Abstract
Cirrhosis is a heterogeneous tissue composed of polyclonal regenerative and monoclonal neoplastic, potentially malignant nodules from which hepatocellular carcinoma (HCC) might develop. The aim of this study was to investigate proteomic profile changes associated with clonal expansion of cirrhotic nodules and malignant transformation of monoclonal nodules. Seventy-one cirrhotic nodules from 10 female patients with six HCC were dissected from liver surgical specimen by laser capture microdissection. Clonal status of each nodule was assessed by the study of X-chromosome inactivation pattern using the human androgen receptor. Protein profiles were determined by surface-enhanced laser desorption ionisation-time-of-flight technology using Q10 arrays (Cyphergen ProteinChip®). Molecular weight of differentially expressed protein peaks was assessed. An average of 50 protein peaks was obtained for each nodule's profile. Comparison of protein profiles in polyclonal (n=45) and monoclonal cirrhotic nodules (n=26) identified three differentially expressed protein peaks (10 092, 54 025 and 62 133 Da). All were upregulated in monoclonal nodules. Twelve peaks were differentially expressed between monoclonal nodules and HCC with nine proteins upregulated in cancer samples. This study confirms that proteome analysis can be achieved from a limited number of microdissected cells, and provides further insight into the process of clonal expansion and malignant transformation of cirrhotic nodules.
Similar content being viewed by others
Main
Hepatocellular carcinoma (HCC) is a major public health problem worldwide, with an estimated 564 000 new cases per year. It nearly always develops in the setting of chronic hepatitis or cirrhosis related to various origins, including hepatitis virus infection, high alcohol intake or metabolic diseases.1 Until now, diagnosis of HCC has been made too late. Indeed, at the time of discovery, prognosis is poor, with a 5-year survival rate of less than 5%.2 Detection of smaller tumors and even preneoplastic lesions is therefore becoming a topic of major importance, in order to improve prognosis and therapeutic possibilities. In this context, identification of early molecular steps in HCC development might be useful in characterizing efficient, clinically useful biomarkers in cirrhotic patients.
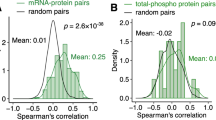
Molecular steps in liver carcinogenesis have been widely investigated. Most studies have focused on full-blown HCC, showing deregulation of several major pathways, such as p53 and β-catenin mutations.3 In contrast, only a limited number of studies have paid attention to the earliest stages of HCC development in cirrhotic tissue. It has been suggested that HCC might develop via slow progression from regenerative to dysplastic to malignant macronodules. This hypothesis is based mainly on morphological changes. However, up until now, the hypothesis of slow progressive transformation has been lacking in functional characterization. Interestingly, based on molecular pathology approaches, several authors have conversely shown that cirrhotic liver is a heterogeneous tissue composed of a mixture of polyclonal regenerative and monoclonal neoplastic, potentially malignant nodules without any significant difference based on pathological criteria.4, 5, 6, 7, 8, 9 It is also presumed that, whereas most monoclonal nodules will not progress to carcinoma, a limited number of them, under the influence of additional oncogenic events, will become malignant. This finding is of major importance, since clonal expansion of cirrhotic nodules might represent very early steps in carcinogenesis, followed by malignant transformation of monoclonal nodules. By studying gene expression, we recently characterized a set of genes, the expression level of which is deregulated during clonal expansion.10 These data provide clues to the theory of clonal expansion, but it is also known that, especially in the liver, mRNA and protein levels do not strictly correlate.11 For this reason, characterization of protein profiles associated with clonal expansion and malignant transformation of cirrhotic nodules in HCC is of particular interest.
Surface-enhanced laser desorption ionization time-of-flight mass spectrometry (SELDI-TOF-MS®, Ciphergen Biosystems) has recently proven useful in discovering biomarkers for diagnosis of various cancers in biological fluids and tissues.12, 13, 14, 15, 16 Using this technical approach, proteins are selectively retained on chromatographic surfaces, and after desorption, they are detected and analyzed by mass spectrometry. This easy-to-use technology has numerous advantages. It offers an adequate sensitivity and high resolution at low mass ranges. Furthermore, it allows easy differential analysis by comparison of raw data in groups of different phenotypes. Using this technology, we recently characterized a new biomarker of HCC in the serum of cirrhotic patients.17 Finally, the small amounts of material required for protein analysis enables the application of the SELDI-TOF ProteinChip technique with laser capture microdissection (LCM) technologies, to enriched and selected cell populations.
The aim of the present study was to determine whether comprehensive proteomic profiling of microdissected liver cirrhotic nodules, coupled with the ProteinChip system, could identify proteomic signatures associated with clonal expansion and malignant transformation of monoclonal cirrhotic nodules into HCC.
Materials and methods
Human Tissue Samples
Frozen tissue blocks from 10 female patients were retrieved from the files of the Department of Pathology of Beaujon Hospital between 2002 and 2004. All patients had cirrhotic livers, which were explanted or resected for an associated HCC. The mean age of the patients was 55 years (range 38–70 years). Cirrhosis was related to alcohol abuse (three cases), chronic hepatitis C infection (five cases) and chronic hepatitis B infection (two cases). Seven of these cases were associated with HCC. The mean size of the HCC was 4.2±2.2 cm (range 3–7.5 cm). According to Edmonson's histological grade, two HCC were grade 3, three were grade 2, one was grade 1 and one was unavailable for evaluation because of total necrosis. Patient characteristics are shown in Table 1.
In each case, liver tissue blocks from cirrhosis and HCC were immediately frozen in nitrogen and stored at −80°C until use for molecular studies. Samples were taken on cirrhotic tissue at distance from HCC or from any suspicious macronodule at gross examination. The remainder of the liver was routinely fixed and paraffin-embedded for routine histopathologic examination.
LCM of Cirrhotic Nodules
We used LCM to limit the heterogeneity of cirrhotic tissue and HCC and to separately study cirrhotic nodules. Four serial sections 7 μm thick were cut from each of the frozen blocks of cirrhotic livers. The first section was stained with hematoxylin and eosin for histological analysis and cirrhotic nodule landmarks and selection. Histologic analysis included assessment of main pathologic features of cirrhotic nodule: size (in mm), inflammation, iron deposit, steatosis, large liver cell change, small liver cell change and cholestasis. Each histologic feature was semiquantitatively assessed (presence or absence) independently for each nodule prior to microdissection.
The next three serial sections were immediately dehydrated for 30 min in 100% ethanol at −20°C and for 20 min in xylene at room temperature. Each selected nodule or HCC was microdissected individually by LCM with a Pixcell II instrument (Acturus Engineering, Mountain View, CA, USA). The parameters used included a laser diameter of 30 μm, laser power of 80 μm and a pulse length of 20 ms. The microdissected material obtained separately from each cirrhotic nodule or HCC was used for DNA extraction and clonal analysis (2nd section), and for protein extraction and SEDI-TOF analysis (3rd and 4th sections).
Clonal Analysis of Microdissected Cirrhotic Nodules by the Human Androgen Receptor Gene Polymorphic Marker Assay (HUMARA)
Assessment of clonality
From each microdissected cirrhotic nodule, DNA was extracted using a standard procedure. The clonal status of each cirrhotic nodule was then analyzed by the pattern of X-chromosome methylation using the HUMARA. Half of the extracted DNA was digested in the presence of 10 U of HpaII and 10 U of RsaI restriction enzymes (Roche Diagnostics). The other half was digested only by RsaI under the same conditions. Both HpaII-digested and -undigested DNAs were subjected to PCR amplification with primers and conditions according to those published previously.10 After a short denaturation, PCR products were electrophoresed through a denaturing gel run on a Abiprism 377® DNA sequencer (Applied Biosystems). The relative amounts of PCR products (area under the curve) were calculated for each allele using Abiprism Genescan™ 2.0.2 software (Applied Biosystems).
Data interpretation
A corrected ratio (CR) was first assessed by dividing the ratio (allele 1/allele 2) of the digested sample obtained after digesting DNA with HpaII by the ratio (allele 1/allele 2) of the nondigested sample. A final ratio for each cirrhotic nodule was determined by dividing the CR obtained from nodule DNA by the CR of the adjacent cirrhotic tissue DNA. This final clonal ratio (clonal index) corrects lionization phenomena. To estimate the sensitivity of the technique, a titration curve was performed by mixing variable amounts of monoclonal cells in a polyclonal background and assessing the clonal index according to the percentage of monoclonal cells in the mix. A cirrhotic nodule was considered to be monoclonal when it consisted of at least 25% of monoclonal cells against a polyclonal background. According to the titration curve, this percentage corresponded to a final ratio of 1.5, and any nodule with a final ratio above 1.5 was considered to be monoclonal.
Proteomic Analysis of Cirrhotic Nodules and HCC
Lysis and proteinchip array protocol
A total of 1200–1800 liver cells for each microdissected cirrhotic nodule or HCC (approximately 800–1200 LCM laser transfer pulses) were transferred to an LCM cap. To each LCM cap, 9 μl of lysis buffer was added (7 M urea, 2 M thiourea, 1% dithioerythritol, protease inhibitors) and incubated for 20 min at room temperature in a humidity chamber. Each sample was transferred to a 500 μl tube and frozen immediately at −80°C for a maximum of one day. The protein lysates were analyzed on a strong anionic exchanger array (Q10; Ciphergen Biosystems, Fremont, CA). This ProteinChip array was selected because, in a preliminary study of selected cases and in comparison with other array surfaces, Q10 ProteinChip gave the protein profile with the highest number of peaks of the highest intensity.
ProteinChip Q10 arrays were assembled into a deep-well-type bioprocessor assembly (Ciphergen Biosystems) and samples were run according to the manufacturer's instructions. Briefly, prior to sample loading, Q10 arrays were equilibrated with 200 μl binding/washing buffer (50 mM Tris-HCl, pH 8.4, 0.05% Triton X-100) for 5 min at room temperature. Each spot was then incubated with 100 μl of  diluted sample with the binding/washing buffer for 30 min on a shaker. Then, arrays were washed two times with 200 μl of binding buffer and one time with 200 μl of binding buffer without Triton. After rinsing with 200 μl of HEPES buffer 1 mM, the arrays were removed from the bioprocessor assembly and air-dried. In each spot of the arrays, 2 × 0.5 μl of saturated sinapinic acid with acetonitrile 50% and trifluoroacetic acid 0.5% was added.
diluted sample with the binding/washing buffer for 30 min on a shaker. Then, arrays were washed two times with 200 μl of binding buffer and one time with 200 μl of binding buffer without Triton. After rinsing with 200 μl of HEPES buffer 1 mM, the arrays were removed from the bioprocessor assembly and air-dried. In each spot of the arrays, 2 × 0.5 μl of saturated sinapinic acid with acetonitrile 50% and trifluoroacetic acid 0.5% was added.
ProteinChip array analysis
The ProteinChip Q10 arrays were analyzed in the ProteinChip biology SELDI-TOF system reader (model PBS II, Ciphergen Biosystems). An external calibration with ‘All-in-One proteins standard II’ (Ciphergen Biosystems) was performed before each measurement. Each spot was analyzed twice, with separate optimization for the low (3–30 kDa) and high (30–70 kDa) mass ranges using, respectively, low (190) and high (225) laser intensities. Protein profiles for each microdissected cirrhotic nodule or HCC where each protein was characterized by a peak with a specific mass/charge ratio (m/z) and intensity were collected.
Finally, all mass spectra generated for each cirrhotic nodule and HCC were normalized together to total ion current and analyzed using ProteinChip software version 3.0 (Ciphergen Biosystems).
Data analysis
SELDI mass spectra were compared between polyclonal and monoclonal cirrhotic nodules, and between monoclonal nodules and HCC using the Biomarker Wizard program (Ciphergen Biosystem). Clonal status and protein profiles were also compared according to histopathologic features. Raw data were also analyzed with NCSS 2004 software (Statistical Systems, Kayville, UT) after exportation to a standard Excel file format. Quantitative values in different phenotype groups were compared using the nonparametric Mann–Whitney test. A P-value of less than 0.05 was considered as significant.
Results
Histological Analysis of Microdissected Nodules
Before extraction, an H&E-stained slide was performed on each frozen tissue block to assess main relevant histological feature of cirrhotic micronodules. Prevalence of histopathologic features and their relationship with clonal status of micronodules are reported in Table 2.
Clonal Analysis of Microdissected Cirrhotic Nodules
Each of the 10 female patients included in the study were heterozygous for the gene polymorphic sequence of HUMARA providing information in clonal status analysis. A total of 117 nodules and six HCC were microdissected with five to 14 cirrhotic nodules per case.
DNA from the cirrhotic nodules and adjacent liver was successfully extracted in all cases. Among the 117 nodules studied, the clonality index was obtained for 78 nodules. Twenty-seven had a final ratio equal to or greater than 1.5, indicating a monoclonal pattern, and 51 had a final ratio of less than 1.5, indicating a polyclonal pattern according to the titration curve. The mean size of monoclonal nodules was 3.1±1.4 mm, not statistically different from polyclonal nodules (2.7±1.3 mm).
ProteinChip Analysis
Since clonal status was known for 78 cirrhotic nodules, protein profiles from the 39 nodules with unknown clonal status were excluded from the study. The 78 protein profiles from poly- and monoclonal cirrhotic nodules and from microdissected HCC were listed in a global file. After normalization, six polyclonal and one monoclonal cirrhotic nodules were also excluded because the normalization coefficient fell out of the acceptable range. Finally, 45 polyclonal nodules, 26 monoclonal nodules and six HCC were used for ProteinChip analysis.
For each cirrhotic nodule or HCC, the protein profile comprised between 30 and 50 different protein peaks.
Comparison of protein profiles of polyclonal and monoclonal cirrhotic nodules
Comparison of protein mass spectra in polyclonal (n=45) and monoclonal (n=26) cirrhotic nodules yielded three protein peaks that were significantly differentially expressed (P<0.05). Indeed, all were overexpressed in monoclonal vs polyclonal nodules. These peaks had an m/z ratio of 10 092, 54 025 and 62 133 Da, respectively. A comparison of protein profiles between a monoclonal and polyclonal cirrhotic nodule is shown in Figure 1. These results were not significantly different whether viral or alcohol cirrhosis was considered. Results are summarized in Table 3. Interestingly, when the quantitative value of the clonal index was correlated with the expression level of the three markers of clonality (Spearman's correlation test), they all displayed significant correlation (P<0.01). The correlation between the clonal index and the 54 025 Da peak (r2=0.53, P<0.01) is shown in Figure 2.
Comparison of protein profiles of monoclonal cirrhotic nodules and HCC
Protein mass spectra comparison of monoclonal cirrhotic nodules (n=26) and microdissected HCC (n=6) showed 12 protein peaks that were significantly and differentially expressed (P<0.05), and ranged between 3.3 and 70.7 kDa. Except for one (m/z=54025), none of them appeared deregulated when comparing poly- and monoclonal nodules. Among these 12 protein peaks, nine were higher in HCC compared to monoclonal nodules. A comparison of the protein profile between a monoclonal cirrhotic nodule and HCC is shown in Figure 3. Figure 4 shows the mean value of protein peaks that differentiated mono- vs polyclonal nodules and monoclonal cirrhotic nodules vs HCC. There was no correlation between the different selected peaks when intensity was compared.
Discussion
Cirrhosis is a premalignant condition, since most HCC develop from cirrhosis. It has been clearly shown that cirrhotic tissue is composed of a mixture of poly- and monoclonal nodules.4, 5, 6, 7, 8, 9 One of the earliest molecular mechanisms that drive cirrhosis to HCC is clonal expansion of cirrhotic nodules. Several technical approaches have been used to assess clonality at the tissular level. Analysis of integration site of HBV genome is an option, but is limited to those cases of HBV-related diseases. The HUMARA assay, which is based on the analysis of the pattern of X-chromosome methylation, using the polymorphic human androgen receptor gene limits this approach to female patients but can be used in every etiology. Furthermore, it has been used successfully in studies related to liver pathology.6, 7, 18, 19, 20
It is believed that HCC arises from the malignant transformation of a limited number of monoclonal cirrhotic nodules related to additional oncogenic events. Therefore, insight into early molecular mechanisms of HCC can only be obtained from analysis of the two earliest steps in liver carcinogenesis: clonal expansion of polyclonal nodules and malignant transformation of clonal nodules. The aim of the present study was to explore protein expression associated with these two initial events, by combining differential proteomic analysis and DNA clonality assessment of microdissected cirrhotic nodules. This objective is in line with a previous work showing that a set of genes is differentially expressed when polyclonal and monoclonal nodules are compared using a real-time RT-PCR approach.10
Proteomics has now gained wide acceptance for studying the diagnosis and prognosis of human malignancy.21, 22, 23 Thus, such approach has already provided additional insight into the pathogenesis of HCC, with the identification of new tumor-associated proteins in fully developed HCC. Indeed, members of the aldo-keto reductase (AKR) family, tissue ferritin light chain and aldehyde dehydrogenase have recently been proposed as valuable diagnostic markers of HCC.24, 25, 26, 27 All of these candidates were isolated by two-dimensional (2D) polyacrylamide gel electrophoresis and identified by mass spectrometry. Although they are very accurate, it is noteworthy that such techniques are time-consuming, requiring substantial technical expertise and significant amounts of proteins. The recent development of proteomic array technology, including protein profiling, which couples the ProteinChip array with laser desorption ionization time-of-flight mass spectrometry, provides a potentially powerful tool for global visualization of proteome. Comparison of profiles according to phenotypes of samples enables the identification of peaks characteristic of the condition under study.28, 29, 30, 31 Using such technology, by separately comparing protein profiles obtained from polyclonal and monoclonal cirrhotic nodules, and HCC samples, we were able to point out three and 12 protein peaks that were differentially and sequentially expressed during clonal expansion and transformation of monoclonal nodules into HCC, respectively. These results show that emergence of HCC from cirrhosis is not a continuous, progressive molecular process, but rather a stepwise process associated with different molecular events.
It is interesting to note that none of the histopathological features of cirrhotic micronodules correlated either with their clonal status or with significant changes in proteomic profiles. These data suggest that the earliest molecular events in liver carcinogenesis have no morphologic counterpart and that clonal expansion is not related to phenotypic changes.
To compare accurately monoclonal and polyclonal nodules, we used LCM microdissection, which enables accurate selection of cell types of interest. Indeed, this procedure overcomes the problem of tissue heterogeneity. Cell selection is of crucial importance in the context of liver carcinogenesis, since various cell types (hepatocytes, mesenchymal, endothelial and inflammatory cells) are intimately mixed in cirrhotic tissue as well as in HCC. Although the capture of a pure cell population such as hepatocytes cannot be totally achieved because of the close association of liver cells with sinusoidal cells, LCM microdissection dramatically increases liver cell purity by excluding fibrous septa. This approach undoubtedly contributes to the enrichment of biological samples resulting in adequate and representative material. A combination of proteomic analysis with LCM has already been achieved in several studies. Using LCM, Li et al31 were able to identify at least 600 proteins from 50 000 to 100 000 cells of HCC and non-HCC by isotope-coded affinity tag technology and 2D liquid chromatography. Nevertheless, the picking up of so many cells under adequate conditions is time-consuming. This technical shortcoming can be overcome by the use of SELDI-TOF technology. Indeed, due to the high sensitivity of this technique, we evaluated that a mean of 1000 microdissected cells was sufficient to display reproducible and informative protein spectra.
In this study, we have assessed specific protein profiles associated with the multistep process of hepatocarcinogenesis. Thus, several protein peaks were shown to be significantly deregulated with respect to the different phenotypes of cirrhotic nodules and HCC. In fact, three and 12 protein peaks were associated with the clonal expansion and malignant transformation of cirrhotic nodules, respectively. Based upon their m/z ratio, it appears that different proteins, except for one, are involved in these two main steps of progression. This suggests that development of HCC requires various proteins that are specifically induced or repressed during clonal expansion and malignant transformation of cirrhotic nodules. According to the design of this study, mainly restricted to protein analysis of the epithelial compartment using LCM microdissection, the resulting protein peaks have certainly a molecular relevance in the cellular process of malignancy. These results are in line with a recent study that assessed molecular changes with gene expression profiling in dysplastic nodule and overt HCC.32 These authors sampled and compared macronodules that differ according to their macroscopic and microscopic features and they also show molecular demarcations between the different groups, suggesting also that liver carcinogenesis is a multistep molecular process. Our study differs from the study of Nam et al, since we focus our efforts on cirrhotic nodules that were very similar on a morphological basis (cirrhosis regenerative nodules) and differ only on the basis of their clonal status. Our study demonstrates that even at this stage and in the absence of morphological markers, molecular changes can be detected at the protein expression level. It would be of great interest to investigate whether these proteomic changes are present even at an earlier stage according to the hypothesis of Theise et al,33 who suggests that clonal areas resistant to scarring should expand at the precirrhotic stage and result in neoplastic islands of hepatic parenchyma
These results are encouraging, and will lead to further characterization of some of these peaks that could represent potential key factors in the pathogenesis of development of HCC, as well as being accurate markers of the different steps of the disease.
In conclusion, this study confirms that proteome analysis, as evaluated by SELDI-TOF, can be achieved from a limited number of microdissected cells, and it provides additional evidence for the multistep process of malignant transformation of cirrhotic tissue.
References
Schafer DF, Sorrell MF . Hepatocellular carcinoma. Lancet 1999;353:1253–1257.
Hiroashi S, Ishak KG, Kojiro M, et al. World Health Organization Classification of Tumors. Pathology and Genetics of Tumors of the Digestive System. IARC Press: Lyon, 2000, pp 159–172.
Thorgeirsson SS, Grisham JW . Molecular pathogenesis of human hepatocellular carcinoma. Nat Genet 2002;31:339–346.
Aihara T, Noguchi S, Sasaki Y, et al. Clonal analysis of regenerative nodules in hepatitis C virus-induced liver cirrhosis. Gastroenterology 1994;107:1805–1811.
Ochiai T, Urata Y, Yamano T, et al. Clonal expansion in evolution of chronic hepatitis to hepatocellular carcinoma. Hepatology 2000;31:615–621.
Paradis V, Laurendeau I, Vidaud M, et al. Clonal analysis of macronodules in cirrhosis. Hepatology 1998;28:953–958.
Paradis V, Dargere D, Bonvoust F, et al. Clonal analysis of micronodules in virus C-induced liver cirrhosis using laser capture microdissection (LCM) and HUMARA assay. Lab Invest 2000;80:1553–1559.
Aihara T, Noguchi S, Sasaki Y, et al. Clonal analysis of precancerous lesion of hepatocellular carcinoma. Gastroenterology 1996;111:455–461.
Yeh SH, Chen PJ, Shau WY, et al. Chromosomal allelic imbalance evolving from liver cirrhosis to hepatocellular carcinoma. Gastroenterology 2001;121:699–709.
Colombat M, Paradis V, Bièche I, et al. Quantitative RT-PCR in cirrhotic nodules reveals gene expression changes associated with liver carcinogenesis. J Pathol 2003;201:260–267.
Anderson L, Seilhamer J . A comparison of selected mRNA and protein abundances in human liver. Electrophoresis 1997;18:533–537.
Vlahou A, Schellhammer PF, Mendrinos S, et al. Development of a novel proteomic approach for the detection of transitional cell carcinoma of the bladder in urine. Am J Pathol 2001;158:1491–1502.
Petricoin EF, Ornstein DK, Paweletz CP, et al. Serum proteomic patterns for detection of prostate cancer. J Natl Cancer Inst 2002;94:1576–1578.
Qu Y, Adam BL, Yasui Y, et al. Boosted decision tree analysis of surface-enhanced laser desorption/ionization mass spectral serum profiles discriminates prostate cancer from noncancer patients. Clin Chem 2002;48:1835–1843.
Cazares LH, Adam BL, Ward MD, et al. Normal, benign, preneoplastic, and malignant prostate cells have distinct protein expression profiles resolved by surface enhanced laser desorption/ionization mass spectrometry. Clin Cancer Res 2002;8:2541–2552.
Zhang Z, Bast RC, Yu Y, et al. Three biomarkers identified from serum proteomic analysis for the detection of early stage ovarian cancer. Cancer Res 2004;64:5882–5890.
Paradis V, Degos F, Dargere D, et al. Identification of a new marker of hepatocellular carcinoma by serum protein profiling of patients with chronic liver diseases. Hepatology 2005;41:40–47.
Zhang SH, Cong WM, Wu MC . Focal nodular hyperplasia with concomitant hepatocellular carcinoma: a case report and clonal analysis. J Clin Pathol. 2004;57:556–559.
Tang LH, Hui P, Garcia-Tsao G, et al. Multiple angiomyolipomata of the liver: a case report. Mod Pathol 2002;15:167–171.
Gaffey MJ, Iezzoni JC, Weiss LM . Clonal analysis of focal nodular hyperplasia of the liver. Am J Pathol 1996;148:1089–1096.
Banks RE, Dunn MJ, Hochstrasser DF, et al. Proteomics: new perspectives, new biomedical opportunities. Lancet 2000;356:1749–1756.
Wulfkuhle JD, Paweletz CP, Steeg PS, et al. Proteomic approaches to the diagnosis, treatment, and monitoring of cancer. Adv Exp Med Biol 2003;532:59–68.
Wulfkuhle JD, Liotta LA, Petricoin EF . Proteomic applications for the early detection of cancer. Nat Rev Cancer 2003;3:267–275.
Yokoyama Y, Kuramitsu Y, Takashima M, et al. Proteomic profiling of proteins decreased in hepatocellular carcinoma from patients infected with hepatitis C virus. Proteomics 2004;4:2111–2116.
Zeindl-Eberhart E, Haraida S, Liebmann S, et al. Detection and identification of tumor-associated protein variants in human hepatocellular carcinomas. Hepatology 2004;39:540–549.
Park KS, Cho SY, Kim H, et al. Proteomic alterations of the variants of human aldehyde dehydrogenase isozymes correlate with hepatocellular carcinoma. Int J Cancer 2002;97:261–265.
Yoon GS, Lee H, Jung Y, et al. Nuclear matrix of calreticulin in hepatocellular carcinoma. Cancer Res 2000;60:1117–1720.
Issaq HJ, Veenstra TD, Conrads TP, et al. The SELDI-TOF MS approach to proteomics: protein profiling and biomarker identification. Biochem Biophys Res Commun 2002;292:587–592.
Petricoin EF, Liotta LA . SELDI-TOF-based serum proteomic pattern diagnostics for early detection of cancer. Curr Opin Biotechnol 2004;15:24–30.
Petricoin EF, Ornstein DK, Liotta LA . Clinical proteomics: applications for prostate cancer biomarker discovery and detection. Urol Oncol 2004;22:322–328.
Li C, Hong Y, Tan YX, et al. Accurate qualitative and quantitative proteomic analysis of clinical hepatocellular carcinoma using laser capture microdissection coupled with and two-dimensional liquid chromatography mass spectrometry. Mol Cell Proteomics 2004;3:399–409.
Nam SW, Park JY, Ramasamy A, et al. Molecular changes from dysplastic nodule to hepatocellular carcinoma through gene expression profiling. Hepatology 2005;42:809–818.
Theise ND, Park YN, Kojiro M . Dysplastic nodules and hepatocarcinogenesis. Clin Liver Dis 2002;6:497–512.
Acknowledgements
This study was supported by a grant from ARC (Association de la recherche sur le cancer) and CAMPLP (Caisse d’Assurance Maladie PLP).
Author information
Authors and Affiliations
Corresponding author
Additional information
Duality of interest
None declared.
Rights and permissions
About this article
Cite this article
Guedj, N., Dargere, D., Degos, F. et al. Global proteomic analysis of microdissected cirrhotic nodules reveals significant biomarkers associated with clonal expansion. Lab Invest 86, 951–958 (2006). https://doi.org/10.1038/labinvest.3700450
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700450