
Nature Materials (2010)
-
A Reexamination of Correlations of Amino Acids with Particular Secondary Structures
The Protein Journal (2009)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.