Abstract
To investigate the effects of Rho GTPase inactivation on lens fiber cell cytoskeletal and morphological integrity, a transgenic mouse model expressing C3-exoenzyme (a bacterial toxin) in a lens-specific manner was utilized. Cryosections of whole eyes from C3 transgenic mice and littermate controls were stained for F-actin with rhodamine–phalloidin or immunostained for β-catenin, aquaporin-0 or connexin-50, and confocal images were recorded. Lens fiber cell morphology was examined at both light and electron microscopic levels. To investigate the influence of Rho GTPase inactivation on the profiles of gene expression, cDNA libraries generated from transgenic and littermate control mouse lenses were screened by cDNA microarray analysis. In contrast to the wild-type lens, fiber cells of the transgenic lens were grossly swollen and disorganized, with abnormal membrane architecture. Staining of F-actin, β-catenin, aquaporin-0 and connexin-50 was reduced dramatically in the C3 transgenic lens as compared to controls. Western blot analysis and cDNA microarray analysis did not reveal any noticeable decreases in actin, β-catenin and aquaporin-0 protein levels or expression in C3 transgenic lenses, indicating that altered cytoskeletal organization in response to Rho GTPase inactivation might underlie the noted changes in staining for these proteins. Additionally, cDNA microarray analysis of C3 lens revealed altered expression (at least two-fold, compared to littermate controls) of 44 genes. These include genes encoding extracellular matrix and basement membrane proteins, cell survival and apoptotic pathways, and ion and protein transport. These data indicate that disruption of Rho GTPase function in the developing mouse lens results in abnormal cytoskeletal organization, fiber cell interactions, impaired lens fiber cell morphology and altered gene expression of cellular proteins involved in diverse functions. This work reveals that the morphological and cytoskeletal abnormalities triggered upon Rho GTPase inactivation in lens could be one of the important insults associated with cataract formation in C3 transgenic mouse lens.
Similar content being viewed by others
Main
Members of the Rho subfamily of Ras-related GTP-binding proteins act as molecular switches regulating the organization of the actin cytoskeleton, cell adhesion, transcription, cell cycle progression, apoptosis and membrane trafficking.1, 2, 3, 4 The activity of these GTPases is regulated by signals originating from different classes of cell surface receptors including G-protein-coupled receptors, tyrosine kinase receptors, cytokine receptors and adhesion receptors.1, 2, 3, 4 The actin cytoskeleton is considered a primary target of the small GTP-binding protein Rho GTPase, which through its downstream effectors Rho-kinase and LIM kinase, regulates the formation of actin stress fibers, focal adhesions and adherens junctions via a molecular mechanism involving myosin light-chain phosphorylation and cellular contraction.1, 2, 3, 4, 5 Rho GTPases also regulate the assembly of integrin-containing signaling complexes at the plasma membrane, which generate adhesion-dependent intracellular signals essential for cell growth, differentiation and survival.3, 6 Additionally, they activate signaling pathways that regulate transcription, cell cycle progression, apoptosis and organogenesis.7, 8, 9, 10, 11, 12
The normal development and growth of the vertebrate lens is a complex process that requires the precise spatial and temporal control of epithelial cell proliferation and the subsequent differentiation of epithelial cells into fiber cells. Fiber cell differentiation is accompanied by cell elongation and remodeling of cytoskeletal structures.13, 14, 15, 16, 17, 18 Our knowledge of lens fiber cell elongation and cytoskeletal reorganization is limited, and the role of intracellular signaling pathways in these cellular processes remains to be elucidated. We have hypothesized that Rho GTPases (Rho and Rac) play a critical role in lens growth and function. Lens tissue expresses various Rho GTPases, and inhibition of their activity affects lens transparency, lens epithelial cell morphology and cytoskeletal organization.19, 20, 21 Various growth factors have been shown to activate Rho and Rac GTPases in lens epithelial cells.22 Recently, we have demonstrated the significance of Rho GTPase function in lens growth and function using a transgenic mouse model in which Rho GTPase function was impaired, in an organ-specific manner.23 This inactivation of Rho GTPase was achieved by utilizing the lens-specific αA-crystallin promoter to drive the expression of C3-exoenzyme in transgenic lenses. In the presence of NAD, C3-exoenzyme from Clostridium botulinum ADP-ribosylates and irreversibly inactivates the Rho GTPase A, B and C isoforms.24 Transgenic mice expressing C3-exoenzyme exhibit a severe ocular phenotype, including cataract and microphthalmia.23
To obtain further insights into the significance of Rho GTPase-mediated signaling on lens growth and structural integrity, we have performed histological and immunohistochemical studies on lenses from C3 transgenic and wild-type mice. These studies focus on the effects of Rho GTPase function on morphology, actin cytoskeletal organization, adherens junction formation, and the distribution of water channel and gap junction proteins in lens fiber cells. Further, changes in gene expression profiles were also evaluated to elucidate the significance of Rho GTPase activity in maintenance of lens growth and function.
Materials and methods
Mice
C3 transgenic mice with FVB/N genetic background were bred inhouse and used in this study.23 Transgene positive mice were screened by genomic analysis of tail DNA, using C3-exoenzyme-specific oligonucleotide-based PCR analysis as described previously.23 The transgenic mouse line used in this study was ODO4-11, in which only male animals are transgene positive, and age-matched littermate females were used as controls. Animal studies were conducted in accordance with the ARVO statement for the use of animals in ophthalmic and vision research under an approved animal protocol.
Histological Evaluation
Whole eyes from 7-day-old postnatal transgenic and littermate control mice were dissected and immersed in 2.5% glutaraldehyde in 50 mM cacodylate buffer (pH 7.2) containing 4% sucrose and 2 mM CaCl2 for 2 h, and were then transferred to 10% buffered formalin for 48 h. The specimens were then embedded in methyl methacrylate, and 2-μm sections were cut and stained with hematoxylin and eosin (H&E) or with PAS (periodic acid-Schiff) using conventional methods and micrographs were captured using Zeiss light microscope. For electron microscopy, lenses were fixed in modified Karnovsky's cacodylate buffer (0.1 M) containing 2.5% glutaraldehyde, 2% paraformaldehyde and 1% tannic acid at pH 7.2. Sections were cut with a vibrotome and ultramicrotome, and staining was performed as described previously.25 Specimens (70 nm thick sections) were examined using the FEI Philips Tecnai 12 TEM (FEI Company, Hillsboro, OR, USA) at 80 kV. Images were captured using the Gatan 794 digital camera and Digital Micrograph software.
Immunohistochemistry
Freshly obtained mouse tissues (whole heads or eyes) were fixed in 4% paraformaldehyde in phosphate-buffered saline (PBS) at 4°C for 18 h. Fixed tissues were then transferred sequentially into 5% and 30% sucrose in PBS, each at 4°C for 24 h. Cryostat sections (7 μm) of the whole head or whole eyes from both transgenic and littermate controls were cut and mounted on gelatin-coated glass slides and stored at −80°C until use. Blocking was performed by incubating the sections with 5% goat serum in PBS with 0.3% Triton X-100 (PBST), for 1 h at room temperature. Tissue sections were incubated overnight at 4°C with polyclonal antibodies raised against β-catenin (from Sigma-Aldrich), MIP-26 (a kind gift from Dr Sam Zigler) or Connexin-50 (from Alpha Diagnostics, San Antonio, TX, USA), at a dilution of 1:1000. These sections were washed three times with PBST with an interval of 15 min and subsequently incubated with fluorescein isothiocyanate (FITC)-conjugated goat anti-rabbit secondary antibody (from Sigma-Aldrich), for 2 h at room temperature. After thorough washing with PBST, sections were coverslipped and mounted using 50% glycerol, for viewing with a Zeiss LSM confocal fluorescence microscope. For actin staining, the preblocked sections were labeled with phalloidin conjugated with tetra rhodamine isothiocyanate (TRITC) (500 ng/ml; Sigma-Aldrich) for 2 h at room temperature; these sections were then washed thrice in PBST for 15 min each. To assess specificity of antibody reactivity, sections were labeled either with FITC-conjugated secondary antibody alone or with rhodamine alone.
RNA Extraction
Total RNA was extracted from five pairs of 7-day-old C3 transgenic or littermate control mouse lenses using the Qiagen RNeasy Mini kit (QIAGEN, Valencia, CA, USA), following the manufacturer's instructions. During the extraction, lens homogenates were passed through a 21-gauge needle several times to shear DNA, and the crude RNA was treated with DNase-1 to eliminate contamination from genomic DNA. Purified RNA was quantitated spectrophotometrically at 260/280 nm. Since several lenses needed to be pooled in order to obtain sufficient RNA for microarray analysis, lenses from 7-day-old animals were used. Furthermore, gender determination was found to be much easier in 7-day-old mice compared to younger animals, making it easier to pool lenses based on gender, as only males are transgene positive in the ODO4-11 line.
Microarray Analysis
Affymetrix gene chip-based transcript profiling was performed at the Duke University Microarray Core Facility by using MG-U74Av2 chips that contained immobilized probes for 12 488 genes (6000 genes from Unigene database and more than 6000 EST clusters), by following the manufacturer's protocol. Briefly, 8 μg of total RNA from wild type and C3 transgenic mouse lens was reverse transcribed using T7 RNA polymerase and an oligo (dT) primer, as per the manufacturer's instructions (Affymetrix). Biotinylated complementary RNA (cRNA) generated from cDNA was hybridized at 45°C for 16 h with the Affymetrix gene chips. The hybridized gene chips were incubated with phycoerythrin–streptavidin, and after several washings, the signal was further amplified by incubating the chips with antistreptavidin antibody coupled to phycoerythrin. Hybridization efficiency was quantified using a Gene chip scanner and scanned signals were processed using the Affymetrix microarray analysis suite, version 5.0 software. In such experiments, cRNA probes were first hybridized to an Affymetrix test chip to ensure the quality of the preparation.
Reverse Transcription-Polymerase Chain Reaction
To obtain further confirmation of differentially expressed genes identified by the microarray analysis, RNA (1 μg) derived from the different sets of C3 or wild-type lenses was reverse transcribed and amplified using the Advantage RT-PCR kit obtained from BD Bioscience Clonetech (Palo Alto, CA, USA). RNA samples were treated with DNase1 (GIBCO BRL, Gaithersburg, MD, USA) prior to performing RT reactions, and controls lacking RT were also set up to confirm the absence of genomic DNA-generated signals. The sequences of primers (forward and reverse) used and the expected product lengths for the different transcripts amplified in this study are listed below:
JNK-1: (GAAAACAGGCCTAAATACGCTGG/CCC-TCTTATGACTCCATTCTTAGT TCG; 341 bp), Collagen IV: (GCCACTGGTACAAGAATGCGA/CATTCTTTCTGGTTCAGCGAA; 611 bp), Integrin-β7: (GTGCCTGGATGGCTACTACGG/ATTTCCACAGAGAGCCGGTAAG; 378 bp), EDG-2: (GATGTTCAATACAGGACCTAATACCCG/GAACATAGCCAAAGATGTGAGCGTA; 382 bp) and ABCtransporter-1: (TCAGATTACCTGGTGAAGACGTACG/AGATAGACACGAGGACGTCGACA; 462 bp).
PCR amplifications were performed within the linear range of DNA product formation. The housekeeping gene glyceraldehyde 3-phosphate dehydrogenase (G3PDH) with the following primer pairs (ACCACAGTCCATGCCA TCAC/TCCAC CACCCTGTTGCTGTA; 452 bp) was amplified as an internal control for normalizing the DNA content of control and transgenic samples in Reverse transcription-polymerase chain reaction (RT-PCR) reactions.
Western Blot Analysis
Whole lenses from both control and transgenic mice (7 days old) were dissected free of other tissue and homogenized in 20 mM Tris buffer, pH 7.4 containing 1 mM sodium orthovanadate, 0.2 mM EDTA, 0.2 mM PMSF, 0.1 M NaCl, 50 mM NaF, aprotinin (25 μg/ml), leupeptin (25 μg/ml) and centrifuged at 800 g for 10 min at 4°C. Protein concentration was estimated in 800 g supernatants by the Bradford method.26 Equal amounts of lens protein (60 μg) were dissolved in Tris buffer containing 5 M urea, 2 M thiourea and 2% CHAPS (3-[3-cholamidopropyl)dimethylammonio]-1-propanesulfonate and separated on sodium dodecyl sulfate (SDS)-polyacrylamide gels (12.5% acrylamide), followed by electrophoretic transfer to nitrocellulose membrane and electrochemiluminescence (ECL) detection using antibodies raised against actin (anti-smooth muscle actin, mouse monoclonal from Sigma, Saint Louis, MO, USA), paxillin (anti-paxillin, mouse monoclonal from sigma, Saint Louis, MO, USA), gelsolin (antigelsolin, mouse monoclonal from Transduction Laboratories, Lexington, KY, USA), β-catenin (anti-β-catenin, raised in rabbit from Sigma, Saint Louis, MO, USA) and aquaporin-0 (antiaquaporin-0, a kind gift from Dr Zigler, raised in rabbit).
Results
Histological Changes in C3 Transgenic Lens
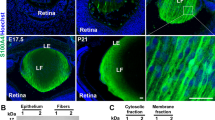
For comparison of lens integrity, lens sections obtained from 7-day-old C3 mice from line ODO4-11 and wild-type littermates were stained with H&E or PAS. As reported previously,23 severe morphological abnormalities were observed in the C3 transgenic lens, including large extracellular spaces in the cortical regions, thickened anterior lens capsule, and ruptured posterior capsule with degraded lens material (indicated by asterisk) leaking out into the vitreous chamber (Figure 1b). The epithelium in C3 lenses was intact, exhibiting no obvious morphological changes. However, unlike wild-type lens fiber cells, which are compact with an anterior to posterior ribbon-like fiber cell organization (Figure 1a), both primary and secondary fibers of C3 lenses appear shorter in length (Figure 1b) at the equatorial regions, displaying stunted growth and accumulation of large vacuoles, particularly in the anterior and equatorial regions of the lens. Further, the cortical region in C3 lenses displayed areas of definite PAS-positive staining, which is rather unusual, as it would indicate the presence of basement membrane material in the cortical region of the lens (indicated with an arrow in Figure 1b). The lens weight of C3 transgenic mice was 40–50% lower than that of controls, and many of the lens fiber cells in the differentiated nuclear area of C3 lenses retained nuclei (indicated with arrow heads in Figure 1b).
Histological changes in C3 transgenic mouse lenses. To evaluate histological integrity of the C3 transgenic lens, eyes from 7-day-old C3 transgenic (b) and littermate control mice (a) were fixed, embedded in methyl methacrylate and stained with PAS. As can be seen from the figure, severe morphological abnormalities were evident in the transgenic lens including deformed lens shape, thin and mostly ruptured posterior lens capsule, thickened anterior lens capsule, distorted fiber cell organization with evidence of degrading lens proteins (indicated by asterisk) leaking into the posterior chamber. Further, at the equatorial regions lens fiber cells in C3 mouse appear much shorter than in the wild-type lenses. Vacuole accumulation beneath the epithelium, and at the equatorial regions, presence of traces of basement membrane (indicated by arrow) in lens fiber mass and retention of nuclei (indicated by arrow heads) in differentiated lens fibers are some of the obvious histological changes found in C3 transgenic mouse lens.
EM Histological Changes
Although the epithelium of the C3 mouse lens appears normal under a light microscope, electron microscopy of C3 lens epithelium reveals large extracellular spaces and accumulation of large vacuoles (Figure 2, right panel) as compared to wild-type lens epithelium (left panel). The cytoplasm of C3 and wild-type lens epithelium, however, appear very similar. The lens fiber membrane is very abnormal with irregular cell borders (Figure 3b). Further, these abnormal fiber cells exhibit large vacuole-like structures, which may represent either degraded cell bodies or vacuoles (Figure 3b). Figures 3c and d depict the presence of F-actin (c) and a microtubule (d) in elongating fibers, indicating that these cytoskeletal proteins are organized in their native form in fibers from C3 lenses. The interaction of F-actin with other cytoskeletal proteins such as myosin, however, might not be normal in C3 transgenic mouse lens. The anterior capsule of C3 lenses is usually very thick compared to that from a control lens (Figure 1b). EM morphology of the C3 lens capsule, however, did not reveal any structural abnormality when compared with the control lens (Figure 2, right panel and indicated as c).
C3 transgenic mouse lens epithelial cell morphology. As compared to wild type (left panel), lens epithelial cells from C3 transgenic mice (right panel) exhibit large extracellular spaces (indicated with stars). Although the anterior lens capsule (C) is consistently thicker in C3 lenses, it does not exhibit any abnormal morphology compared to wild type. N indicates nucleus.
C3 transgenic mouse lens elongating fiber cell morphology. Fiber cells from the equatorial region of C3 lenses (panel b) exhibited abnormal morphology in size and shape with accumulation of large cell bodies along with abnormal cell borders compared to wild-type lenses (panel a). Panels c and d depict the presence of actin structure (bar is × 90 000) and microtubule (bar is × 45 000) in C3 lens shown with arrows, respectively. Arrows in panel a indicate the fiber cell membrane.
Lens Cytoskeletal Architecture in C3 Mouse
Staining of F-actin with rhodamine–phalloidin yielded a continuous staining pattern throughout the lens epithelium and fibers of wild-type mice, with intense staining at the interface of lens epithelium and anterior lens fibers and at the posterior ends of lens fibers where they attach to the lens capsule (Figure 4a, b). Actin staining was much reduced in lens fibers in both the cortical and nuclear regions of the C3 transgenic lens (Figure 4c,d), while the staining pattern of the C3 lens epithelium (Figure 4d) was comparable, to that observed in control lenses. Interestingly, in both C3 and wild-type lenses, actin staining was very intense at the interface of lens epithelium and anterior lens fibers and at the contacts of posterior lens fiber tips and capsule, indicating that actin organization at these regions is less influenced by transgene expression, as compared to that observed in cortical and nuclear regions from the day 1 lens. Whether this difference is due to differential expression and distribution of the C3 transgene is not clear. Fiber cells in the cortical and nuclear regions of C3 lens were found to be swollen, exhibiting a distorted morphology (indicated with arrows in Figure 4c, d). C3 fiber cells also stained very weakly for F-actin, with the staining very diffusely distributed in the lumen of the fiber cell body (Figure 4d).
Organization and distribution of rhodamine–phalloidin-stained F-actin in 1-day-old C3 transgenic mouse lens. To address the role of Rho GTPase activity in actin cytoskeletal architecture, wild-type and C3 lenses were labeled for filamentous actin (F-actin) with rhodamine–phalloidin. Confocal images of F-actin showed positive staining throughout the lens cross-section, including the epithelium, and the cortical and nuclear lens fibers. F-actin staining in wild-type lenses was very uniform along the fiber cells with intense staining at the anterior and posterior terminals, which interact with either the lens epithelium or lens capsule, respectively (Panels a and b). In contrast, actin staining in the C3 lens fibers was very weak, and the distribution appears very diffuse in the disorganized and swollen lens fibers of the C3 lens (indicated with arrows, panel d). Stars in panels a and c indicate the magnified area shown in panels b and d.
The immunostaining of β-catenin was found to be very strong in the epithelium and in the elongating lens fibers of equatorial and cortical regions in the wild-type lens (Figure 5a, b). In the epithelium, β-catenin staining was localized predominantly to the intercellular junctions (Figure 5b), with intense staining at the equatorial epithelial cells and the early differentiating fiber cells. β-Catenin staining was markedly reduced in the primary and secondary fibers (Figure 5c, d) of the C3 transgenic lens, particularly in regions where fiber cell organization was severely compromised (Figure 5d, indicated with arrows). Again as seen in Figures 4 and 5d, lens fibers appear swollen and shorter (indicated with arrows) in C3 lenses as compared to fiber cells in wild-type lenses. Lens sections obtained from the 7-day-old C3 mouse showed similar but more severe reductions in actin and β-catenin staining profiles (data not shown).
Distribution of β-catenin in 1-day-old C3 transgenic mouse lens. To study the effects of Rho GTPase inactivation on adherens junction distribution in C3 lens, cryosections obtained from wild-type and C3 transgenic lenses were labeled with anti-β-catenin and FITC-conjugated secondary antibody. β-Catenin is localized to intercellular junctions (adherens junctions) in the epithelium (indicated with arrow head) and is distributed both in epithelium and lens fibers of the cortex and nucleus (panels a and b) in the wild-type lens. In C3 lenses, staining for β-catenin was dramatically reduced, particularly in lens fibers that exhibit distorted and swollen morphology (indicated with arrow, panel d). The epithelium and those fiber cells in the C3 lens exhibiting a normal morphology, however, stained normally for β-catenin (indicated with arrow heads). Stars in panels a and c indicate the magnified area shown in panels b and d.
Distribution of Water Channel and Gap Junctions Proteins
As expected, the major intrinsic protein (MIP-26), a water channel-associated protein called aquaporin-0, was distributed primarily in lens fiber cells (Figure 6a–c). Fiber cells throughout the wild-type lens immunostained very strongly for aquaporin-0 (Figure 6a, b) with no staining in epithelium (indicated by arrow head). In contrast, the staining in C3 lens fibers (particularly fibers with abnormal morphology) was very weak and diffuse (Panel d indicated with arrows). Corneal epithelium from both wild-type and C3 transgenic eyes showed some nonspecific staining for aquaporin-0. Connexin-50, a major gap junction protein, known to be present in fibers, but not in lens epithelial cells displayed discontinuous punctate distribution in wild-type fiber cells (Figure 7a). C3-lenses revealed a dramatic reduction in staining for connexin-50, as depicted in Figure 7b. The 7-day-old lens also exhibited a similar effect of C3 transgene expression on the distribution of connexin-50. The staining pattern for connexin-50, however, was observed to be strong, punctate and continuous (data not shown) in the 7-day-old wild-type lens. Differentiating cortical lens fibers are known to express higher levels of connexins during lens maturation.27
Immunohistochemical localization of MIP 26/aquaporin-0 in C3 lens. To analyze the distribution of lens-specific water channel-associated protein in C3 lens, anti-MIP-26 polyclonal antibody was utilized to label frozen sections from 1-day-old lenses. Being a fiber-specific protein, MIP-26 showed positive staining only in the fiber cells of control (panels a and b) and C3 lenses (panels c and d) and immunostaining was detected along the fiber membranes (indicated with arrows in panel b) and all through the fiber cells (panels a and b). Reduced staining was noted in the swollen fiber cells of 1-day-old C3 lenses with diffused cytosolic distribution in distorted fiber cells (panels c and d, indicated with arrow). The arrowhead in panel b indicates lack of MIP 26 staining in lens epithelium. Mouse corneal epithelium showed nonspecific staining with anti-rabbit secondary antibody (panels a and c). Stars in panels a and c indicate the magnified area shown in panels b and d.
Immunolocalization of connexin-50 in C3 transgenic mouse lens. Connexin-50, a gap-junction protein was immunolocalized using polyclonal antibody in conjunction with FITC-conjugated secondary antibody. The distribution of this protein appears punctate and specific to only lens fibers in 1-day-old wild-type lens (indicated with arrows in panel a). In C3 lens, the staining of connexin-50 was also punctate but is dramatically reduced (panel b). As expected, the lens epithelium showed no staining for connexin-50 in either wild-type or C3 lenses (indicated with arrow head).
Levels of Cytoskeletal Proteins in C3 transgenic Lens
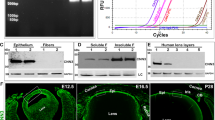
Western blot analysis of actin and actin-interacting proteins including actin, paxillin, gelsolin, β-catenin and aquaporin-0 in 7-day-old C3-transgenic and wild-type lenses revealed not much difference between the two groups. The levels of these proteins were not decreased in C3 lenses as compared to wild-type lenses (Figure 8); indeed there appeared to be a moderate increase in the levels of actin and beta-catenin in C3 transgenic lenses (Figure 8).
Cytoskeletal protein profile in C3 transgenic and control mouse lenses. Equal amounts of total lens protein (60 μg) obtained from 7-day-old C3 transgenic (right panel) and control lenses (left panel) were subjected to Western blot analysis to determine the effects of Rho GTPase inactivation on the levels of selected cytoskeletal interacting proteins in lens tissue.
Differential Expression of Genes in C3 transgenic Lens
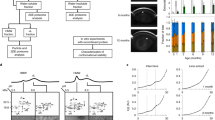
To identify the mechanism(s) accounting for the altered staining patterns of cytoskeletal, water channel and gap junction proteins, and to investigate the status of gene expression in general in C3 transgenic mouse lens, we analyzed the expression profiles of various genes in C3 lens. Towards this end, we performed a cDNA microarray analysis using a chip containing over 12 000 specific gene tags. Comparison of gene expression profiles of 7-day-old C3-transgenic mouse lens and littermate wild-type lens by the Affymetrix cDNA microarray analysis detected a total of 44 differentially expressed genes, which exhibited a two-fold minimum difference in expression between C3 and wild-type lenses. Of the 44 differentially expressed genes, four were ESTs and the remaining 40 encoded proteins of known function (from the Unigene databank). Among these 44 differentially expressed genes, 19 were downregulated, and 25 were upregulated in C3 lenses. Interestingly, many of the genes whose expression appeared to be downregulated in response to inactivation of Rho GTPase function in C3 lenses, encoded proteins involved in transport functions. These include a member of the solute carrier family 12 (U20973), potassium voltage-gated channel (AF032099), Ran GTP binding protein (AV043011), kinesin-like protein KIF3A (AV324729) and endoplasmic reticulum protein 29 (AV292321). AV379327 and AV358218, which were downregulated by 8- and 16-fold, respectively, in the C3 mouse lens, represent ESTs with no known function. This analysis did not reveal any significant changes in the expression profile of actin, β-catenin, connexins, aquaporins and other cytoskeletal genes.
Expression of a group of 25 genes encoding proteins, belonging to diverse functional clusters was increased in the C3 lens. This group includes CD44 (X66084), latent TGF-beta binding protein-3 (L40459), dermatopontin (AA717826), C1q-related factor (AF095155), integrin beta-7 (M68903) and collagen Type IV alpha-3 (Z35166), all of which are involved in extracellular matrix and basement membrane production and organization. Importin alpha3 (AF020772), ABC1-transporter (X75926) and Lin-7 (AF087695) are all proteins with role(s) in intracellular and nuclear transport. Interestingly, genes encoding proapoptotic response proteins, including JNK1 (AB005663), quaking protein-7 (AV347370) and granzyme F (J03257), were among those whose expression was strongly upregulated in C3 lenses. Lysophospholipid (LPA) receptor-edg2 (U13370), A kinase anchor protein-1 (U95145), proline–serine–threonine phosphatase interacting protein (AV 312229), which participate in intracellular signaling and cell survival, were also upregulated in C3-lenses. All the differentially expressed genes (both upregulated and downregulated) have been clustered based on their known function and listed in Table 1. These genes fall into one of four functional clusters: extracellular matrix and basement membrane related, transport and trafficking, cell survival and death, and protein degradation.
RT-PCR Analysis of Differentially Expressed Genes in C3 Lens
To obtain independent confirmation for the differential profiles of gene expression deduced from microarray analysis of cDNA from C3 lenses, semiquantitative RT-PCR was performed to analyze the expression of selected genes including JNK1, collagenase IV alpha3, LPA receptor, ABC1 transporter and integrin beta-7. The RNA used in this analysis was derived from samples independent from those used for microarray analysis. The housekeeping protein G3PDH was amplified as an internal control. All of the genes selected for RT-PCR analysis were those found to be upregulated by a minimum of two - fold by microarray analysis. Figure 9 depicts the expression profile of genes in C3 and control lenses analyzed by RT-PCR. As can be seen from the figure, expression levels of JNK1, collagen IV alpha3, LPA receptor (edg2), ABC1 transporter and integrin beta-7 were much higher in C3 lenses compared to their littermate controls, while expression levels of G3PDH showed no difference between C3 and control lenses. These RT-PCR results were confirmed in two independent analyses using RNA derived from the independent samples.
Semiquantitative RT-PCR analysis of selected genes in C3 transgenic lens. To obtain independent confirmation of the differential expression of genes in C3 lenses, identified by the cDNA microarray method (Table 1), semiquantitative RT-PCR amplification was performed for selected genes. G3PDH (a house keeping gene) was amplified as an internal control for normalizing the RNA template content in PCR amplification reactions, Reactions lacking reverse transcriptase (−RT) were included as controls. All the genes evaluated by RT-PCR confirmed the results obtained by microarray analysis, and were found to be upregulated in C3 lenses as compared to control lenses.
Discussion
Expression of the C3 ADP-ribosyl transferase gene in developing lenses of transgenic mice was found to result in disruption of fiber cell morphology, as indicated by the presence of swollen and shortened fiber cells, abnormal fiber cell borders and a ruptured lens capsule. Lenses from C3 transgenic mice also revealed altered profiles of gene expression, extensive changes in the organization of actin and β-catenin, and altered distribution of gap junctional and water channel proteins. Normal lens development is predicated upon a temporally and spatially controlled sequence of events involving interfiber interactions and fiber cell organization. Lens function is also critically dependent on intact intercellular-junctions (gap and adherens junctions), cell–extracellular matrix (ECM) interactions, and water channel function.13, 14, 15, 16, 17, 28, 29, 30 Given the extent of the morphological, cellular and biochemical deficits noted in the C3 lens, it is clearly evident that Rho GTPase functions as a critical determinant of normal lens growth, development and function.
Rho GTPases regulate a wide range of biological processes including actin cytoskeleton organization, cell–cell adhesion, cell–ECM interactions, intracellular trafficking and transcriptional regulation through various signaling mechanisms.1, 2, 3, 4, 5 Rho GTPase has been shown to have a critical role in organogenesis and cell survival as well.7, 8, 9, 10, 11, 12 The lenses of C3 mice evaluated in this study revealed striking abnormalities of fiber cell organization (Figure 3), with accumulation of large vacuoles inside and beneath the lens epithelium and at the equatorial regions. C3 mice consistently exhibit a ruptured posterior capsule with a very thick anterior capsule (Figure 1b). Furthermore, the cortical region in C3 lenses exhibited positive staining for the presence of basement membrane material. Deficits in immunostaining intensity, as well as altered distribution of immmunostaining were noted for both F-actin and β-catenin, an adherens junction-cadherin binding protein (Figures 4 and 5). Inhibition of Rho GTPase by C3 exoenzyme, and of Rho-kinase, the downstream effector of Rho GTPase, in various cell types including a human lens epithelial cell line, interferes with actin polymerization and stress fiber organization, formation of focal adhesions and adherens junctions, and leads to changes in cell morphology and cell death.1, 2, 3, 4, 20, 21, 22, 31 In C3 lens fibers, the staining of actin and β-catenin appeared rather diffuse in the fiber cell cytosol (Figures 4 and 5). In control lenses, these proteins are predominantly localized to the fiber cell membrane. To understand whether the reduced staining of actin and β-catenin observed in C3 lenses owes solely to abnormal organization of actin and disrupted adherens junction formation, or occurs in response to reduced levels of expression of genes encoding for cytoskeletal proteins, we compared the gene expression profiles of C3 and wild-type lenses by cDNA microarray analysis, and quantitated the levels of selected cytoskeletal protein by Western blot analysis. Neither the microarray results (Table 1) nor the Western blots (Figure 8) demonstrated a difference between C3 and wild-type lenses, suggesting that the reduced staining of actin and β-catenin most likely occur as a consequence of the impairments in cytoskeletal organization and/or the interaction of these proteins with other proteins of the cytoskeletal network. Further, the abnormal cytoskeletal organization observed in C3 lenses might also influence the distribution and function of water channels (such as aquaporin-0) and gap junctional proteins (connexin-50), which are important not only for maintaining normal fiber cell function and communication but also for lens transparency (Figures 6 and 7).28, 32, 33, 34 Unlike in wild-type lenses, aquaporin-0 localization in C3 lenses appears diffuse and spread throughout the fiber cell lumen (Figure 6d, indicated with arrows). It is also possible that the altered fiber cell morphology and abnormal cell borders that were observed in C3 lenses might potentially impair water channel and gap junction organization and/or their function. However, since Rho GTPase activity is thought to be a master switch controlling actin cytoskeletal organization and thereby cell–cell interactions, the morphological changes observed in C3 transgenic mouse lens fibers (Figure 3) may be the consequence of impaired cytosketal organization and cell adhesion.
Other genes contributing to the transport function were downregulated in the C3 lenses as evidenced by the microarray analysis including the solute carrier family 12 transporter (Na–K–Cl cotransporter NKCC2) which was downregulated 16-fold. This transporter is reported to be involved in transport of Na, K and Cl ions into and out of cells in an electrically neutral manner, and is involved in maintenance of cell volume.35, 36 The reduced expression of NKCC2, in conjunction with lowered expression of the potassium voltage-gated channel in C3 lenses, might be expected to affect the osmotic balance leading to abnormal fiber swelling (Figures 4, 5 and Figures 6) and vacuole accumulation as seen in C3 lenses (Figures 1 and 3).
One consistent and interesting characteristic of C3 mouse lens is a thickened anterior lens capsule. This abnormally thickened capsule might be a consequence of compositional change(s) deriving from altered synthesis of capsule components. With the lens capsule being composed of collagens and various basement membrane proteins,37 it is interesting to note that the expression of collagen type IV alpha3, dermatopontin, and latent TGF-beta binding protein were significantly upregulated in the C3 lens. Higher levels of expression of these genes, together with the marginally upregulated expression of nidogen2/entactin-2 (AB017202), osteoglycin (D31951) and collagen VIII related gene (AF095155), could potentially be involved in the increased capsule thickening noted in C3 lenses.
Additionally, the cDNA microarray analysis has also provided evidence for the upregulation of some of the genes known to be associated with the cell death program. Genes encoding several proapoptotic proteins, including JNK1, lymphotoxin beta-receptor, quaking protein-7 and granzyme F,38, 39, 40 were strongly upregulated in C3 lenses. C3 toxin has been reported to activate the JNK stress-signaling pathway and increased apoptosis in cell culture systems.41, 42 Consistent with this, we have also obtained some direct evidence for the induction of programmed cell death in 1-day-old C3 lenses, as evidenced by the increase in number of TUNEL (terminal deoxynucleotidyltransferase dUTP nick end labeling)-positive lens fiber cells (data not shown).
Thus, the data from this study clearly establish an important role for Rho GTPase function in maintaining lens fiber cell organization, cytoskeletal integrity and interfiber interactions, all of which are critical for normal lens growth and transparency. Although it is technically challenging to prove that the changes observed in this study are directly due to the inhibition of Rho GTPase, it is reasonable to conclude that inhibition of Rho GTPase activity would have a major impact on the actin cytoskeleton, lens fiber cell morphology and lens transparency, and that the other events may be sequelae of this primary insult.
Accession codes
Accessions
GenBank/EMBL/DDBJ
References
Etienne-Manneville S, Hall A . Rho GTPases in cell biology. Nature 2002;420:629–635.
Aelst VL, D'Souza-Schorey C . Rho GTPases and signaling network. Genes Dev 1997;11:2295–2322.
Kaibuchi K, Kuroda S, Amano M . Regulation of the cytoskeleton and cell adhesion by the Rho family GTPases in mammalian cells. Annu Rev Biochem 1999;68:459–486.
Takai Y, Sasaki T, Matozaki T . Small GTP-binding proteins. Physiol Rev 2001;81:153–208.
Braga VM, Machesky LM, Hall A, et al. The Small GTPases Rho and Rac are required for the establishment of cadherin-dependent cell-cell contacts. J Cell Biol 1997;137:1421–1431.
Arthur WT, Noren NK, Burridge K . Regulation of Rho family GTPases by cell–cell and cell–matrix adhesion. Biol Res 2002;35:239–246.
Hill CS, Wynne J, Treisman R . The Rho family GTPases RhoA, Rac1 and CDC42Hs regulate transcriptional activation by SRF. Cell 1995;81:1159–1170.
Perona R, Montaner S, Saniger L, et al. Activation of the nuclear factor-kappaB by Rho, cdc42 and rac-1 proteins. Genes Dev 1997;11:463–475.
Olson MF, Paterson HF, Marshall CJ . Signals from Ras and Rho GTPases interact to regulate expression of p21Waf1/Cip1. Nature 1998;394:295–299.
Welsh CF, Roovers K, Villanueva J, et al. Timing of cyclin D1 expression within G1 phase is controlled by Rho. Nat Cell Biol 2001;3:950–957.
Coleman ML, Sahai EA, Yeo M, et al. Membrane blebbing during apoptosis results from caspase-mediated activation of ROCK1. Nat Cell Biol 2001;3:339–345.
Wei L, Roberts W, Wang L, et al. Rho kinases play an obligatory role in vertebrate embryonic organogenesis. Development 2001;128:2953–2962.
Piatigorsky J . Lens differentiation in vertebrates: a review of cellular and molecular features. Differentiation 1981;19:134–153.
McAvoy JW, Chamberlain CG, de Iongh RU, et al. Lens Development. Eye 1999;13:425–437.
Bassnett S, Missey H, Vucemilo I . Molecular architecture of the lens fiber cell basal membrane complex. J Cell Sci 1999;112:2155–2165.
Lee A, Fischer RS, Fowler VM . Stabilization and remodeling of the membrane skeleton during lens fiber cell differentiation and maturation. Dev Dyn 2000;217:257–270.
Quinlan RA, Sandilands A, Procter JE, et al. The eye lens cytoskeleton. Eye 2002;13:409–416.
Menko AS . Lens epithelial cell differentiation. Exp Eye Res 2002;75:485–490.
Maddala RL, Peng YM, Rao PV . Selective expression of the small GTPase RhoB in the early developing mouse lens. Dev Dyn 2001;222:534–537.
Rao PV, Robison WG, Bettelheim F, et al. Role of small GTP-binding proteins in lovastatin-induced cataract. Invest Ophthalmol Vis Sci 1997;38:2313–2321.
Maddala RL, Reddy VN, Rao PV . Lovastatin-induced cytoskeletal reorganization in lens epithelial cells: role of Rho GTPases. Invest Ophthalmol Vis Sci 2001;42:2610–2615.
Maddala R, Reddy VN, Epstein DL, et al. Growth factor induced activation of Rho and Rac GTPases and actin cytoskeletal reorganization in human lens epithelial cells. Mol Vision 2003;9:329–336.
Rao PV, Wawrousek E, Tamm ER, et al. Rho GTPase inactivation impairs lens growth and integrity. Lab Invest 2002;82:231–239.
Rubin EJ, Gill DM, Boquet P, et al. Functional modification of a 21-kilodalton G protein when ADP-ribosylated by exoenzyme C3 of Clostridium botulinum. Mol Cell Biol 1988;8:418–426.
Freel CD, Gilliland KO, Mekeel HE, et al. Ultrastructural characterization and Fourier analysis of fiber cell cytoplasm in the hyperbaric oxygen treated guinea pig lens opacification mode. Exp Eye Res 2003;76:405–415.
Bradford MM . A rapid and sensitive method for the quantification of microgram quantity of protein utilizing the principle of protein dye-binding. Anal Biochem 1976;72:248–254.
White TW, Goodenough DA, Paul DL . Targeted ablation of connexin50 in mice results in microphthalmia and zonular pulverulent cataracts. J Cell Biol 1998;143:815–825.
Beebe DC, Vasiliev O, Guo J, et al. Changes in adhesion complexes define stages in the differentiation of lens fiber cells. Invest Ophthalmol Vis Sci 2001;42:727–734.
Agre P, King LS, Yasui M, et al. Aquaporin water channels from atomic structure to clinical medicine. J Physiol 2002;542:3–16.
Shiels A, Bassnett S . Mutations in the founder of the MIP gene family underlie cataract development in the mouse. Nat Gent 1996;62:212–215.
Khurana RN, Maddala RL, Shimokawa, et al. Inhibition of Rho kinase induces αB-crystallin expression in lens epithelial cells. Biochem Biophys Res Commun 2002;294:981–987.
Shiels A, Mackay D, Ionides A, et al. A missense mutation in the human connexin50 gene (GJA8) underlies autosomal dominant ‘zonular pulverulent’ cataract, on chromosome 1q. Am J Hum Genet 1998;62:526–532.
Francis P, Chung JJ, Yasui M, et al. Functional impairment of lens aquaporin in two families with dominantly inherited cataracts. Hum Mol Genet 2002;9:2329–2334.
White TW . Unique and redundant connexin contributions to lens development. Science 2002;295:319–320.
Alvarez LJ, Candia OA, Turner HC, et al. Localization of a Na(+)–K(+)–2Cl(−) cotransporter in the rabbit lens. Exp Eye Res 2001;73:669–680.
Haas M, Forbush B . The Na–K–Cl cotransporter of secretory epithelia. Annu Rev Physiol 2000;62:515–534.
Heathcote JG, Grant ME . The macromolecular composition of the embryonic chick lens capsule. Preliminary biosynthetic studies on the collageneous and non-collageneous glycoproteins. Exp Eye Res 1982;34:985–1000.
Weston CR, Davis RJ . The JNK signal transduction pathway. Curr Opin Genet Dev 2002;12:14–21.
Pilotte J, Larocque D, Richard S . Nuclear translocation controlled by alternatively spliced isoforms inactivates the QUAKING apoptotic inducer. Genes Dev 2001;15:845–858.
Trapani JA . Granzymes: a family of lymphocyte granule serine proteases. Genome Biol 2001;2:3014–3018.
Beltman J, Erickson JR, Martin GA, et al. C3 toxin activates the stress signaling pathways, JNK and p38, but antagonizes the activity of AP-1 in rat cells. J Biol Chem 1999;274:3772–3780.
Bokock GM . Regulation of cell function by Rho GTPases. Immunol Res 2000;21:139–148.
Acknowledgements
We thank the staff of Duke University Microarray Service facility, particularly Dr Holly Dressman for her suggestions and comments on the array analysis. This work was supported by NIH RO1-EY12201 and EY013573 grants (P.V.R).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Maddala, R., Deng, PF., Costello, J. et al. Impaired cytoskeletal organization and membrane integrity in lens fibers of a Rho GTPase functional knockout transgenic mouse. Lab Invest 84, 679–692 (2004). https://doi.org/10.1038/labinvest.3700105
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700105
Keywords
Keywords
This article is cited by
-
Gap Junctions and Wnt Signaling in the Mammary Gland: a Cross-Talk?
Journal of Mammary Gland Biology and Neoplasia (2019)
-
Modulation of connexin signaling by bacterial pathogens and their toxins
Cellular and Molecular Life Sciences (2011)
-
Inside Lab Invest
Laboratory Investigation (2004)