Abstract
During pregnancy some maternal cells reach the fetal circulation. Microchimerism (Mc) refers to low levels of genetically disparate cells or DNA. Maternal Mc has recently been found in the peripheral blood of healthy adults. We asked whether healthy women have maternal Mc in T and B lymphocytes, monocyte/macrophages and NK cells and, if so, at what levels. Cellular subsets were isolated after fluorescence activated cell sorting. A panel of HLA-specific real-time quantitative PCR assays was employed targeting maternal-specific HLA sequences. Maternal Mc was expressed as the genome equivalent (gEq) number of microchimeric cells per 100 000 proband cells. Thirty-one healthy adult women probands were studied. Overall 39% (12/31) of probands had maternal Mc in at least one cellular subset. Maternal Mc was found in T lymphocytes in 25% (7/28) and B lymphocytes in 14% (3/21) of probands. Maternal Mc levels ranged from 0.9 to 25.6 and 0.9 to 25.3 gEq/100 000 in T and B lymphocytes, respectively. Monocyte/macrophages had maternal Mc in 16% (4/25) and NK cells in 28% (5/18) of probands with levels from 0.3 to 36 and 1.8 to 3.2 gEq/100 000, respectively. When compared to fetal Mc, as assessed by quantification of male DNA in women with sons, maternal Mc was substantially less prevalent in all cellular subsets; fetal Mc prevalence in T and B lymphocytes, monocyte/macrophages and NK cells was 58, 75, 50 and 62% (P=0.01, P=0.005, P=0.04, P=0.05) respectively. In summary, maternal Mc was identified among lymphoid and myeloid compartments of peripheral blood in healthy adult women. Maternal Mc was less frequent than fetal Mc in all cellular subsets tested. Studies are needed to investigate the immunological effects and function of maternal Mc and to explore whether maternal Mc in cellular subsets has biological effects on her progeny.
Similar content being viewed by others
Main
During normal human pregnancy, some bidirectional cell trafficking occurs between the mother and the fetus.1 Fetal cells have been identified in the maternal circulation,2 and maternal cells in umbilical cord blood.3, 4, 5 The type of maternal cells transferred is unknown although limited information describes maternal T cells in cord blood.4, 6 Long-term persistence of fetal cells in the mother and maternal cells in her immune competent progeny have been described, referred to as fetal and maternal microchimerism (Mc), respectively.7, 8 Most studies of fetal Mc have employed assays that detect male DNA in women who gave birth to sons. Other approaches are necessary for identification and quantification of maternal Mc. For this purpose, we developed a panel of real-time quantitative PCR (Q-PCR) assays that target nonshared maternal-specific HLA sequences. In initial studies of unfractionated peripheral blood mononuclear cells (PBMC) we found maternal Mc in 22% of healthy women.9
The biological significance of long-lasting Mc to the benefit and/or detriment of the host is currently unknown. The potential breadth of maternal Mc effects in her progeny is considerable including, for example, effects on development, tissue repair, diversity in response to infections, immune surveillance and autoimmunity. We previously investigated peripheral blood samples from women with the autoimmune disease systemic sclerosis and found elevated levels of fetal Mc compared to healthy women,10 but not of maternal Mc.9 In other studies maternal Mc prevalence was increased in peripheral blood and in muscles of children with dermatomyositis11 and idiopathic myositis12 compared to controls. Maternal Mc was examined in cellular subsets in one study but for only two healthy subjects13 and in the other for nine healthy subjects in CD4 and CD8 only.12 Thus, information regarding maternal Mc in cellular subsets is needed for at least three reasons including the potential for a wide variety of effects on her progeny, as a foundation for comparison with disease states and because maternal Mc persistence into adult life in the circulation and in tissues intrinsically raises questions regarding immunological ‘self’ tolerance. We therefore conducted studies to determine the prevalence and quantitative levels of maternal Mc among T lymphocytes, B lymphocytes, monocyte/macrophages and NK cells in healthy adults.
Materials and methods
Subjects
Healthy women probands were recruited if the proband's mother was also willing to participate. (The term proband is used to refer to the index study subject to obviate any potential confusion in discussing maternal Mc in that women who are parous are also mothers.) A further requirement for probands who were parous was that all children be willing to participate. The inclusion criteria for a woman as a ‘healthy’ proband in our studies was that she did not have an autoimmune disease as defined by Rose and Bona.14 Family demographic, health issues and pregnancy history were collected by means of a questionnaire filled out by probands. A total of 110 subjects were recruited including 31 probands, 31 mothers and 48 children. The mean age of the proband at the time of the blood draw was 39 years (range 14–62) and corresponding mean age of the proband's mother 66 years (36–91). Most probands were Caucasian (30 of 31, one of whom was Hispanic Caucasian) with one woman for whom one parent was Asian and the other Caucasian and all were recruited from the Seattle area (Washington State). Twenty-two probands were parous women with an average number of children of 2.2 (mean age 12.6 years). Probands were selected from a large HLA family study database in which 80% of the probands were informative for maternal Mc. An additional requirement for these studies was that maternal HLA polymorphism was not shared with one of the proband's children. The Internal Review Board of the Fred Hutchinson Cancer Research Institute approved the study. All subjects provided informed consent.
For comparison purposes we also studied 27 healthy women for fetal Mc, as assessed by male DNA in women with sons, as some cellular subsets have not previously been studied for fetal Mc. For these studies, the mean age of the proband at the time of the blood draw was 43.8 years (range 26–57) and the mean age of the proband's children was 13.8 years (range 1–34).
HLA Genotyping
HLA genotyping was performed for the probands, their mothers and all the proband's children. HLA genotyping was included for all children of probands to assure that testing for a nonshared maternal HLA allele could not be confounded by a similar HLA allele from fetal Mc. HLA genotyping was conducted from DNA extracted from whole blood for probands and from mouthwash samples for family members, as previously described.15 All subjects were genotyped for HLA-B, DRB1, DRB3, DRB4, DRB5, DQA1 and DQB1 loci. DNA-based typing was conducted with sequence-specific oligonucleotide probe panels as previously described10 or alternatively DRB and DQB1 strip detection (Dynal RELI™ SSO, UK) was used for initial determination of DRB and DQB1 families followed by identification of specific alleles by sequencing (Applied Biosystems, Foster city, CA, USA). HLA-B typing was performed with HLA-B strip detection kits using sequence-specific oligonucleotide probe (Dynal RELI™ SSO, UK).
Fluorescence-Activated Cell Sorting for Isolation of Cellular Subsets
Fifty ml whole blood samples were drawn into acid citric dextrose solution A-vacutainer tubes and processed to isolate PBMC by Ficoll–Hypaque (Pharmacia Biotech AB, Uppsula, Sweden) with density gradient centrifugation 1.077 g/ml. PBMC were resuspended in phosphate-buffered saline (PBS)/1% fetal bovine serum. Staining was performed as previously described:16 10–15 × 106 cells were stained with anti-CD3-FITC and anti-CD56/16-PE or anti-CD14-FITC and anti-CD19-PE monoclonal antibodies (BD Biosciences, Mountain View, CA, USA). Cells were sorted on FACSVantage™ SE Cell Sorter (BD Biosciences, Mountain View, CA, USA). An aliquot of the sorted cells was collected and the percentage of sorted cells correctly gated was calculated to assess purity. The majority (91%) of PBMC subset populations were sorted to purity ≥97%, with most (59%) ≥99%. Sorted samples with purity below 95% were excluded from analysis. Purity check was not conducted for the CD3+CD56/16+ population because these cells are infrequent, the number of events collected was low, and this population is described as enriched for NK T cells. Cellular subsets were washed and then frozen as dry pellets for DNA extraction. T lymphocytes (CD3+), B lymphocytes (CD19+), monocyte/macrophages (CD14+), NK (CD56/CD16+) and NK T (CD3+CD56/CD16+) represented, respectively, 58±8, 8±3, 16±7, 10±4 and 2.4±1.7% of the PBMC sorted (median±standard deviation, all probands).
DNA Extraction for Q-PCR
DNA was extracted from dry pellets using a Wizard® Genomic DNA Purification Kit (Promega, Madison, WI, USA) according to the manufacturer's instructions and resuspended in 50 μl of water.
Detection and Quantification of Maternal Mc by HLA-Specific Real-Time Q-PCR. Quantification of Male Mc by Q-PCR for a Y-Chromosome-Specific Sequence
Twelve HLA-specific Q-PCR assays were employed to identify and quantify maternal Mc. The panel of Q-PCR assays targets sequences that are specific to DRB1*01, DRB1*15/16, DRB1*0301, DRB1*04, DRB1*11, DRB1*1001, DQA1*01, DQA1*03, DQB1*02, DQB1*0301/0304, DRB4*01 and HLA-B*44. Development and validation of the panel of HLA-specific Q-PCR assays was recently reported9 with the addition of four new assays. Briefly, real-time Q-PCR reactions were setup in a reaction volume of 50 μl. In all assays TaqMan Universal Master Mix was used as recommended by the manufacturer (Applied Biosystems, Foster City, CA, USA), with 300 nM of each amplification primer and 100 nM of the dual-labeled probe. The amplification conditions consisted of an initial incubation at 50°C for 2 min, followed by incubation at 95°C for 10 min, 45 cycles of 95°C for 15 s and 60°C for 1 min (except for DRB1*15/16 and DQB1*02, for which the annealing temperature was 62°C, and DRB1*04, DRB1*11, DQA1*01 and DQA1*03 for which the annealing temperature was 64°C). Amplification data were collected by an Applied Biosystems 7000 sequence detector and analyzed by the Sequence Detection System software (Applied Biosystems, Foster City, CA, USA).
Specificity was validated by testing each HLA-specific Q-PCR primer pairs and probe with an extended panel of 47 well-characterized HLA cell lines from the 13th International HLA Working Group representing all HLA–DRB1, DRB3, DRB4, DRB5, DQA1, DQB1 and B allele groups.9 Sensitivity was determined by testing dilutions of the equivalent DNA of cells homozygous for the HLA-specific polymorphism in a background of cells negative for that HLA polymorphism. All HLA-specific assays had similar sensitivity of one cell in a background of 20 000 negative cell equivalents (sensitivity 0.005%).
Between four and six aliquots of DNA from cellular subsets and PBMC were tested by HLA-specific Q-PCR. Two additional DNA aliquots were amplified concurrently for a housekeeping gene, β-globin, to define the total DNA concentration of the sample tested using a β-globin standard curve. For quantitative measurement of maternal DNA, two HLA-specific calibration curves were run on each plate using dilutions of equivalent DNA of 500, 100, 50, 10, 5, 1, 0.5 cells homozygous for the specific HLA polymorphism in a background of 20 000 cell equivalents negative.9 The conversion factor of 6.6 pg of DNA per cell was used.17 To assure the intra- and interassay homogeneity, HLA-specific standard curves and β-globin standard curve were run simultaneously on all plates. For all the standard curves, the correlation coefficient was 0.99. Samples were considered positive if at least one well gave an amplification curve crossing the threshold. The majority of the samples (67%) had more than two wells positive. The median total number of genome equivalents (gEq) tested for CD3, CD19, CD14, NK cells and PBMC was respectively: 88 705; 65 193; 99 147; 70 575 and 98 969 gEq. To be included in analysis it was required that any aliquot tested not exceed the DNA gEq of 30 000 cells (198 ng) because we have previously found that larger quantities of DNA can be inhibitory for the Q-PCR reaction.9 As some sorted subsets yielded small numbers we also included the requirement that at least 20 000 gEq be tested for inclusion in analysis. As each well is assumed to have a Poisson distribution of Mc, for each subject, the gEq number of microchimeric cells were averaged combining information across aliquots of DNA as previously described.15 Results were reported as the DNA gEq number of maternal cells per 100 000 proband cells (gEq/100 000).
Additional studies evaluated fetal Mc, as measured by male Mc in women who had given birth to sons, employing Q-PCR for the Y-chromosome-specific sequence DYS14 as previously reported.15 The DYS14 and all HLA-specific Q-PCR assays were developed to have the same sensitivity as described above.
Anticontamination Procedures
Rigorous precautions against PCR contamination were taken. Aerosol-resistant pipette tips and clean gloves were always used. Isolation of DNA and setup of amplification reactions were accomplished in separate locations. The use of dUTP instead of dTTP and the addition of uracil-N-glycosylase in TaqMan Universal Master Mix were used for prevention of carryover PCR product. Negative controls were included in each Q-PCR plate with no DNA and with background DNA from an individual not carrying the HLA polymorphism tested. Negative controls were consistently negative across all the experimental plates.
Statistical Analysis
Comparisons in prevalence of maternal vs fetal Mc were made using logistic regression models, with adjustment for the total number of cells collected. Reported P-values are two-sided, and based on the Wald statistic.
Results
Characteristics of the 31 healthy women probands studied for maternal Mc are provided in Table 1. After HLA genotyping all study subjects, one of a panel of twelve different HLA-specific Q-PCR assays was employed to test DNA extracted from cellular subsets for maternal Mc. Maternal Mc was quantified by targeting a nontransmitted, nonshared HLA allele of the proband's mother and if the proband was a parous woman, also not shared by any of her children. Among T lymphocytes, maternal Mc was found in 25% of probands (7/28) (Table 2). Maternal Mc levels ranged from 0.9 to 25.6 gEq/100 000. Among B lymphocytes, 14% of probands had maternal Mc (3/21) (Table 2) with similar levels as observed in T lymphocytes, ranging from 0.9 to 25.3 gEq/100 000. In monocyte/macrophages, 16% of the probands had maternal Mc (4/25) and in NK cells 28% (5/18). Maternal Mc levels in monocyte/macrophages were 0.3–36 gEq/100 000 and in NK cells, 1.8–3.4 gEq/100 000. Thus overall, the prevalence of maternal Mc was somewhat greater in T lymphocytes and NK cells compared to monocyte/macrophages and B lymphocytes while levels were generally similar with the possible exception of NK cells. Overall, more than one-third of probands had maternal Mc in at least one cellular subset (39%, 12/31). In limited studies of NK T cells, a rare population within PBMC so that fewer sorted cells were available for testing (see Materials and methods), two of nine probands (22%) had maternal Mc (data not shown).
Samples from 14 probands yielded sorted cell numbers that permitted testing in all of the four populations, CD3, CD19, CD14 and NK from the same blood sample; six (43%) were positive for at least one subset (two in three of four subsets). Maternal Mc was also assayed in DNA from unselected PBMC from the same blood draw used for FACS sorting. Seventeen percent (5/30) of probands had maternal Mc in PBMC (Table 2). Most probands with maternal Mc in at least one subset did not have detectable maternal Mc in PBMC (8/12) and of those with positive results in both unselected PBMC and cellular subsets had lower maternal Mc levels in PBMC than in cellular subsets (3/4) (Table 2). Both of these observations are consistent with relative enrichment of maternal Mc in cellular subsets after FACS when compared to total PBMC.
The presence of maternal Mc was not correlated with any particular HLA allele of the proband or of her mother nor was there any suggestion that results differed according to the specific Q-PCR assay used (all Q-PCR assays have similar sensitivities; see Materials and methods). There was the suggestion that maternal Mc may be more prevalent among parous than nulliparous women, as maternal Mc was found in at least one subset in 10 of 22 (45%) compared to two of nine (22%), respectively (for those with all four subsets tested five of 10 (50%) vs one of four (25%) had maternal Mc, respectively).
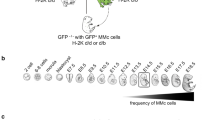
For comparison purposes similar cellular subsets as studied for maternal Mc were studied for fetal Mc, as assessed by male DNA in women who had at least one son. Fetal Mc was quantified among T lymphocytes, B lymphocytes, monocyte-macrophages and NK cells for 27 women using previously described methods.15 Fetal Mc was more prevalent than maternal Mc in all subsets (Figure 1). Within T lymphocytes, 58% (14/24) had fetal Mc compared to the 25% observed for maternal Mc (P=0.01). Among B lymphocytes, 75% (6/8) had fetal Mc compared to 14% for maternal Mc (P=0.005). Among monocyte/macrophages, fetal Mc was found in 50% (6/12) of women compared to 16% for maternal Mc (P=0.04) and in NK cells, 61.5% (8/13) had fetal Mc compared to 28% with maternal Mc (P=0.05). Overall 78% (21/27) of women had fetal Mc in at least one subset compared to 39% (12/31) with maternal Mc (P=0.002). Fetal Mc levels ranged, respectively, from 0.03 to 2.7; 0.09 to 4.1; 0.01 to 23.8 and 0.06 to 1076 gEq/100 000 for T lymphocytes, B lymphocytes, monocyte/macrophages and NK cells. Among women who had three or four subsets tested for fetal Mc all had at least one positive result and most had positive results in more than one subset (6/10). Thus, when the prevalence of maternal Mc was compared to that of fetal Mc as assessed by male DNA in women with sons, maternal Mc was substantially less prevalent than fetal Mc both overall and in all the cellular subsets.
Prevalence of maternal Mc compared to fetal Mc, as measured by male Mc in women with sons, in cellular subsets in healthy adult women. A total of 31 women were tested for maternal Mc and 27 women with sons were tested for fetal Mc in cellular subsets by Q-PCR targeting maternal-specific HLA sequences or the male DNA sequence DYS14, respectively (see Materials and methods). Seven parous women were tested for both maternal and fetal Mc on two different blood draws, two of whom were positive for both in the same cellular subset, four negative for both and one woman had a positive result in one subset for fetal Mc but negative for maternal Mc. (1) P=0.01, (2) P=0.005, (3) P=0.04, (4) P=0.05. *Results previously reported.15
Discussion
Maternal Mc has recently been described in the fetal circulation in the third and early second trimester.18, 19 We previously found that maternal Mc persists into adult life in immune competent adults in the circulation of healthy subjects in addition to patients with systemic sclerosis.8, 9 In the current study we asked whether healthy adult women harbor maternal Mc among their T and B lymphocytes, monocyte/macrophages and NK cells, and if so at what levels.
We found maternal Mc in each of the cellular subsets tested in some study subjects. Given the detection of maternal Mc in circulating peripheral blood cells with widely variable half-lives from hours to months, healthy women likely harbor long-surviving populations of maternal microchimeric hematopoietic stem cells. Maternal Mc levels generally ranged from 1 to 30 gEq/100 000 proband cells. Considering T lymphocytes as an example, the total number of circulating maternal T lymphocytes may be estimated by considering 5 l of blood with 4–10 billion white blood cells per liter,20 half of which are PBMC; therefore with 59% of PBMC being T lymphocytes (see Materials and methods), 1–26 gEq/100 000 would correspond to 100 000–2 600 000 circulating maternal T lymphocytes. Thus, the quantitative assessment of maternal Mc in cellular subsets indicates maternal Mc is present at levels with the potential for immunological effects. Peptide-specific CD4 T cells, for example, have been reported at 3–5/100 00021 and virus-specific CD8 T cells at 0.03 to 2% of CD8 T cells.22, 23
We recently identified female cells in the heart tissues of male infants with neonatal lupus who died from congenital heart block that lacked hematopoietic markers instead expressing the phenotype of cardiac myocytes.24 In studies by others, female cells (presumed maternal) have been identified in multiple tissues of newborn males including the liver, spleen, thymus, thyroid and skin.25 Our current results when considered along with other observations suggest maternal Mc is a constitutive component of normal biology, represented within circulating cellular subsets and contributing to some tissues. This consideration raises interesting questions such as whether maternal Mc plays a role in development in her progeny26 and whether an autoimmune disease may sometimes result from breakdown in tolerance to maternal Mc. Maternal Mc could also be recruited to sites of inflammation and secondarily function in tissue repair or maternal effector cells might sometimes gain reactivity to autologous tissues.
While the explanation for maternal Mc being in cellular subsets in some women but not others is not known, that maternal Mc persists in adult women leads to the question of whether there are effects of maternal Mc when the daughter becomes pregnant, thus acquiring a second source of Mc from the fetus. Although we cannot address this question as part of the current studies, we did note a tendency for women who were parous to be positive for maternal Mc more often than nulliparous women, and the number of prior births was higher in women with vs those without maternal Mc (Table 3).
When compared to similar studies of fetal Mc, as assessed by male DNA in women with sons, maternal Mc was less prevalent in each cellular subset. Although the reasons for this difference are not known it bears resemblance to maternal-fetal trafficking during pregnancy where greater fetus to mother than mother to fetus trafficking has been reported1 and it is possible that levels of Mc are tightly regulated thereafter. Although the mechanisms by which maternal and fetal Mc persist in the face of HLA-disparity are unknown it is reasonable to anticipate that mechanisms permitting maternal and fetal Mc persistence would differ because the former is acquired during and the latter after immune system development.
It is important to emphasize that the current studies provide an estimate of maternal Mc in cellular subsets in proband–mother pairs in which there is at least one HLA-disparity, the usual situation in outbred populations. In a recent report of a murine model, differences in maternal Mc levels were sometimes observed in tissues and in the circulation in association with major histocompatibility gene sharing.27 The interesting issue of whether maternal Mc in cellular subsets differs in situations where the mother is fully HLA-identical or HLA-homozygous could be addressed in future studies using a similar approach as the current studies but developing a panel of Q-PCR assays that target nonallelic polymorphisms on other chromosomes. A recent report using Q-PCR assays targeting genetic polymorphism other than HLA suggested an association between maternal Mc and maternal histocompatibility at class II loci from the perspective of the fetus.28
There are a number of limitations to our study. The approach that we used to detect and quantify maternal Mc is labor intensive because HLA genotyping in complete family studies must first be performed, and the number of study subjects is modest. The effect of age, gender and pregnancy history in women is not known. Among parous women in our study there was no apparent difference in maternal Mc according to age; women who were positive in at least one subset had a mean age of 43.7 years and women negative a mean age of 44 years (Table 3). However, nulliparous women who were younger than parous women were also mostly negative for maternal Mc indicating additional studies are needed. A potential source of confounding in testing for Mc is a blood transfusion.29 This is unlikely to impact our findings as there was no history of blood transfusion for 28 of our study subjects; for two history was unknown and although one subject reported a transfusion and had maternal Mc in NK cells, it is not likely that the transfusion donor shared the maternal HLA. Another limitation is that measuring male DNA in women who previously gave birth to a son is an imperfect measure of fetal Mc as some women without sons may have male DNA in peripheral blood.30 Male DNA may potentially derive from early pregnancy loss, a vanished male twin or other sources. While it is possible that some male DNA originated from sources other than women's sons, whereas it might diminish differences, it could not explain the marked difference in maternal and fetal Mc prevalence observed in the cellular subsets. Finally, caution is warranted in interpretation as Q-PCR results could be both under and over-representative, the former due to testing relatively small numbers of cell equivalents and the latter because subsets were sorted to high but not 100% purity and because of the potential for PCR contamination despite rigorous preventive measures.
In conclusion, maternal Mc was found in more than one-third of healthy women within lymphoid and myeloid lineages. Maternal Mc was found among cell populations that have immunologic activity functioning in innate and/or adaptive immune responses. To our knowledge, the current study is the first to specifically address maternal Mc in cellular subsets in healthy adults. These results indicate that further studies are needed to explore whether there are biological effects of maternal Mc in cellular subsets, and to determine whether beneficial and/or detrimental effects sometimes result for her progeny.
Supplementary information
Supplementary information providing a representative HLA-specific Q-PCR assay is available on the Laboratory Investigation's website.
References
Lo YM, Lau TK, Chan LY, et al. Quantitative analysis of the bidirectional fetomaternal transfer of nucleated cells and plasma DNA. Clin Chem 2000;46:1301–1309.
Herzenberg LA, Bianchi DW, Schroder J, et al. Fetal cells in the blood of pregnant women: detection and enrichment by fluorescence-activated cell sorting. Proc Natl Acad Sci USA 1979;76:1453–1455.
Socie G, Gluckman E, Carosella E, et al. Search for maternal cells in human umbilical cord blood by polymerase chain reaction amplification of two minisatellite sequences. Blood 1994;83:340–344.
Hall JM, Lingenfelter P, Adams SL, et al. Detection of maternal cells in human umbilical cord blood using fluorescence in situ hybridization. Blood 1995;86:2829–2832.
Lo YM, Lo ES, Watson N, et al. Two-way cell traffic between mother and fetus: biologic and clinical implications. Blood 1996;88:4390–4395.
Wernet P, Kogler G, Somville T . The rapid detection of the quantity (genotype) and cell lineage (immunophenotype) of contaminating maternal white cells in cord blood samples by fluorescence in situ hybridization combined with confocal laser scanning microscopy. Blood Cells 1994;20:296–302.
Bianchi DW, Zickwolf GK, Weil GJ, et al. Male fetal progenitor cells persist in maternal blood for as long as 27 years postpartum. Proc Natl Acad Sci USA 1996;93:705–708.
Maloney S, Smith A, Furst DE, et al. Microchimerism of maternal origin persists into adult life. J Clin Invest 1999;104:41–47.
Lambert NC, Erickson TD, Yan Z, et al. Quantification of maternal microchimerism by HLA-specific real-time polymerase chain reaction: studies of healthy women and women with scleroderma. Arthritis Rheum 2004;50:906–914.
Nelson JL, Furst DE, Maloney S, et al. Microchimerism and HLA-compatible relationships of pregnancy in scleroderma. Lancet 1998;351:559–562.
Reed AM, Picornell YJ, Harwood A, et al. Chimerism in children with juvenile dermatomyositis. Lancet 2000;356:2156–2157.
Artlett CM, Ramos R, Jiminez SA, et al. Chimeric cells of maternal origin in juvenile idiopathic inflammatory myopathies. Childhood Myositis Heterogeneity Collaborative Group. Lancet 2000;356:2155–2156.
Reed AM, McNallan K, Wettstein P, et al. Does HLA-dependent chimerism underlie the pathogenesis of juvenile dermatomyositis? J Immunol 2004;172:5041–5046.
Rose NR, Bona C . Defining criteria for autoimmune diseases (Witebsky's postulates revisited). Immunol Today 1993;14:426–430.
Lambert NC, Lo YM, Erickson TD, et al. Male microchimerism in healthy women and women with scleroderma: cells or circulating DNA? A quantitative answer. Blood 2002;100:2845–2851.
Evans PC, Lambert N, Maloney S, et al. Long-term fetal microchimerism in peripheral blood mononuclear cell subsets in healthy women and women with scleroderma. Blood 1999;93:2033–2037.
Saiki RK, Gelfand DH, Stoffel S, et al. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 1988;239:487–491.
Petit T, Dommergues M, Socie G, et al. Detection of maternal cells in human fetal blood during the third trimester of pregnancy using allele-specific PCR amplification. Br J Haematol 1997;98:767–771.
Lo ES, Lo YM, Hjelm NM, et al. Transfer of nucleated maternal cells into fetal circulation during the second trimester of pregnancy. Br J Haematol 1998;100:605–606.
CDC Desk Reference for Hematology. Reference Range Tables. In: Shinton NK (ed). Table 1: Reference Ranges of Hematological Values for Adults. CRC Press LLC: Boca Raton, London, New York, Washington, DC, 1998, p 710.
Novak EJ, Liu AW, Nepom GT, et al. MHC class II tetramers identify peptide-specific human CD4(+) T cells proliferating in response to influenza A antigen. J Clin Invest 1999;104:R63–R67.
He XS, Mahmood K, Maecker HT, et al. Analysis of the frequencies and of the memory T cell phenotypes of human CD8+ T cells specific for influenza A viruses. J Infect Dis 2003;187:1075–1084.
Kern F, Bunde T, Faulhaber N, et al. Cytomegalovirus (CMV) phosphoprotein 65 makes a large contribution to shaping the T cell repertoire in CMV-exposed individuals. J Infect Dis 2002;185:1709–1716.
Stevens AM, Hermes HM, Rutledge JC, et al. Myocardial-tissue-specific phenotype of maternal microchimerism in neonatal lupus congenital heart block. Lancet 2003;362:1617–1623.
Srivatsa B, Srivatsa S, Johnson KL, et al. Maternal cell microchimerism in newborn tissues. J Pediatr 2003;142:31–35.
Rinkevich B . Human natural chimerism: an acquired character or a vestige of evolution? Human Immunol 2001;62:651–657.
Kaplan J, Land S . Influence of maternal-fetal histocompatibility and MHC zygosity on maternal microchimerism. J Immunol 2005;174:7123–7128.
Berry SM, Hassan SS, Russell E, et al. Association of maternal histocompatibility at class II HLA loci with maternal microchimerism in the fetus. Pediatr Res 2004;56:73–78.
Lee TH, Paglieroni T, Ohto H, et al. Survival of donor leukocyte subpopulations in immunocompetent transfusion recipients: frequent long-term microchimerism in severe trauma patients. Blood 1999;93:3127–3139.
Yan Z, Lambert NC, Guthrie KA, et al. Male microchimerism in women with no prior birth of a son: quantitative assessment and correlation with pregnancy history. Am J Med 2005;118:899–906.
Acknowledgements
We thank Shelly Heimfeld, PhD for his useful comments and critique. We thank Allison J Porter, Jöelle Lucas and Jeff Redinger for their valuable technical contribution and Jennifer Brackensick for study subject recruitment. This work was supported by NIH Grants AI 45659 and AI 41721.
Author information
Authors and Affiliations
Corresponding author
Additional information
Duality of interest
None declared.
Supplementary Information accompanies the paper on the Laboratory Investigation website (http://www.nature.com/labinvest)
Supplementary information
Rights and permissions
About this article
Cite this article
Loubière, L., Lambert, N., Flinn, L. et al. Maternal microchimerism in healthy adults in lymphocytes, monocyte/macrophages and NK cells. Lab Invest 86, 1185–1192 (2006). https://doi.org/10.1038/labinvest.3700471
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700471
Keywords
This article is cited by
-
The ‘communicatome’ of pregnancy: spotlight on cellular and extravesicular chimerism
EMBO Molecular Medicine (2024)
-
Cord blood maternal microchimerism following unrelated cord blood transplantation
Bone Marrow Transplantation (2021)
-
Immune dysfunction in developmental programming of type 2 diabetes mellitus
Nature Reviews Endocrinology (2021)
-
Breastfeeding-related maternal microchimerism
Nature Reviews Immunology (2017)
-
The maternal microbiome during pregnancy and allergic disease in the offspring
Seminars in Immunopathology (2017)