Abstract
The question of whether any genetic differences exist between primary and colorectal cancers (CRCs) and their metastatic foci is controversial. To look for genetic aberrations involved in metastasis of CRCs to the liver, we performed subtractive comparative genomic hybridization (CGH) experiments using paired samples from 20 CRC patients with primary tumors and synchronous or metachronous liver metastases. Relatively frequent gains in DNA copy number were detected at 6p, suggesting the presence of one or more metastasis-related genes in the region. Analysis of 11 CRC cell lines using array-based CGH (CGH-array) revealed one 6p candidate gene, CCND3. Quantitative reverse transcriptase-polymerase chain reaction experiments showed that CCND3 was significantly upregulated in liver-metastatic lesions compared with primary lesions (P<0.0152). In addition, immunohistochemical analysis of 120 primary CRC tumors demonstrated that cyclin D3 expression in the region of rolled edge was significantly associated with total recurrence, especially hematogenous recurrence (P=0.0307). The results implied involvement of cyclin D3 in liver metastasis of CRC, and the data may contribute to the development of a novel therapy or diagnostic agent for this currently intractable disease. Our experiments also confirmed the power of subtractive CGH and CGH-array analysis for identifying cancer-related genes.
Similar content being viewed by others
Main
Colorectal cancer (CRC) is one of the most common forms of human malignancy, especially in industrialized countries, and metastasis of those tumors to the liver is a major cause of death among patients with CRC. The progression of cancer cells to a metastatic phenotype is thought to occur in multiple steps that include cytoskeletal changes, loss of adhesion, enhanced mortality, expression of proteolytic enzymes that degrade the basement membrane, adhesion to endothelial cells, anchorage-independent growth, and angiogenesis. Given the highly complicated nature of the molecular pathogenesis of metastasis, much remains to be learned about this lethal process.
Multiple genetic alterations occurring sequentially in a cell lineage underlie the carcinogenetic process in solid tumors.1 Among those genetic alterations, amplification of chromosomal DNA is one of the mechanisms capable of activating genes whose aberrant expression is critical in the development and progression of cancer. Comparative genomic hybridization (CGH) analysis2 has proven to be useful for identifying novel regions of gene amplification in various types of cancers.3, 4, 5, 6 Moreover, a recent development, the CGH-array technique, allows high-throughput and quantitative analysis of copy-number changes at high resolution throughout cancer genomes, providing many advantages over conventional methods and often allow precise and rapid identification of tumor suppressor genes as well as oncogenes.7, 8, 9, 10
Whether any genetic differences exist between primary colorectal carcinomas and liver metastases remains controversial even in the postsequence era. CGH analysis has been performed in primary tumors and metastatic tumors of CRC to compare CGH profiles between those two groups. However, only a few published studies have analyzed paired samples, and only in limited numbers.11, 12, 13, 14 To determine genetic aberrations responsible for the metastatic phenotype in CRC, it is necessary to compare paired tumor tissues obtained from original and metastatic sites in the same patient.
In the work reported here, we examined 20 primary CRC tumors and their corresponding metastatic foci in the liver by conventional CGH and subtractive CGH analyses, to explore genomic alterations that might be associated with metastasis of CRC. With this approach, we identified 6p as a candidate region for harboring one or more metastasis-related genes. A CGH-array analysis of CRC cell lines using our custom-made array (‘MCG Cancer Array-800’; Inazawa et al,8 Sonoda et al9 and Takada et al10), and analysis of gene expression by means of real-time quantitative reverse transcriptase-polymerase chain reaction (RT-PCR) experiments revealed CCND3 as the most probable target of additional amplification during metastasis. To clarify the significance of cyclin D3 overexpression in colorectal carcinogenesis and metastasis to the liver, we examined associations between the expression of cyclin D3 protein and clinicopathological characteristics of 120 primary CRCs.
Materials and methods
Cell Lines and Primary Tumors
Among the 11 human CRC cell lines selected for our experiments, COLO205, HCT-15, and SW480 were generously provided by Cell Resource Center for Biomedical Research, Institute of Development, Aging and Cancer, Tohoku University (Sendai, Japan). CCK-81, CoCM-1, COLO201, COLO320DM, DLD-1, CaR-1, and WiDr were purchased from the Japanese Collection of Research Bioresources (Osaka, Japan), and HT-29 from the American Type Culture Collection (Manassas, VA, USA). CCK-81, COLO320DM, and WiDr were maintained in Dulbecco's modified Eagle's MEM supplemented with 10% fetal bovine serum (FBS) and 100 U/ml penicillin/100 μg/ml streptomycin (P/S). HT-29 and CoCM-1 were maintained in McCoy's 5a and RPMI-1640 (50%)/F-12 (50%), respectively, each supplemented with 10% FBS and P/S. All others were maintained in RPMI-1640 supplemented with 10% FBS and P/S. HSC-4 and HO-1-u-1 cell lines derived from oral squamous cell carcinoma were maintained in Dulbecco's modified Eagle's MEM supplemented with 10% FBS and P/S.
Paired samples of primary CRCs and corresponding normal colonic mucosa and metastatic foci (14 with synchronous liver metastases and six with metachronous liver metastases) for CGH experiments were obtained from 20 unrelated patients being treated at the Tokyo Medical and Dental University Hospital, after written consent from each patient in the formal style and approval by the local ethics committee. Tissues from these patients were frozen immediately in liquid nitrogen and stored at −80°C until required. Clinicopathological data were collected on the basis of General Rules for Clinical and Pathological Studies on Cancer of the Colon, Rectum, and Anus by the Japanese Society for Cancer of the Colon and Rectum, and clinical stages of the disease were classified according to the tumor-node metastasis (TNM) classification of the International Union Against Cancer (Table 1).
Paraffin-embedded specimens of primary pT3 CRCs to be used for immunohistochemistry (IHC) were obtained from 120 unrelated patients treated at the National Defense Medical College Hospital, with written consent from each patient in the formal style and after approval by the local ethics committee. Clinicopathological data were collected on the basis of General Rules for Clinical and Pathological Studies on Cancer of the Colon, Rectum, and Anus by the Japanese Society for Cancer of the Colon and Rectum. Clinical stages of the disease were classified according to the TNM classification of the International Union Against Cancer; 58 patients were at stage II, 39 at stage III, and 23 at stage IV. All patients were periodically followed up at outpatient clinics and monitored for postoperative recurrence every 3 months by chest X-rays and measurements of serum carcinoembryonic antigen and carbonhydrate antigen 19-9 levels; every 6 months by abdominal ultrasonography; and every year by colonoscopy. Contrast-enhanced computed tomography was performed when recurrence of cancer was suspected. Whenever no findings suggestive of cancer relapse appeared during 5 years of follow-up, the procedure was changed to an annual physical check without other detailed examinations. Once a year, we confirmed by telephone the conditions of patients who did not visit the clinic. The duration of overall survival was calculated for each patient from the date of primary surgery to the date of the last follow-up or death. Of the 120 patients, 48 died of cancer during the study period, with a median interval of 26.3 months (range 1.8–72.7 months) from surgery to death. The median follow-up period of the 72 survivors was 79.0 months (range 40.0–135.9 months). The others were dead, but not by cancer.
CGH and Subtractive CGH
We used directly fluorochrome-conjugated DNA for CGH experiments as described by Kallioniemi et al,2 with minor modifications.3 DNAs extracted from normal colonic mucosa corresponding to tumor samples were used as reference. Briefly, test and reference DNAs were labeled, respectively, with Spectrum Green- and Orange-dUTP (Vysis, Chicago, IL, USA) by nick translation, denatured, and hybridized to normal male metaphase chromosome spreads together with Cot-1 DNA. Shifts in CGH profiles were rated as gains and losses if they reached at least the 1.2 and 0.8 thresholds, respectively, and over-representations were considered to be high-level gains (HLGs) when the fluorescence ratio exceeded 1.5, as described elsewhere.3 In subtractive CGH, DNAs from metastatic tumors and primary tumors were labeled, respectively, with Spectrum Green- and Orange-dUTP by nick translation. Heterochromatic regions near centromeres, and Y chromosomes, were excluded from these analyses.
CGH-Array Analysis
Our MCG Cancer Array-8008, 9, 10 was used for CGH-array experiments, which were carried out as described elsewhere.9, 10 DpnII-restricted test and reference DNAs were labeled by random priming with Cy3- and Cy5-dCTP (Amersham Biosciences, Tokyo, Japan), respectively, precipitated together with ethanol in the presence of Cot-1 DNA, redissolved in a hybridization mix (50% formamide, 10% dextran sulfate, 2 × standard saline citrate (SSC), 4% sodium dodecyl sulfate (SDS), pH 7), and denatured at 75°C for 10 min. After incubation at 37°C for 30 min, the mixture was applied to array slides set up in custom-made hybridization chambers, and incubated at 42°C on a rocking table for 48–72 h. The hybridized slides were washed once in a solution of 50% formamide, 2 × SSC (pH 7.0) for 15 min at 50°C, once in 2 × SSC, 0.1% SDS for 15 min at 50°C, and once in a 0.1 mol/l sodium phosphate buffer containing 0.1% Nonidet P-40 (pH 8) for 15 min at room temperature.
The arrays were scanned with a GenePix 4000B (Axon Instruments, Foster City, CA, USA), and the acquired images were analyzed with GenePix Pro 4.1 imaging software (Axon Instruments). Fluorescence ratios were normalized so that the mean of the middle third of log 2 ratios across the array was zero. Average ratios that deviated significantly (>2 s.d.) from zero were considered abnormal.
Fluorescence In Situ Hybridization (FISH)
Metaphase chromosome slides were prepared in the manner described previously.3 BACs containing the CCND3 gene (RP11-720D9) were labeled with biotin-16-dUTP by nick translation (Roche Diagnostics, Tokyo, Japan), and hybridized to metaphase chromosome spreads together with Cot-1. The chromosomes were counterstained with 4′,6-diamidino-2-phenylindole.
Real-Time Quantitative RT-PCR
Expression levels of CCND1, CCND2, CCND3, and CDK6 mRNA were measured by means of a real-time fluorescence detection method.5 Single-stranded cDNAs were generated from total RNAs using the SuperScript™ First-Strand Synthesis System (Invitrogen, Carlsbad, CA, USA). Real-time quantitative PCR was performed with an ABI PRISM 7900HT Sequence Detection System (Applied Biosystems, Foster City, CA, USA) using SYBR Green, according to the manufacturer's protocol; primer sequences are available on request. The glyceraldehyde-3-phosphate dehydrogenase (GAPDH) gene served as an endogenous control (Applied Biosystems); the level of mRNA expression for each gene in each sample was normalized on the basis of the respective GAPDH content and recorded as a relative expression level. PCR amplification was performed in duplicate for each sample.
Immunohistochemistry
Indirect IHC was performed on formalin-fixed, paraffin-embedded tissue sections, as described elsewhere.15 Using tissue blocks prepared from pT3 primary CRCs resected from 120 patients, we constructed tissue-microarray (TMA) blocks of tissue-core specimens taken from the submucosal invasive front, subserosal invasive front, central area, and rolled edge of each tumor.16 To construct TMA blocks, four tissue cores were taken from each representative tissue block, and these cores were transferred to a recipient block using a Tissue Microarrayer (Beecher Instruments, Silver Spring, MD, USA). We used cores 2.0 mm in diameter and arranged them 0.7–0.8 mm apart in a recipient block, to construct a total of 15 TMA sets comprising 480 core specimens. The sections were dewaxed and rehydrated in graded concentrations of ethanol. Antigens were retrieved by microwave pretreatment in 10 mM citrate buffer (pH 6.0) for 10 min. After cooling, the sections were treated with 3% hydrogen peroxide to block endogenous peroxidase, then reacted overnight at 4°C with antihuman cyclin D3 monoclonal antibody (1:50, DSC-22; DAKO, Santa Barbara, CA, USA) or normal rabbit serum. The sections were rinsed, incubated with rabbit EnVision+peroxidase (Dako, Carpinteria, CA, USA), stained with 0.05% hydrogen peroxide and 3,3′-diaminobenzidine, and counterstained with hematoxylin. HSC-4 and HO-1-u-1 cell lines served as positive controls, and also as negative controls where the primary antibody was omitted. In the HSC-4 and HO-1-u-1 cells, cyclin D3 was strongly stained in about 70% and about 5% of carcinoma cell nuclei, respectively.
Expression levels of cyclin D3 were divided into four categories according to the percentage of cyclin D3-positive cells (nuclei) in a sample as follows: irrespective of the intensity of immunoreaction, no positive cells=0; positive in <5% of constituent carcinoma cells=+1; positive in 5–50% of constituent carcinoma cells=+2; and positive in >50% of constituent carcinoma cells=+3. CRC samples that registered levels 0 or +1 were defined as negative expression, and samples containing levels +2 or +3 were defined as positive.
Statistical Analysis
Differences in levels of CCND3 mRNA expression between subgroups were tested by the nonparametric Wilcoxon's rank test with the determination of associated probability (P). Correlations between cyclin D3 expression in primary CRCs and clinicopathological variables were analyzed for statistical significance by χ2 or Fisher's exact tests. Survival data were analyzed according to the method of Kaplan and Meier. The log-rank test was used to compare survival data with cyclin D3 expression patterns. P-values of less than 0.05 were considered significant.
Results
Chromosomal Regions Frequently Involved in DNA Copy-Number Aberrations in CRCs
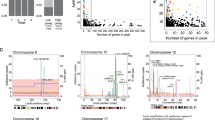
To explore chromosomal regions that might harbor genes critical to metastatic progression of CRC, we first carried out a conventional CGH analysis of 20 primary CRC tumors and their corresponding liver metastases. DNA extracted from normal colonic mucosa corresponding to the tumor sample served as a reference for each case. Overviews of genetic changes in 20 primary (P) and 20 metastatic (M) tumors are shown in Figure 1a and b, respectively. Most of the samples showed copy-number aberrations (P, 15/20, 75%; M, 18/20, 90%). On average, we observed 9.3 (range, 0–27) aberrations in primary tumors per patient: 4.5 (range, 0–10) gains and 4.8 (range, 0–19) losses. Metastatic tumors showed an average of 9.5 (range, 0–19) aberrations: 5 (range, 0–8) gains and 4.5 (range, 0–12) losses.
Summary of genetic imbalances detected by CGH in 20 paired samples. The 22 autosomes and X chromosome are represented by ideograms showing G-banding patterns. As judged by the computerized green-to-red profiles, vertical lines on the left of each ideogram show losses of genomic material indicated by their identification numbers; those on the right correspond to copy-number gains. HLGs are represented as open rectangles. (a) Conventional CGH in 20 primary tumors. (b) Conventional CGH in liver metastases of the same tumors. (c) Subtractive CGH in the 20 paired samples.
Regions with the most frequent copy-number gains in primary tumors were at 13q (70%), 20q (65%), 8q (55%), 20p (40%), 7p and 7q (25% each), and 1q (20%). Common losses in primary tumors were seen at 18q (55%), 18p and 8p (50% each), 1p (40%), 17p (35%), 4q (30%), 4p (25%), and 3p (35%). The smallest regions of HLGs in primary tumors involved 8q23-qter (five cases), 13q32-qter and 20q12-qter (four cases each), and 20p (three cases). Common regions for the most frequent copy-number gains in metastatic tumors were at 8q (70%), 20q and 13q (60% each), 20p (45%), 7p (40%), 7q (35%), Xq (25%), 1q (20%), and 6p (15%). Common losses in metastatic tumors were seen at 8p (60%), 18p and 18q (55% each), 14q (30%), 17p (25%), 4p and 22q (20% each), 4q (15%), and 1p (15%). The smallest regions of HLGs in metastatic tumors involved 8q13-q21.3, 13q14-qter, 20q12-qter and 20p11.2-p12 (four cases each), and 12p12-pter (two cases each).
Subtractive CGH analysis to compare primary CRC tumors with paired metastatic liver tumors revealed no significant differences at chromosomes 7, 8, 13, 18, and 20, where the aberrations noted above were observed in both primary and metastatic tumors. Using subtractive CGH, on the other hand, in three of the 20 patients, we detected gains on the whole arm of 6p (Figure 1b, c) that were not present in the corresponding primary tumors (Figure 1a). Interestingly, liver metastases of all these three cases were metachronous. None of the synchronous liver metastases showed gains on 6p compared with corresponding primary tumors.
Identification of HLGs of CCND3 by CGH-Array in 11 CRC Cell Lines
To screen for amplified genes on 6p, we applied CGH-array analysis to the three metastatic tumor samples that showed gains of 6p, and to 11 CRC cell lines. Using the MCG Cancer Array-800, we identified several genes whose copy numbers were frequently increased or decreased (Tables 2 and 3) without evidence of any homozygous deletions. Among the 19 genes/loci within 6p that were spotted on the MCG Cancer Array-800, CCND3, located at 6p21.1 (log 2 ratio=2.25/2.28), was the only one amplified, and in only two of the cell lines (COLO201, Figure 2a and c, and COLO205, Figure 2c). Amplification of CCND3 was confirmed by FISH analysis: BAC RP11-720D9, which contains the CCND3 gene, generated a remarkably increase in FISH signals in COLO201 (Figure 2b); a similar result was obtained with COLO205 (data not shown). In the three metastatic tumor samples, on the other hand, the pattern of copy-number alterations was similar among all of the 19 6p genes/loci examined (data not shown).
(a) Representative CGH-array image of the COLO201 cell line. An increase in copy number of CCND3 at 6p21.1 was detected as a clear green signal (log 2 ratio=2.25). (b) Representative FISH image, using a CCND3-specific BAC (RP11-720D9) as a probe in the COLO201 cell line. BAC RP11-720D9 generated clear signals as an HSR pattern on these metaphase chromosomes. (c) Quantitative CGH-array results on 6p in the COLO201 and COLO205 cell lines.
Expression of D-Type Cyclins and Cyclin-Dependent Kinase in Primary and Metastatic Foci of CRC
To confirm that expression of cyclin D3 might be associated with liver metastasis, we used real-time quantitative RT-PCR to assess the expression of CCND3, other D-type cyclins, and CDK6, a main partner of cyclin D3, in all of our paired samples. As shown in Figure 3, all of the D-type cyclins were upregulated in primary tumors compared with normal mucosa, but only CCND3 was significantly upregulated further in metastatic lesions (P=0.0152). Increased expression of cyclin D3 was also observed at the protein level using IHC, where nuclear staining of cyclin D3 was observed in paired primary and metastatic lesions of CRC; very weak staining was observed in corresponding normal mucosa (Figure 4). CDK6 was also significantly upregulated in metastatic lesions (P=0.0366), suggesting that the cyclin D3–CDK6 complex may play a hitherto unexpected role in the metastatic process of CRC.
Relative levels of CCND1, CCND2, CCND3, and CDK6 mRNA compared among subgroups. Among the three D-type cyclin genes, only CCND3 mRNA was significantly upregulated in liver metastases compared with the corresponding primary lesions (P=0.0152). In addition, mRNA of CDK6, a partner of D-type cyclins, was also upregulated (P=0.0366). Mean values are indicated by filled circles; vertical bars indicate s.e. Wilcoxon's rank test was used for statistical analysis.
Association of Cyclin D3 Protein with Clinicopathological Features among Primary CRCs
To assess the clinical significance of cyclin D3 overexpression in CRCs, we performed immunohistological examinations using four-point TMA sets from 120 primary pT3 CRCs, and compared expression patterns among different tumor phenotypes. Table 4 summarizes relationships between expression status of cyclin D3 protein and clinicopathological features. Representative immunostaining patterns of cyclin D3 are shown in Figure 5a. The expression of cyclin D3 in rolled-edge regions was significantly related to total recurrence (P=0.0318), hematogenous recurrence (P=0.0307), and venous invasion (P=0.0217), although no significant relationship was detected in other regions. Univariate analysis of overall survival by the log-rank test demonstrated that cyclin D3 expression status in regions of rolled edge tended to be associated with poorer prognosis, although the difference did not reach significance (Figure 5b, P=0.1081). Notably, no deaths occurred in the cyclin D3-negative expression group of stage II CRCs during the study period, whereas 16.3% of patients in the positive group died (Figure 5c, P=0.0803). No significant correlation was seen between cyclin D3 expression status and age, sex, location, or histological subtype in any region of the pT3 tumors.
Expression of cyclin D3 protein in primary CRCs. (a) Representative staining patterns of cyclin D3 protein, from experiments using a TMA system to examine 120 cases of CRC. (b) Expression pattern of cyclin D3 in regions of rolled edge vs overall survival in patients with CRC. Cyclin D3 expression was associated with poorer prognosis, although the difference was marginally significant (P=0.1081). (c) Expression pattern of cyclin D3 in regions of rolled edge vs overall survival in patients with stage II CRC. There were no deaths in the cyclin D3-negative expression group of stage II CRCs.
Discussion
Carcinoma is a genetically heterogeneous disease. Animal studies suggest that multiple clones of malignant cells in a primary lesion contribute to the manifestation and evolution of the metastatic phenotype.17 Specific chromosomal alterations responsible for metastasis occur only in certain subpopulations of cells, and may not be distinguishable from the cancer-cell background by standard molecular analysis of the primary tumor. On the other hand, chromosomal alterations with relevance for the metastatic process might be enriched in metastatic foci, suggesting that comparison of genetic differences between primary and corresponding metastases may reveal metastasis-associated genes. In the study reported here, we assessed genetic differences between 20 primary CRCs and their corresponding metastatic tumors in the liver by subtractive CGH, and successfully identified gain of DNA on 6p as a liver metastasis-related chromosomal alteration.
The copy-number aberrations revealed through our CGH analysis were largely consistent with those already reported.11, 12, 13, 14, 18, 19 However, our direct comparisons of 20 paired samples by means of subtractive CGH analysis identified gain in 6p as a metastasis-specific change, but we found no significant differences at chromosomes 7, 8, 13, 18, or 20, where major chromosomal aberrations occurred in both primary and metastatic tumors. Al-Mulla et al11 also reported increased copy-number on 6p in CRCs, but only in tumors of Duke's stage D and liver metastases.
These circumstances prompted us to focus on 6p as a chromosomal region that might harbor genes responsible for liver metastasis, even though we could not identify a specific region of gain on 6p using clinical specimens. Therefore, we carried out CGH-array analysis of 11 CRC cell lines, which sometimes harbor remarkable copy-number changes as landmarks within specific regions. With this approach, we detected high-level amplification of CCND3 (6p21.1) in two cell lines (COLO201 and COLO205), even though most of the CRC cell lines we examined did not show genetic alterations on 6p. To address whether this event was related to liver metastasis, we assessed expression of CCND3 mRNA in our 20 paired clinical samples. Expression levels of CCND3 and CDK6, a partner of D-type cyclins, were relatively low in normal colonic mucosa, but in metastatic lesions, both genes were significantly more elevated than in primary lesions (P=0.0152 and 0.0366, respectively); other D-type cyclins, CCND1 and CCND2, were not upregulated further in metastatic lesions. These observations suggested that activation of CCND3 and CDK6 through overexpression may play a critical role in CRC tumorigenesis, especially in terms of metastasis to the liver.
Immunohistochemical analysis of cyclin D3 using area-specific four-point TMAs of primary CRCs revealed significant association of cyclin D3 expression in the region of rolled edge with total recurrence, especially hematogenous recurrence (P=0.0307), but it was not correlated with metastasis to lymph nodes. On the other hand, cyclin D3 expression in subserosal or submucosal invasion fronts, where malignant cells are recognized as more invasive, was not correlated with lymph-node metastasis or hematogenous recurrence. Recently, expression of laminin-5 gamma 2 chain in the invasive fronts of tumors was found to be an indicator of lymph-node metastasis and patient prognosis, but expression of that gene in the rolled edge was not.16 It appears that molecules that are important for tumor-cell proliferation and hematogenous recurrence, for example, cyclin D3, and those that are important for stromal invasion and lymph node metastasis, for example, laminin-5 gamma 2 chain, are expressed in different areas to allow the tumor to grow and spread with integrity.
The results reported here suggest that evaluation of cyclin D3 expression in rolled-edge regions of tumors, using preoperative biopsy specimens obtained by colonoscopic examination, may anticipate the risk of hematogenous recurrence of CRC. Furthermore, we observed no deaths among cyclin D3-negative patients with stage II CRCs, suggesting that overexpression of cyclin D3 may be a risk factor for recurrence in patients with nonsynchronous metastases.
Among three known D-type cyclins, cyclin D3 is the most widely expressed and, in several cell types, it appears to be the only expressed member of this cyclin subfamily.20 The CCND3 gene is rearranged and cyclin D3 protein is overexpressed in several human lymphoid malignancies21, 22 and in solid tumors as well.23, 24, 25, 26 Cyclin D3−/− laboratory animals fail to undergo normal expansion of immature T-lymphocytes and show greatly reduced susceptibility to T-cell malignancies triggered by specific oncogenic pathways.27 Therefore, cyclin D3 has carcinogenic potential, as do other D-type cyclins. On the other hand, tissue-specific expression patterns, different affinity for CDKs, and the fact that sequence comparisons reveal a higher degree of conservation between species than between other D-type cyclins within the same species, argue for a highly specific role of each D-type cyclin.28, 29 pRb and its relatives p130 and p107,30 which bind repressive E2Fs (E2F4 and E2F5), are substrates for cyclin D3-dependent kinase. Among the D-type cyclins, only cyclin D3 efficiently phosphorylated p130 in an in vitro kinase assay in mouse BALB/c 3T3 fibroblasts.31 p107-labeling indices decline in liver-metastatic foci, and also in large primary colorectal tumors of mucinous type or characterized by venous invasion, lymphatic invasion, poor differentiation, deep invasion, lymph-nodal metastasis, hepatic metastasis, or advanced stage.32 Furthermore, the cyclin D3–CDK6 complex has a unique ability to evade inhibition by p27KIP1 and p21CIP1.33 and is resistant to inhibition by p16INK4a.34 These findings strongly suggest that cyclin D3, when activated in CRC cells, may indeed play an important role in metastasis to the liver. Further investigations along this line may lead to new therapies for this currently intractable disease, such as interventions that would target cyclin D3.
References
Vogelstein B, Kinzler KW . The multistep nature of cancer. Trends Genet 1993;9:138–141.
Kallioniemi A, Kallioniemi OP, Sudar D, et al. Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors. Science 1992;258:818–821.
Fukuda Y, Kurihara N, Imoto I, et al. CD44 is a potential target of amplification within the 11p13 amplicon detected in gastric cancer cell lines. Genes Chromosomes Cancer 2000;29:315–324.
Imoto I, Yang ZQ, Pimkhaokham A, et al. Identification of cIAP1 as a candidate target gene within an amplicon at 11q22 in esophageal squamous cell carcinomas. Cancer Res 2001;61:6629–6634.
Yasui K, Arii S, Zhao C, et al. TFDP1, CUL4A, and CDC16 identified as targets for amplification at 13q34 in hepatocellular carcinomas. Hepatology 2002;35:1476–1484.
Saito-Ohara F, Imoto I, Inoue J, et al. PPM1D is a potential target for 17q gain in neuroblastoma. Cancer Res 2003;63:1876–1883.
Snijders AM, Nowak N, Segraves R, et al. Assembly of microarrays for genome-wide measurement of DNA copy number. Nat Genet 2001;29:263–264.
Inazawa J, Inoue J, Imoto I . Comparative genomic hybridization (CGH)-arrays pave the way for identification of novel cancer-related genes. Cancer Sci 2004;95:559–563.
Sonoda I, Imoto I, Inoue J, et al. Frequent silencing of low density lipoprotein receptor-related protein 1B (LRP1B) expression by genetic and epigenetic mechanisms in esophageal squamous cell carcinoma. Cancer Res 2004;64:3741–3747.
Takada H, Imoto I, Tsuda H, et al. Screening of DNA copy-number aberrations in gastric cancer cell lines by array-based comparative genomic hybridization. Cancer Sci 2005;96:100–110.
Al-Mulla F, Keith WN, Pickford IR, et al. Comparative genomic hybridization analysis of primary colorectal carcinomas and their synchronous metastases. Genes Chromosomes Cancer 1999;24:306–314.
Korn WM, Yasutake T, Kuo WL, et al. Chromosome arm 20q gains and other genomic alterations in colorectal cancer metastatic to liver, as analyzed by comparative genomic hybridization and fluorescence in situ hybridization. Genes Chromosomes Cancer 1999;25:82–90.
Aragane H, Sakakura C, Nakanishi M, et al. Chromosomal aberrations in colorectal cancers and liver metastases analyzed by comparative genomic hybridization. Int J Cancer 2001;94:623–629.
Diep CB, Parada LA, Teixeira MR, et al. Genetic profiling of colorectal cancer liver metastases by combined comparative genomic hybridization and G-banding analysis. Genes Chromosomes Cancer 2003;36:189–197.
Imoto I, Tsuda H, Hirasawa A, et al. Expression of cIAP1, a target for 11q22 amplification, correlates with resistance of cervical cancers to radiotherapy. Cancer Res 2002;62:4860–4866.
Shinto E, Tsuda H, Ueno H, et al. Prognostic implication of laminin-5 gamma 2 chain expression in the invasive front of colorectal cancers, disclosed by area-specific four-point tissue microarrays. Lab Invest 2005;85:257–266.
Bell C, Frost P, Kerbel RS . Cytogenetic heterogeneity of genetically marked and metastatically competent ‘dominant’ tumor cell clones. Cancer Genet Cytogenet 1991;54:153–161.
Ried T, Knutzen R, Steinbeck R, et al. Comparative genomic hybridization reveals a specific pattern of chromosomal gains and losses during the genesis of colorectal tumors. Genes Chromosomes Cancer 1996;15:234–245.
Rooney PH, Boonsong A, McKay JA, et al. Colorectal cancer genomics: evidence for multiple genotypes which influence survival. Br J Cancer 2001;85:1492–1498.
Bartkova J, Lukas J, Strauss M, et al. Cyclin D3: requirement for G1/S transition and high abundance in quiescent tissues suggest a dual role in proliferation and differentiation. Oncogene 1998;17:1027–1037.
Shaughnessy Jr J, Gabrea A, Qi Y, et al. Cyclin D3 at 6p21 is dysregulated by recurrent chromosomal translocations to immunoglobulin loci in multiple myeloma. Blood 2001;98:217–223.
Filipits M, Jaeger U, Pohl G, et al. Cyclin D3 is a predictive and prognostic factor in diffuse large B-cell lymphoma. Clin Cancer Res 2002;8:729–733.
Buschges R, Weber RG, Actor B, et al. Amplification and expression of cyclin D genes (CCND1, CCND2 and CCND3) in human malignant gliomas. Brain Pathol 1999;9:435–442.
Ito Y, Takeda T, Wakasa K, et al. Expression and possible role of cyclin D3 in human pancreatic adenocarcinoma. Anticancer Res 2001;21:1043–1048.
Hedberg Y, Roos G, Ljungberg B, et al. Cyclin D3 protein content in human renal cell carcinoma in relation to cyclin D1 and clinico-pathological parameters. Acta Oncol 2002;41:175–181.
Lopez-Beltran A, Luque RJ, Alvarez-Kindelan J, et al. Prognostic factors in stage T1 grade 3 bladder cancer survival: the role of G1-S modulators (p53, p21Waf1, p27kip1, Cyclin D1, and Cyclin D3) and proliferation index (ki67-MIB1). Eur Urol 2004;45:606–612.
Sicinska E, Aifantis I, Le Cam L, et al. Requirement for cyclin D3 in lymphocyte development and T cell leukemias. Cancer Cell 2003;4:451–461.
Sherr CJ . D-type cyclins. Trends Biochem Sci 1995;20:187–190.
Reed SI . Control of the G1/S transition. Cancer Surv 1997;29:7–23.
Herzinger T, Reed SI . Cyclin D3 is rate-limiting for the G1/S phase transition in fibroblasts. J Biol Chem 1998;273:14958–14961.
Dong F, Cress Jr WD, Agrawal D, et al. The role of cyclin D3-dependent kinase in the phosphorylation of p130 in mouse BALB/c 3T3 fibroblasts. J Biol Chem 1998;273:6190–6195.
Wu F, Li JQ, Miki H, et al. p107 expression in colorectal tumours rises during carcinogenesis and falls during invasion. Eur J Cancer 2002;38:1838–1848.
Lin J, Jinno S, Okayama H . Cdk6–cyclin D3 complex evades inhibition by inhibitor proteins and uniquely controls cell's proliferation competence. Oncogene 2001;20:2000–2009.
Faast R, White J, Cartwright P, et al. Cdk6–cyclin D3 activity in murine ES cells is resistant to inhibition by p16(INK4a). Oncogene 2004;23:491–502.
Acknowledgements
We are grateful to Professor Yusuke Nakamura (Human Genome Center, The Institute of Medical Science, The University of Tokyo) for his continuous encouragement throughout this work. We thank Ai Watanabe for technical assistance. Grant support: Grants-in-aid for Scientific Research (B) and Scientific Research on Priority Areas (C) and a Center of Excellence Program for Research on Molecular Destruction and Reconstruction of Tooth and Bone from the Ministry of Education, Culture, Sports, Science, and Technology of Japan; and from Core Research for Evolutional Science and Technology (CREST) of the Japan Science and Technology Corporation (JST).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tanami, H., Tsuda, H., Okabe, S. et al. Involvement of cyclin D3 in liver metastasis of colorectal cancer, revealed by genome-wide copy-number analysis. Lab Invest 85, 1118–1129 (2005). https://doi.org/10.1038/labinvest.3700312
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700312
Keywords
This article is cited by
-
MicroRNA-138 negatively regulates non-small cell lung cancer cells through the interaction with cyclin D3
Tumor Biology (2016)
-
Copy number alterations and allelic ratio in relation to recurrence of rectal cancer
BMC Genomics (2015)
-
Integrated genomic and functional analyses reveal glyoxalase I as a novel metabolic oncogene in human gastric cancer
Oncogene (2015)
-
SIX1 promotes epithelial–mesenchymal transition in colorectal cancer through ZEB1 activation
Oncogene (2012)
-
Tumor immunosurveillance in human cancers
Cancer and Metastasis Reviews (2011)