Abstract
The cyclin-dependent kinase (CDK) inhibitor p57Kip2 (CDKN1C) is a negative regulator of cell proliferation, binding to a variety of cyclin-CDK complexes and inhibiting their kinase activities in vitro. The p57Kip2 gene is imprinted and the maternal allele is expressed in terminally differentiated cells, including human β-cells. Somatic loss of p57Kip2 expression is associated with increased β-cell proliferation in the focal form of Hyperinsulinism of Infancy. We cloned and sequenced the rat ortholog of p57Kip2, and demonstrate that it is highly homologous to the mouse gene. However, the human and rodent genes are quite divergent. Despite having highly homologous C- and N-terminal domains, the mid-portion of the human gene is entirely different from that of its rodent counterparts. Expression of p57Kip2 was evaluated during fetal and postnatal development, and a highly cell-specific, temporal and spatial expression profile was found. In contrast to other tissues, the expression pattern in rat pancreas was entirely opposite from that previously reported in man, with high levels of expression in rodent exocrine cells, but no expression in β-cells during any stage of development. These findings demonstrate that p57Kip2 expression is highly regulated. In the pancreas, the functional significance of this gene appears to be quite different in humans when compared with rodents, suggesting that a better understanding of the function of this protein may provide new insights into the mechanisms involved in the control of human β-cell mass.
Similar content being viewed by others
Main
The cell cycle and consequently the proliferation of the mammalian cells are regulated by cyclins and cyclin-dependent kinases (CDKs). The CDK inhibitor p57Kip2 (CDKN1C) has the ability to bind with a variety of cyclin-CDK complexes and to inhibit their kinase activities in vitro.1, 2 Thus, p57Kip2 is a negative regulator of cell proliferation and may play a role in the maintenance of the nonproliferative state throughout life.
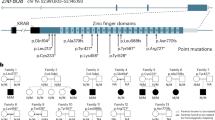
The p57Kip2 gene is an imprinted, maternally expressed gene, and is located within a cluster of imprinted genes on human chromosome 11p15.5 and mouse distal chromosome 7.3, 4, 5 In human and mouse, p57Kip2 is expressed in terminally differentiated cells during embryonic development,3, 4 and in some tissues the expression persists into adulthood. In humans, loss of the maternally derived 11p15.5 region is implicated in Beckwith–Wiedemann syndrome (BWS), which is characterized by congenital malformations and organomegaly, and is associated with an increased risk for the development of childhood neoplasms.6, 7, 8, 9 P57Kip2 mutations have been found only rarely in association with BWS,10, 11, 12 but a decrease in expression levels has been detected in several types of tumors.13, 14, 15, 16
Mice deficient in the p57Kip2 gene show defective endochondral bone formation. Most die soon after birth as a result of severe cleft palate.17, 18, 19 Zhang et al18 report that p57Kip2-deficient neonates have a 100% neonatal mortality and display organomegaly and abdominal wall defects, two of the hallmarks of BWS. However, others report that p57Kip2-deficient mice displayed no features of BWS, and a 10% survival rate was observed.17, 19 Surviving mutant mice were not report to develop hypoglycemia or tumors.17, 19
Hyperinsulinism of infancy (HI) is a rare genetic disease characterized by increased insulin secretion leading to potentially life-threatening hypoglycemia.20, 21 More than 50% of the cases are caused by mutations in either the sulfonylurea receptor-1 (SUR1, ABCC8) or the inward rectifying K+ channel (Kir6.2, KCNJ11), two subunits of the β-cell ATP-sensitive K+ channel (KATP channel). Histologically, the disease can present in two forms, diffuse and focal. The diffuse form is usually recessively inherited and involves all of the β-cells within the pancreas. Focal HI consists of adenomatous hyperplasia within a limited region of the pancreas, and is caused by somatic loss of heterozygosity (LOH) of maternal Ch11p15-ter in a β-cell precursor carrying a germ line mutation in the paternal allele of SUR1 or Kir6.2.22, 23, 24 β-Cells in focal-HI show increased proliferation when compared to normal β-cells or to those from patients with diffuse-HI.25 This region of Ch11 contains the imprinted genes p57Kip2 and IGF-II. Kassem et al26 demonstrated that p57Kip2 is expressed and paternally imprinted in human normal pancreatic β-cells and that β-cells in these focal lesions do not express p57Kip2. These findings led to the hypothesis that manipulation of p57Kip2 expression may result in altered β-cell proliferation. Specifically, suppression of p57Kip2 expression in vivo or in vitro may result in expansion of β-cell mass.25
Given the potential importance of p57Kip2 in cell cycle regulation, and because the rat is a very useful model for study, we characterized the rat ortholog of p57Kip2 and determined its expression profile, with a special emphasis on β-cell expression during development and in rodent β-cell lines. At the time of initiation of this study, no rat p57Kip2 sequence had been published or entered into the NCBI database.
Materials and methods
Cloning rp57Kip2 and Expression Profile
The rat expressed sequence tagged (EST) database was screened for sequences homologous to mouse Cdkn1c (Acc# U22399). Suitable ESTs were ordered, DNA was prepared (QIAGEN, Hilden, Germany), and the inserts were sequenced from the T3 and T7 primers.
The assembled cDNA sequence was confirmed by reverse transcriptase (RT)-PCR analysis. Total RNA was extracted from up to 100 mg samples of rat brain, kidney, heart, liver and pancreatic islets (Rattus norvegicus, Wistar strain, Hebrew University), using EZ-RNA reagent (Biological Industries, Kibbutz Beit Haemek, Israel). Reverse transcription was performed using Superscript II (Life Technologies, Inc., Paisley, UK). PCR reactions (1 cycle at 95°C for 5 min, followed by 34 cycles (p57Kip2) or 21 cycles (actin) at 94°C for 40 s, 54°C for 30 s, 72°C for 40 s, followed by one final cycle at 72° for 7 min) were performed with ReadyMix Taq (AB gene, Epsom, UK) with 4 pmol of each primer (Table 1) in a 10 μl reaction volume. PCR products were purified by Qiaquick PCR or gel purification kits (QIAGEN, Hilden, Germany), sequenced at the Hebrew University core laboratories using Big_Dye Terminator kit (Perkin-Elmer, Wellesley, MA, USA) and the ABI PRISM DNA Analyzer 3700 (Perkin-Elmer, Wellesley, MA, USA). Sequence comparisons and analysis were performed using the Lasergene package (DNASTAR Inc., Madison, WI, USA).
Expression level was determined by semiquantitative RT-PCR with β-actin as reference. We used equal amounts of input cDNA and varied the number of cycles for actin and p57Kip2 to keep the assays within the linear range; β-actin analyses were carried out in separate PCR reactions simultaneously. Primers for β−actin cDNA cross an intron, thus serving as an additional control for the absence of genomic DNA in RNA preparations. PCR products were run on 2% agarose gels containing ethidium bromide and quantified by densitometry using TINA 2.0 software (Raytest, Staubenhardt, Germany). Results are presented as the ratio of the p57Kip2 to β-actin band density.
Cells and Cell Culture
The cells (passages 52–63) were cultivated in a humidified atmosphere (5% CO2, 95% air at 37°C) in monolayer on plates (Nunc Brand Products, Nunc A/S, Roskilde, Denmark). The culture medium was changed twice weekly and the cells were passaged weekly. The INS-1 cells (Wollheim, Geneva, Switzerland) were grown in modified RPMI 1640 medium supplemented with 10% (vol/vol) FCS, 11.2 mM glucose, 10 mM HEPES, 100 IU/ml penicillin, 100 μg/ml streptomycin, 1 mM sodium pyruvate, 2 mM L-glutamate (Biological Industries, Kibbutz Beit Haemek, Israel) and 50 μM ß-mercaptoethanol (Fluka AG, Switzerland). MIN-6 cells were cultured in DMEM medium with 15% FCS, 7.5% NaHCO3 and 75 μM ß-mercaptoethanol. βTC cells were cultured in DMEM supplemented with 15% horse serum, 2.5% FCS and 2 mM L-glutamate.
Immunohistochemistry
Coronal and sagittal sections of whole sabra rat embryos (gestational ages 13.5, 15.5, 17.5, 19.5 and 21.5 days), and major organs dissected from rats aged 1, 2, 3, 7 and 9 weeks were fixed in 4% buffered formalin and embedded in paraffin. At least two rats were examined in each age group. Approval was obtained from the institutional ethics committee for animal experimentation.
Five micron sections were cut, placed on SuperFrost Plus slides (Menzel-Glaser, Germany), dried at 37°C overnight, deparaffinized in xylene and rehydrated in serial concentrations of alcohol (100, 90 and 80%) and double distilled water. Antigen retrieval was accomplished by microwave heating in 0.01 M citrate buffer (pH 6) at full power until boiling, followed by 15 min at 20% power.27 Slides were cooled at room temperature (RT) for 30 min and blocked by nonimmune serum for 10 min at RT prior to application of each primary antibody. Slides were incubated with rabbit anti-rat p57Kip2 (Sigma, St Louis, Missouri) for 1 h at 37°C, dilution 1:200. Bound antibody was detected using the streptavidin biotin-peroxidase kit (Zymed, cat#85-9943) and developed with aminoethylcarbazole (AEC).
Sections that contained pancreatic tissue were further stained for insulin, using guinea pig anti-insulin antibody (Dako, Denmark) for 1 h at 37°C, dilution 1:100. Detection was accomplished using the streptavidine biotin-alkaline phosphatase kit, developed with 5-bromo-4-chloro-3-indolyl phosphate/nitro blue tetrazolium (BCIP/NBT, Sigma). Each batch contained a negative control that consisted of sections incubated with PBS without primary antibody.
The Databases and Computer Programs Used for Analyses
The following databases and computer programs were used in the sequence analyses reported here: http://www.ncbi.nlm.nih.gov/; http://ftp.genome.washington.edu/cgibin/RepeatMasker; http://emboss.md.huji.ac.il and the LASERGENE software package (DNASTAR Inc., Madison, WI, USA).
Results
Rat p57Kip2-Expressed Sequence
Screening the rat EST database for sequences homologous to mouse Cdkn1c, three highly homologous EST clones (BI297597, BI283172, BF525135) were identified. These clones, which were derived from rat embryo cDNA libraries, were obtained from Research Genetics (Invitrogen Corp., University of Iowa). After sequencing the ends of the inserts, only one, BF525135, was confirmed to be similar to mouse p57Kip2, and was shown to contain the entire rat p57Kip2 cDNA, including the 5′ untranslated region (5′-UTR). The entire sequence of the rat gene was thus determined and submitted to Genbank (AC. #AY351679). The computer-assisted alignment of mouse and this clone of rat p57Kip2 revealed a similarity of 93–98% between their respective coding and 3′ untranslated region (3′-UTR) regions, but no similarity in the 5′-UTR.
The accuracy of the gene sequence, as determined from the BF525135 clone, was confirmed by RT-PCR amplification and sequencing from different rat tissues. Attempts were made to PCR amplify rat liver cDNA using several different combinations of primers. Primer sets including the middle portion of the gene frequently failed to produce any product, whereas reactions using primers in the first and last third of the gene consistently resulted in successful amplifications. A close examination of the sequence, using the program REPEAT SEQ, revealed that the region between nucleotides 552 and 964 contains simple repeats (CCCCAC)n from 552 to 684 bp and GA-rich repeat from 804 to 964 bp. These sequences typically make PCR amplification difficult or impossible. In fact, the entire coding region and 5′-UTR of p57Kip2 gene are remarkably GC-rich (67%). In contrast, in 3′-UTR this value is only 43.7%.
The results of automatic and manual sequencing of the PCR fragment amplified using primers p57F903A–p57kip2R showed a mixed pattern. Further analysis revealed alternative splicing corresponding to two messages generated by the use of either of the two distinct 5′ splice donor sites of exon 1 (Figure 1). The first is present in the EST BI283172 clone, and is referred to here as rat p57Kip2 long (AC# AY351678), consisting of an open reading frame from 146 to 1211, coding for 355 amino acids. The second is an in-frame deletion of 36 nucleotides 3′ from 1028 bp of p57Kip2 long and is referred to as p57Kip2sh (abbreviation of short) (AC# AY351679), with an open reading frame from 146 to 1175 bp. The assembled transcript encodes a predicted protein with 343 amino acids. EST BF525135 contains the short isoform of the gene. During preparation of this manuscript, additional mouse ESTs were entered into the NCBI database and repeat alignment analysis revealed the presence of a similar splice variant in the mouse. Mouse published ortholog (U22399) is similar to the short variant of the rat one (91.2% of overall similarity). In contrast, this splice variant was not observed in human p57Kip2.
Schematic representation of the genomic organization of ortholog p57Kip2 genes. Shaded boxes indicate exons with their length indicated within the boxes and the lines are GC-rich introns with their length indicated underneath. Alternative splicing generates amino-terminus heterogeneity that is shown by different hatching. The two-direction dotted arrow shows the alternative intron in the 5′-UTR with its alternative size beneath (see Figure 2 for details). The crosshatched box at the end of exon 2 indicates a segment added by alternative splicing that was found only in rodent p57Kip2. The thick vertical line indicates the location of the major start codon for translation with its base number noted above. The vertical dotted lines in exon 2 of hp57Kip2 indicate four in-frame 12 bp polymorphisms that were previously reported.29 The small horizontal arrows indicated the location of the primers, numbered according to Table 1. The small dotted arrows represent the location of the primers used to span the various splice sites.
While the manuscript was in preparation, we performed periodic in silico EST database searches, and additional Cdkn1c sequences were identified. First, one sequence was published without 5′-UTR (AC XM_341966). This was followed by several other rat Cdkn1c EST sequences, which revealed three different 5′-UTR variants. These three transcripts are generated by alternative splicing at different 3′ splice acceptor sites (Figures 1 and 2). The whole intron is transcribed in variant A1. Alternative splicing in isoforms A2 and A3 results from the use of different acceptor sites. To confirm these in silico results, cDNA was amplified by PCR with two different forward primers and the same reverse primer, which is located inside the coding region of the gene. The first pair was p57F16m and p57R548m, giving a 408 or 445 bp product depending on whether the A2 or the A3 variant was amplified. The second pair was p57F19 and p57R548m, giving a 420 bp product if the A1 variant was present. The resultant products were gel purified and submitted for sequencing, which confirm the in silico analysis and demonstrate three splice variants: variant A1 has three exons whereas variants A2 and A3 each have four exons, with the first intron size of 173 and 135 bp, respectively. The deduced sequences were deposited in GenBank (AC #AY576608 (A2) and #AY586728 (A3)).
(a) Schema of alternative splice isoforms of rat p57Kip2 transcript. Alternative splicing generates amino-terminus heterogeneity. Sequence analysis and ESTs database screening suggests that three different rat p57Kip2 transcripts are generated by intron splicing at different 3′splice acceptor sites. Three types, A1, A2 and A3, are shown. ATG in bold italics indicates the minor transcription site, ATG in bold and underlined indicates the location of the major transcription site. (b) The locations of the PCR primers that were used to detect 5′-UTR alternative isoforms are denoted by horizontal arrows (the numbers above refer to Table 1). The predicted sizes of the respective RT-PCR products are indicated.
The initiation methionine was assigned to the first in-frame ATG translation start codon (Figure 2). This proposed start codon lies within a sequence context (CGACCATGG), which conforms to the Kozak rule.28 The A3 splicing event eliminates this translation start codon and creates a second open reading frame, although its ATG initiation lies within a weaker context (CAGCGATGG) than the first one. EST BF525135 contains the truncated A1 isoform of the p57Kip2 short cDNA.
Screening of mouse and human EST databases revealed strikingly analogous splice isoforms of the 5′-UTR region, with only a few base substitutions in the mouse (95% similarity). In contrast, the human 5′-UTR sequence showed only 80% nucleotide homology with the rat gene, and this was limited to the last 70 bp preceding ATG (Table 2). A similar pattern of 5′-UTR splicing of p57Kip2 was previously reported29 and two splice isoforms are described for 5′-UTR of mouse p57Kip2.1
Interspecies Comparison of Chromosomal Organization of Cdkn1c Orthologs
Alignment of the coding sequences and 3′-UTR of rat p57Kip2 demonstrated 93–98% homology with the mouse ortholog and 77–82% homology with the human gene. Alignment of the different 5′-UTR splice variants also resulted in high homology with the corresponding variants (Table 2).
The exon–intron boundaries of the assigned genomic sequence conform to the conserved GT/AG rule. A high degree of interspecies similarity of the introns was also noted. All three species have unusual small and GC-rich introns of similar size with the exception of mouse intron 2, which is somewhat larger (Figure 1). The introns of rat p57Kip2 show 97% similarity to mouse introns. In contrast, the rat intron 1 is only 77% similar to human intron 1. Intron 2 has only partial similarity with human intron 2, and this similarity is limited to the 5′ and 3′ ends. The middle portion of human intron 2 and all of intron 3 show no resemblance between the species (data not shown).
The p57Kip2 phylogenetic tree, which represents the evolutionary/genealogical relationships between molecular sequences, gives us a clear picture of the evolutionary fork and demonstrates that rodent p57Kip2 is distinct from human p57Kip2 despite the presence of the conserved N- and C-terminal amino-acid domains (Figure 3 and below).
The analysis of the NCBI database indicates that p57Kip2 is located on the long arm of rat chromosome 1q42 in the supercontig WGS (∼1MB), BAC NW043404.1. This chromosome region is syntenic to mouse chromosome region 7F4 and to human chromosome 11p15.5 (Figure 4). Human H19, insulin (INS), and insulin-like growth factor 2 (IGF2) genes are located on the short chromosome arm, centromeric to CDKN1C in the same chromosome band, 15 Mb telomeric to the β-cell KATP channel genes ABCC8 and KCNJ11. As seen in Figure 4, in the rat, the gene cluster is on the long arm, in the same order relative to the centromere. A similar order is also seen in the mouse; however the distance between the two gene clusters is much larger in the rodent (107 and 95Mb, respectively), when compared to the human (15 Mb).
Schematic comparison of chromosomal location of the imprinted genes CDKN1C (p57Kip2), H19, IGF2 as well as β-cell genes INS, ABCC8 (SUR1) and KCNJ11(Kir6.2) in the human, rat and mouse. The centromere of the human and rat chromosome is indicated by a shaded oval. The distance between the two gene clusters is given in megabases (Mb).
Interspecies Comparison of p57Kip2 Proteins
The predicted short isoform of the rat p57Kip2 protein contains 343 amino acids, compared to 335 in the mouse and 316 in the human. An analysis of the protein structure of rodent and human p57Kip2 orthologs reveals marked differences in structural domains (Figures 5 and 6). Rat and mouse p57Kip2 both consist of four structurally distinct domains: an amino-terminal Cdk inhibitory domain that contains the cyclin and the Cdk-binding sites and is the characteristic for all CDKIs, a proline-rich domain, an acidic-repeat domain, and a carboxy-terminal domain conserved with p27Kip1 (QT domain).1, 2 Most of the acidic domain consists of a tandem repeat of a 4-amino-acid motif in which every fourth amino acid is glutamic or aspartic acid. Human p57Kip2 appears to have conserved amino- and carboxy-terminal domains but has replaced the internal region with sequences containing proline–alanine (PAPA) repeats4 (Figure 6b). The rat and mouse proteins show 87.7% identity and 89.2% homology, whereas the rat and human show only 56% identity and 62.6% homology (Table 3).
(a) Amino-acid sequence alignment of rat and mouse p57Kip2sh proteins. The domains are denoted above the sequence, whereas specific consensus sites are denoted under the sequence.  , Cdk inhibitory domain,
, Cdk inhibitory domain,  , proline-rich domain;
, proline-rich domain;  , acidic repeat;
, acidic repeat;  , QT domain;
, QT domain;  , MAP kinase consensus phosphorylation site (PATP);
, MAP kinase consensus phosphorylation site (PATP);  , nuclear localization site (KRKR);
, nuclear localization site (KRKR);  , CDK consensus phosphorylation site (TPRK). (b) Amino-acid sequence alignment of rat p57Kip2sh and human p57Kip2 proteins. The location of the nuclear localization site (KRKR) and the CDK consensus phosphorylation site (TPRK) are indicated under the sequence (
, CDK consensus phosphorylation site (TPRK). (b) Amino-acid sequence alignment of rat p57Kip2sh and human p57Kip2 proteins. The location of the nuclear localization site (KRKR) and the CDK consensus phosphorylation site (TPRK) are indicated under the sequence ( and
and  , respectively).
, respectively).
The 12 amino-acid insertion that differentiates between the short and long isoforms of rat and mouse p57Kip2 is located in the QT domain, which is likely to be important in protein–protein interactions30 and contains putative CDK phosphorylation and nuclear localization sites (Figures 5 and 6a). In the rodent, p57Kip2 Cdk inhibitory domains are completely identical but there are sequence differences in the proline-rich and acidic-repeat domains. Also there is the deletion (absence) of five residues in the rat QT domain when compared to mouse Cdkn1c.
Expression Pattern of Rat p57Kip2 Splice Variants
Specific primers were designed to determine the expression pattern of the two p57Kip2 splice variants that alter the expressed protein: (a) ratkipshF (1170–1188 bp of p57sh) and 1149R and (b) ratkiplongF and 1149R (Table 1). RatkipshF primer spans nucleotides before and after donor splice site (lower case and upper case letters, respectively, Table 1), and ratkiplongF spans the part of sequence that is absent in p57Kip2sh (upper case letters).
Figure 7 shows the representative pattern of p57sh and p57long expression in different rat organs. Although the expression of both short and long p57Kip2 transcripts is higher in the newborn rat pups than in the 1-month-old animal, the total expression of p57Kip2 appears to be low, since many more PCR cycles were required for optimal amplification of p57Kip2 (32 cycles) compared to actin (21 cycles).
Tissue Expression of p57Kip2 during Development
As a result of the putative role of p57Kip2 in the control of cell proliferation, we determined the tissue expression of the protein in various organs during different stages of development (Figure 9). At embryonic age 13.5 days, heart and liver tissue could be clearly demonstrated. Skeletal muscle was identifiable at 15.5 days. All other organs discussed here were examined from 17.5 embryonic days.
Hepatocytes first expressed p57Kip2 at day E19.5. Maximal expression was demonstrated at day 21.5, whereas low expression was demonstrated thereafter. In the heart, both myocardiocytes and endocardiocytes expressed p57Kip2 at high levels at the age of E13.5 days. Thereafter, expression rapidly decreased until only minimal expression was demonstrable at birth. High levels of p57Kip2 expression was demonstrated in striated muscle until the age of E21.5 days, after which expression declined and almost disappeared in the older age groups. A similar pattern of p57Kip2 expression was demonstrated in smooth muscle (data not shown).
In the kidney, medullary interstitial cells expressed p57Kip2 throughout all stages of rat development, with strongest staining demonstrated at ages E17.5 and E19.5. No expression was demonstrated in the tubular epithelium at any point. In the renal cortex, no expression was detected in the early glomerular structures including S-shaped bodies, vesicles and comma-shaped bodies. Epithelial cells in the inner part of the pronephron expressed high levels of p57Kip2 at ages E17.5 and 19.5. After birth, p57Kip2 was expressed in some but not all glomerular podocytes. This expression persisted into adulthood.
High p57Kip2 expression was demonstrated in the interstitial cells of the testis at the age of E17.5. This declined at ages E19.5 and E21.5 but was still detected in the adult rat. No expression was demonstrated in the seminiferous tubules until birth. At 1 week after birth, spermatogonia demonstrated very high expression of p57Kip2, which persisted through the age of 2 weeks, and then subsequently declined. Spermatides clearly demonstrated strong staining for p57Kip2 in the adult testes.
Cells lining the lumen of the major bronchi stained positive for p57Kip2 throughout rat development with strongest expression in days E17.5 and E19.5. Alveolar cells were negative during all stages examined (data not shown). P57Kip2 was expressed at the tips of villi of the small intestine at day E19.5. No immunoreactivity was demonstrated in the large intestine (data not shown). This pattern persisted into adulthood.
A unique and unexpected expression pattern was seen in the pancreas. During the early stages of pancreatic development (E17.5), the fibrous stroma highly expressed p57Kip2 protein. This stromal expression decreased progressively during development and was essentially absent after birth. In contrast, during fetal life there was no expression in the exocrine or endocrine parenchyma of the pancreas. At 1 week after birth, p57Kip2 was visualized in low levels in the exocrine tissue. Expression increased at the age of 2 weeks and remained elevated into adulthood. All cell types in the Islets of Langerhans, including the β-cells, were negative for p57Kip2 expression throughout rat development. To confirm this lack of p57Kip2 immunostaining in rodent pancreatic endocrine cells, we used semiquantitative RT-PCR to amplify RNA extracted from rodent β-cell-derived cell lines and freshly isolated rat islets. In mouse cell lines (MIN6 & βTC), the p57Kip2 expression was low even after 34 PCR cycles (data not shown), whereas in rat INS-1E, it was entirely absent. Since high confluence cell culture conditions may promote CDKN1C expression,31, 32 we cultured INS-1E cells to 85% confluence. However, this manipulation did not influence p57Kip2 expression (Figure 8). Similarly, in freshly isolated rat islets, p57Kip2 expression is also very low (data not shown).
Discussion
In this study, we describe the sequence and expression profile of the rat ortholog of Cdkn1c, the gene coding for p57Kip2, a tight-binding inhibitor of CDK complexes and therefore a negative regulator of the cell cycle.2 Our previous studies suggested that p57Kip2 may be an important regulator of human β-cell proliferation.26 Surprisingly, in this study, we demonstrate that this gene is not expressed in rat β-cells during any stage of development, a finding that is in sharp contrast to our previous observations in man.
The coding sequence of rat p57Kip2 is about 95 and 90% identical to that of the mouse at the DNA and the protein level, respectively. Computer analysis of the rat p57Kip2 suggests that its domain structure is very similar to that of the mouse, consisting of four structurally distinct domains: an amino-terminal Cdk inhibitory domain, a proline-rich domain, an acidic-repeat region, and a carboxy-terminal domain conserved with p27Kip1 that contains glutamine and threonine (QT domain).1, 2 In contrast, the homology with the human ortholog is much lower, only about 80% at the nucleotide level, and about 50% at the protein level. This low value is caused primarily by a marked difference in the structure of the mid-portion of the human protein. The biological significance of highly polymorphic PAPA domain in the middle of human CDKN1C or the proline-rich and acidic domains of rodent Cdkn1c is not clear. The polymorphic deletion of four amino acids in the PAPA repeat domain reduced the ability of p57Kip2 to inhibit growth, suggesting that this domain may have functional importance.29
Phylogenetic tree analysis of DNA sequences shows that rodent p57Kip2 is evolutionary quite distant from human p57Kip2 despite the presence of the conserved N- and C-terminal amino-acid domains. This gene divergence may be the result of repeat elements within the gene itself that are likely to be involved in gene evolution from a remote ancestor. Human, rat, and mouse CDKN1C genes are positioned at the terminal parts of the appropriate chromosomes, regions thought to have increased levels of recombination making them prone to rearrangements.33
5′-UTR of p57Kip2 from all three species shows the same complex pattern, consisting of three different splice variants, A1–A3. Furthermore, rat p57Kip2, like the mouse and human orthologs, has at least two different in-frame start sites for their open reading frames. These structural similarities are unexpected, given the relatively low nucleotide similarity in this region of the human gene when compared to the rodent orthologs (see Table 2). The functional significance of these 5′ splice variants is not known. In the human, the A3 variant, which utilizes a secondary translation start site, appears to be the primary transcript in the tissues examined.29 However, the relative abundance of the different transcripts requires further study, since this may vary in different tissues and at different times in development.
Both rat and mouse have alternative splice variants that result in the exclusion of 12 amino acids in the QT domain. The mouse splice variant was not previously reported, but is evident from alignment studies of published mouse EST sequences. A similar splice variant has not been described in the human, although to the best of our knowledge, it has not been specifically looked for. The functional significance of this variant is not known. We performed semiquantitative RT-PCR to determine if the relative expression of the two variants differed in different tissues and at different times in development. Our findings show that total p57Kip2 expression is somewhat higher in fetal compared to adult tissue with an apparent relative abundance of the short isoform in fetal tissue, a finding that requires further confirmation.
Rat p57Kip2 is situated on the distal portion of the long (q) arm of chromosome 1. This region contains closely linked imprinted genes, including maternally expressed H19 and paternally expressed IGF234 and is syntenic to regions of human chromosome 11p and mouse chromosome 7. Like its mouse and human ortholog, rat p57Kip2 has several features commonly seen in imprinted genes, such as small introns, high GC content and the presence of direct repeats.35 Interestingly, although the nucleotide homology between human and rat introns is low, their size and GC content are remarkably similar.
In the adult human and mouse, p57Kip2 is expressed in the heart, brain, lung, skeletal muscle, kidney and pancreas (human). The highest levels are seen in the placenta, whereas the lowest levels are seen in the liver.2 However, most of these tissues are highly heterogeneous and previous studies relying on RT-PCR and Northern blotting failed to provide any information as to which cell types in a given tissue express the gene. Furthermore, the transcriptional activity may not accurately reflect protein levels.36 For these reasons, we used immunostaining to determine specific cell-type expression in different tissues throughout rat development.
We demonstrate a unique temporal and cell-specific expression of p57Kip2 in various stages of rat development. The extent of its expression in the different tissues during embryonic life suggests that p57Kip2 may play an important role in rat morphogenesis. These findings are consistent with the fact that most of the p57Kip2 knockout mice died shortly after birth.17, 18, 19 In general, the expression pattern of rat and mouse p57Kip2 appears to be quite similar and, with the exception of pancreatic β-cells, the expression profile of rodent p57Kip2 appears to be similar to that previously reported in man.
In hepatocytes, p57Kip2 expression is limited to a relatively short time-window, peaking at E21.5 and then declined rapidly, as previously reported.36, 37 Similarly, both myocardium and endocardial cells expressed high levels of p57Kip2 at E13.5, decreasing progressively to low levels at birth. These findings are partially consistent to those previously reported in rat,38 which showed no expression in cardiac tissue after E17. This difference may be due to different methodologies used. In mouse, Kochilas et al39 found an expression pattern that was similar to that reported here, whereas others37, 40 described expression restricted to the endocardial cells of mouse at comparative embryonic stages. This discrepancy can be attributed to observational differences. As described in the liver, the expression in the fetal heart is limited to a relatively short period, suggesting that other cell cycle controlling genes are involved in liver and heart development, and is consistent with the lack of anomalies in these organs in p57Kip2 deficient mice.17, 18, 19
The high expression rates of p57Kip2 demonstrated in skeletal muscle during early stages of rat development are similar to those reported in mouse.37, 40 However, here we demonstrate relatively high levels that persisted until birth. After birth, the levels of p57Kip2 decreased but were still expressed at low levels in adult skeletal muscle, consistent with previous findings in the adult mouse.2 The extensive expression of p57Kip2 in skeletal muscle cells during all stages of rat embryonic development suggests an important role in the development of this tissue. Mice lacking p57Kip2 exhibited defective closure of the abdominal wall muscles.18 Thus, p57Kip2 may be involved in cell migration during development. A mechanism involving translocation of LIMK-1 from the cytoplasm into the nucleus, which in turn results in a reorganization of actin fiber, was suggested by Yokoo et al.41
In the kidney, interstitial cells in the medulla and epithelial cells of the glomerulus expressed p57Kip2 during all stages of rat development. These findings are similar to findings reported during mouse development.37, 42 In the adult kidney, persistent p57Kip2 expression was demonstrated in interstitial cells and podocytes, findings similar to previously reported data in human, rat and mouse.43 P57Kip2-deficient mice exhibited renal medullary dysplasia, with an increased number of stromal cells and a decrease in the amount of tubular cells in the medulla.18 A similar defect was also reported in BWS. Increased stromal cell proliferation may be due to the lack of p57Kip2 induced cell cycle arrest in this tissue.44 The lack of anomalies found in the renal cortex of p57Kip2-deficient mice and patients with BWS is surprising. P27Kip1, which is known to be expressed in glomerular podocytes,43 may compensate for the lack of p57Kip2 expression.45
In the testis, the cessation of p57Kip2 expression in spermatogonia at the age of 7 weeks coincides with sexual maturity when these cells begin to proliferate to produce sperm. As demonstrated in Figure 9, spermatides, which are visible in the lumen of 7-week-old rat semineferous tubules stain positive for p57Kip2, supporting the hypothesis that the gene is expressed in differentiated cells. Adult p57Kip2 null mice exhibited immaturity of the testis, prostate and seminal vesicles, but nevertheless were fertile.19 It is possible that the lack of p57Kip2 expression in interstitial cells results in delayed testosterone production and thus delayed sexual maturity. Most interstitial cells, and spermatogonia in the adult testis, do not express p57Kip2, suggesting that this protein does not play an important role in the process of spermatogenesis per se.
Our findings in the rat lung are consistent with the published data in the mouse40 and hamster,46 in that p57Kip2 staining was confined to columnar cells of major and segmental bronchi. However, in the mouse, Westbury et al described gradual decay in p57Kip2 expression after day E13.5, while in the hamster, Ikoma et al demonstrated weak expression in bronchi on day E14.5, which disappeared afterwards. In contrast, we demonstrated strong p57Kip2 expression in E17.5 and E19.5, which decreased to low levels but was still demonstrable in older ages. Mouse, hamster and rat alveolar cells do not express p57Kip2 during any stage of development.
In the intestines, cells at the tips of the villi stained positive for p57Kip2. This finding is similar to that reported in the mouse and further supports the hypothesis that p57Kip2 is expressed in terminally differentiated cells.37
We have previously reported a unique p57Kip2 expression pattern in the human pancreas.26 Cells of the fibrous stroma expressed p57Kip2 during embryogenesis in both the human and rat. In man, exocrine tissue never expresses p57Kip2, whereas β-cells demonstrated continuous expression of p57Kip2 throughout development and into adulthood. In sharp contrast, we now demonstrate that, in the rat pancreas, p57Kip2 is expressed in exocrine tissue and no staining at all was detected in the endocrine portion during any stage of development. These findings are confirmed using semiquantitative RT-PCR in that we demonstrated essentially no p57Kip2 expression in any of the β-cell lines studied or in isolated rat islets. Nagahama et al reported that p57Kip2 is expressed in the pancreas during a limited period of mouse embryogenesis,37 however, they did not differentiate between cell populations. BWS patients exhibit β-cell hyperplasia and hypoglycemia,47 while in p57Kip2 knockout mice, neither hypoglycemia nor β-cell hyperplasia was reported. This species difference may be explained by the lack of p57Kip2 expression in rodent β-cells.
The mechanism by which tissue expression of p57Kip2 is regulated during development is as yet unknown. Park and Chung48 determined that decrease of p57Kip2 message level in skeletal muscle and lung during mouse aging involved neither a change of promoter methylation nor an alteration of the imprinting status in aged mice.
In conclusion, we demonstrated unique temporal and spatial expression of p57Kip2 during rat development. In the pancreas, the expression profile of this protein is markedly different in rodents when compared to man. This is in contrast to other tissues for which human data are available, where the expression profile appears similar in the different species. The pattern of p57Kip2 expression in human pancreas, and its loss in patients with BWS and with the focal form of HI, suggests a crucial role for the gene in controlling β-cell turnover in man. Other mechanisms must compensate for the lack of this protein in rodent β-cells.
References
Lee MH, Reynisdottir I, Massague J . Cloning of p57KIP2, a cyclin-dependent kinase inhibitor with unique domain structure and tissue distribution. Genes Dev 1995;9:639–649.
Matsuoka S, Edwards MC, Bai C, et al. p57KIP2, a structurally distinct member of the p21CIP1 Cdk inhibitor family, is a candidate tumor suppressor gene. Genes Dev 1995;9:650–662.
Hatada I, Mukai T . Genomic imprinting of p57KIP2, a cyclin-dependent kinase inhibitor, in mouse. Nat Genet 1995;11:204–206.
Matsuoka S, Thompson JS, Edwards MC, et al. Imprinting of the gene encoding a human cyclin-dependent kinase inhibitor, p57KIP2, on chromosome 11p15. Proc Natl Acad Sci USA 1996;93:3026–3030.
Taniguchi T, Okamoto K, Reeve AE . Human p57(KIP2) defines a new imprinted domain on chromosome 11p but is not a tumour suppressor gene in Wilms tumour. Oncogene 1997;14:1201–1206.
Engel JR, Smallwood A, Harper A, et al. Epigenotype–phenotype correlations in Beckwith–Wiedemann syndrome. J Med Genet 2000;37:921–926.
Hatada I, Mukai T . Genomic imprinting and Beckwith–Wiedemann syndrome. Histol Histopathol 2000;15:309–312.
Maher ER, Reik W . Beckwith–Wiedemann syndrome: imprinting in clusters revisited. J Clin Invest 2000;105:247–252.
Algar EM, Deeble GJ, Smith PJ . CDKN1C expression in Beckwith–Wiedemann syndrome patients with allele imbalance. J Med Genet 1999;36:524–531.
Lee MP, DeBaun M, Randhawa G, et al. Low frequency of p57KIP2 mutation in Beckwith–Wiedemann syndrome. Am J Hum Genet 1997;61:304–309.
O’Keefe D, Dao D, Zhao L, et al. Coding mutations in p57KIP2 are present in some cases of Beckwith–Wiedemann syndrome but are rare or absent in Wilms tumors. Am J Hum Genet 1997;61:295–303.
Bhuiyan ZA, Yatsuki H, Sasaguri T, et al. Functional analysis of the p57KIP2 gene mutation in Beckwith–Wiedemann syndrome. Hum Genet 1999;104:205–210.
Hatada I, Inazawa J, Abe T, et al. Genomic imprinting of human p57KIP2 and its reduced expression in Wilms’ tumors. Hum Mol Genet 1996;5:783–788.
Bourcigaux N, Gaston V, Logie A, et al. High expression of cyclin E and G1 CDK and loss of function of p57KIP2 are involved in proliferation of malignant sporadic adrenocortical tumors. J Clin Endocrinol Metab 2000;85:322–330.
Oya M, Schulz WA . Decreased expression of p57(KIP2)mRNA in human bladder cancer. Br J Cancer 2000;83:626–631.
Tsugu A, Sakai K, Dirks PB, et al. Expression of p57(KIP2) potently blocks the growth of human astrocytomas and induces cell senescence. Am J Pathol 2000;157:919–932.
Yan Y, Frisen J, Lee MH, et al. Ablation of the CDK inhibitor p57Kip2 results in increased apoptosis and delayed differentiation during mouse development. Genes Dev 1997;11:973–983.
Zhang P, Liegeois NJ, Wong C, et al. Altered cell differentiation and proliferation in mice lacking p57KIP2 indicates a role in Beckwith–Wiedemann syndrome. Nature 1997;387:151–158.
Takahashi K, Nakayama K . Mice lacking a CDK inhibitor, p57Kip2, exhibit skeletal abnormalities and growth retardation. J Biochem (Tokyo) 2000;127:73–83.
Glaser B . Hyperinsulinism of the newborn. Semin Perinatol 2000;24:150–163.
Dunne MJ, Cosgrove KE, Shepherd RM, et al. Hyperinsulinism in infancy: from basic science to clinical disease. Physiol Rev 2004;84:239–275.
Verkarre V, Fournet JC, de Lonlay P, et al. Paternal mutation of the sulfonylurea receptor (SUR1) gene and maternal loss of 11p15 imprinted genes lead to persistent hyperinsulinism in focal adenomatous hyperplasia. J Clin Invest 1998;102:1286–1291.
Glaser B, Ryan F, Donath M, et al. Hyperinsulinism caused by paternal-specific inheritance of a recessive mutation in the sulfonylurea-receptor gene. Diabetes 1999;48:1652–1657.
Ryan FD, Devaney D, Joyce C, et al. Hyperinsulinism: the molecular aetiology of focal disease. Arch Dis Child 1998;79:445–447.
Kassem SA, Ariel I, Thornton PS, et al. Beta-cell proliferation and apoptosis in the developing normal human pancreas and in hyperinsulinism of infancy. Diabetes 2000;49:1325–1333.
Kassem SA, Ariel I, Thornton PS, et al. p57(KIP2) expression in normal islet cells and in hyperinsulinism of infancy. Diabetes 2001;50:2763–2769.
Cattoretti G, Becker MH, Key G, et al. Monoclonal antibodies against recombinant parts of the Ki-67 antigen (MIB 1 and MIB 3) detect proliferating cells in microwave-processed formalin-fixed paraffin sections. J Pathol 1992;168:357–363.
Kozak M . Adherence to the first-AUG rule when a second AUG codon follows closely upon the first. Proc Natl Acad Sci USA 1995;92:2662–2666.
Tokino T, Urano T, Furuhata T, et al. Characterization of the human p57KIP2 gene: alternative splicing, insertion/deletion polymorphisms in VNTR sequences in the coding region, and mutational analysis. Hum Genet 1996;97:625–631.
Reynaud EG, Leibovitch MP, Tintignac LA, et al. Stabilization of MyoD by direct binding to p57(Kip2). J Biol Chem 2000;275:18767–18776.
Blakely EA, Bjornstad KA, Chang PY, et al. Growth and differentiation of human lens epithelial cells in vitro on matrix. Invest Ophthalmol Vis Sci 2000;41:3898–3907.
Deschenes C, Vezina A, Beaulieu JF, et al. Role of p27(Kip1) in human intestinal cell differentiation. Gastroenterology 2001;120:423–438.
Mah N, Stoehr H, Schulz HL, et al. Identification of a novel retina-specific gene located in a subtelomeric region with polymorphic distribution among multiple human chromosomes. Biochim Biophys Acta 2001;1522:167–174.
Overall M, Bakker M, Spencer J, et al. Genomic imprinting in the rat: linkage of Igf2 and H19 genes and opposite parental allele-specific expression during embryogenesis. Genomics 1997;45:416–420.
Morisaki H, Hatada I, Morisaki T, et al. A novel gene, ITM, located between p57KIP2 and IPL, is imprinted in mice. DNA Res 1998;5:235–240.
Awad MM, Sanders JA, Gruppuso PA . A potential role for p15(Ink4b) and p57(Kip2) in liver development. FEBS Lett 2000;483:160–164.
Nagahama H, Hatakeyama S, Nakayama K, et al. Spatial and temporal expression patterns of the cyclin-dependent kinase (CDK) inhibitors p27Kip1 and p57Kip2 during mouse development. Anat Embryol (Berl) 2001;203:77–87.
Burton PB, Yacoub MH, Barton PJ . Cyclin-dependent kinase inhibitor expression in human heart failure. A comparison with fetal development. Eur Heart J 1999;20:604–611.
Kochilas LK, Li J, Jin F, et al. p57Kip2 expression is enhanced during mid-cardiac murine development and is restricted to trabecular myocardium. Pediatr Res 1999;45:635–642.
Westbury J, Watkins M, Ferguson-Smith AC, et al. Dynamic temporal and spatial regulation of the cdk inhibitor p57(kip2) during embryo morphogenesis. Mech Dev 2001;109:83–89.
Yokoo T, Toyoshima H, Miura M, et al. p57Kip2 regulates actin dynamics by binding and translocating LIM-kinase 1 to the nucleus. J Biol Chem 2003;278:52919–52923.
Hiromura K, Haseley LA, Zhang P, et al. Podocyte expression of the CDK-inhibitor p57 during development and disease. Kidney Int 2001;60:2235–2246.
Shankland SJ, Eitner F, Hudkins KL, et al. Differential expression of cyclin-dependent kinase inhibitors in human glomerular disease: role in podocyte proliferation and maturation. Kidney Int 2000;58:674–683.
Lee MH, Yang HY . Negative regulators of cyclin-dependent kinases and their roles in cancers. Cell Mol Life Sci 2001;58:1907–1922.
Zhang P, Wong C, DePinho RA, et al. Cooperation between the Cdk inhibitors p27(KIP1) and p57(KIP2) in the control of tissue growth and development. Genes Dev 1998;12:3162–3167.
Ikoma T, Ito T, Okudela K, et al. Modulation of the expression of the Cip/Kip family of cyclin-dependent kinase inhibitors in foetal developing lungs of hamsters. Cell Prolif 2001;34:233–241.
Stefan Y, Bordi C, Grasso S, et al. Beckwith–Wiedemann syndrome: a quantitative, immunohistochemical study of pancreatic islet cell populations. Diabetologia 1985;28:914–919.
Park CW, Chung JH . Age-dependent changes of p57(Kip2) and p21(Cip1/Waf1) expression in skeletal muscle and lung of mice. Biochim Biophys Acta 2001;1520:163–168.
Acknowledgements
The studies reported here were funded by an EFSD/JDFR/Novo Nordisk Focused Research Grant, a grant from the Horowitz Foundation and grant #601/01 from the Israel Science Foundation. This study is related to a research program on the pathophysiology and treatment of Hyperinsulinism of Infancy, which is supported by an EU-funded Concerted Action Grant (QLG1-2000-00513). TP is a recipient of a Kamea fellowship from the Israeli Ministry of Science and Ministry of Absorption. EH is a recipient of Shapiro fellowship from the Israeli Ministry of Absorption. The authors would like to thank Dr Sara Zangen for providing the fetal tissue used in this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Duality of Interest
None declared.
Rights and permissions
About this article
Cite this article
Potikha, T., Kassem, S., Haber, E. et al. p57Kip2 (cdkn1c): sequence, splice variants and unique temporal and spatial expression pattern in the rat pancreas. Lab Invest 85, 364–375 (2005). https://doi.org/10.1038/labinvest.3700229
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700229
Keywords
This article is cited by
-
EGR1 decreases the malignancy of human non-small cell lung carcinoma by regulating KRT18 expression
Scientific Reports (2014)