Abstract
Many molecular alterations are known to occur in urothelial carcinoma of the bladder, but their significance for tumor progression is poorly understood. Deletions of chromosome 8p are frequently found in several tumor types and are often associated with progressive disease. In all, 99 bladder tumors were screened for deletions at 8p using loss of heterozygosity (LOH) and multicolor fluorescence in situ hybridization FISH analyses. Allelic loss on chromosome 8p in at least one marker was found in 25/99 (25%) tumors. There was a significant correlation of 8p deletions with invasive tumor growth and a highly significant association with papillary growth pattern in patients with invasive disease. cDNA array analyses revealed that secreted Frizzled-related protein 1 (sFRP1), an antagonist of Frizzled receptors and Wnt pathway activation on chromosome 8p12–11.1, is frequently downregulated in bladder cancer. To investigate sFRP1 as a candidate for a putative progression-related gene on 8p, urothelial cell lines and primary urothelial carcinomas were screened for sFRP1 expression using quantitative real-time PCR, Northern blot, immunofluorescence and immunohistochemistry (IHC). Of the investigated bladder cancers, 38% showed loss of sFRP1 expression by quantitative RT-PCR. Evaluation of the protein expression by IHC using tissue microarrays containing 776 bladder cancers revealed loss or strong reduction of sFRP1 expression in 66% of cases. SFRP1 loss was associated with higher tumor stage and grade and shorter overall survival. In addition, loss of sFRP1 was an independent indicator of poor survival in patients with papillary but not with muscle invasive bladder cancer. There were neither mutations in the coding region of sFRP1 nor homozygous deletions at 8p12–11.21. However, promoter methylation was detected using methylation-specific PCR in 29% of cases. In conclusion, we could show a close correlation of chromosome 8p deletions and progression of papillary bladder tumors. The sFRP1 gene on chromosome 8p12–11.1 could be a candidate gene for the predicted, progression-related tumor suppressor gene in bladder cancer and could contribute to urothelial carcinogenesis.
Similar content being viewed by others

Main
Bladder cancer is the second most common malignancy of the genitourinary tract and the fourth most common cancer in men in the United States.1 Two different types of bladder cancer can be distinguished both clinically and histopathologically: (i) mostly superficial, papillary tumors (80%) with high rates of recurrence but only a low tendency for progression to invasive disease, and (ii) highly malignant, invasive tumors (20%) with a poor prognosis for the individual patient. The different clinical course is reflected by different oncogenetic pathways (reviewed by Cordon-Cardo et al2 and Knowles3). Deletions of several regions of both arms of chromosome 9 and activating mutations in the fibroblast growth factor receptor 3 (FGFR3) gene are very early and frequent events in the development of papillary tumors.4, 5 In contrast, solid invasive tumors frequently show deletions on chromosome 17p and mutations of the p53 gene, changes that can already be found in dysplasia or carcinoma in situ (CIS).6, 7, 8 Invasive tumors accumulate many additional genetic alterations, for example, losses on chromosomes 3p, 4p, 10q or 13q, changes that are not found in superficial papillary tumors.9, 10, 11, 12 Although there are several clinical and histopathological features that could indicate a higher progression risk of papillary tumors, for example, high tumor grade, multifocality and concomitant CIS, new molecular prognostic markers for papillary bladder cancers are needed.13
Deletions at chromosome 8p are frequently found in several tumor types, for example, colorectal, prostate, lung or hepatocellular cancer,14, 15 and are often associated with progressive disease.16 In bladder cancer, deletions on chromosome 8p are frequently detected in advanced tumor stages.3, 9, 10, 11, 12, 17, 18, 19
Detailed deletion mapping in bladder cancer indicates that there are at least two common deleted regions, one at 8p23.318 and another at 8p12–p11.2.20 Similar deletion regions were also detected in oral21 and ovarian cancer,22 suggesting the presence of one or more tumor suppressor genes (TSG) on this chromosome arm. Several candidate genes (eg WRN, TRAIL-R2, Dematin) have been identified on chromosome 8p.23, 24, 25 Whereas FEZ1/LZTS1 at 8p22 is frequently downregulated in bladder cancer,26 the DNA polymerase β gene (POLB) and an isoform of the catalytic subunit of protein phosphatase 2A (PPP2CB) do not seem to represent the TSG for bladder cancer.27
Another candidate gene on chromosome 8p12–11.1 is the secreted Frizzled-related protein 1 gene (sFRP1), an antagonist of Frizzled receptors and Wnt pathway activation.28 The Wnt proteins comprise a highly conserved, multimember ligand family, which play important roles in patterning and cell fate determination. The Wnt pathway (see, for review, Eastman and Grosschedl29 and Polakis30) involves binding of Wnt proteins to Frizzled receptors and stabilization of cytoplasmic beta-catenin. Stabilization leads to nuclear and cytoplasmatic accumulation of uncomplexed beta-catenin that binds to TCF/LEF transcription factors, subsequently activating the expression of nuclear target genes (eg cyclin D1 and c-myc) to establish the oncogenic phenotype.31, 32, 33
Accumulating evidence links Wnt signaling to tumorigenesis in several organs. Constitutive Wnt pathway exists in several human cancer types through mutations of one or more pathway members, as found in the genes for APC, axin and beta-catenin.34, 35
Several antagonists (eg sFRP1, FrzB, WIF-1)33 of the Frizzled receptors can also bind to Wnt molecules36 providing a precise adjustment of the crucial initiation of the Wnt pathway. sFRP1 directly binds to Wnt proteins and can actively regulate Wnt activity.37 Upregulation of sFRP1 has been associated with apoptosis,38, 39 and downregulation has been found in more than 80% of invasive breast tumors.40 Recently, epigenetic inactivation by promoter hypermethylation of several sFRP proteins, including sFRP1, could be demonstrated in colorectal and gastric cancers.41
In this study, we describe deletion mapping of chromosome 8p in 99 bladder tumors using loss of heterozygosity (LOH) and multicolor fluorescence in situ hybridization (FISH) analysis. A strong correlation of 8p deletions with invasive growth of papillary bladder tumors could be found. The loss of sFRP1 expression was frequent (mRNA and protein) and was associated with high stage and grade and poor survival, especially in papillary muscle invasive tumors.
Materials and methods
Tissue Samples
Fresh tumor tissue snap-frozen in liquid nitrogen in the operating room was obtained by transurethral resection or cystectomy from the Department of Urology, LMU Munich, Germany, as part of a clinical study evaluating 5-ALA-mediated fluorescence endoscopy.42 All patients gave informed consent. Histological evaluation of the resected specimens was carried out on frozen sections by two pathologists (AH, RK), and all tumors were diagnosed according to the WHO/ISUP classification of urothelial tumors43 and staged according to the TNM system.44 Papillary tumors of low malignant potential were excluded from the study. In all, 99 urothelial carcinomas (13 pTaG1, 14 pTaG2, one pTa G3, 12 pT1G2, 35 pT1G3, one pT2aG3, one pT2bG3, 16 pT2G3, one pT3aG3, one pT3G3, one pT3bG3, two pT4aG3, one pT4G3) were investigated. The growth pattern of the tumors was designated either as papillary (n=75, with at least 5% papillary growth in all sections of the tumor) or solid (n=21, no evidence of papillary growth). Three tumors were squamous cell carcinoma.
Cell Lines
Three different bladder cancer cell lines, representing various differentiation states of urothelial carcinogenesis, were used throughout this study. RT4 is derived from a well-differentiated, recurrent urothelial papillary low-grade tumor.45 RT112 is from a moderately differentiated transitional cell carcinoma of the urinary bladder.46 J82 is derived from an invasive, poorly differentiated high-grade urothelial carcinoma of the bladder.47 For comparison, the cell line UROtsa, derived from normal urothelium by immortalization with SV40 large T antigen,48 was investigated. Cells were maintained in RPMI supplemented with 1% L-glutamine, 1% sodium pyruvate and 5% fetal calf serum at 37°C in a humidified atmosphere containing 5% CO2.
DNA Isolation
DNA was isolated from frozen tumor tissue sections (5 μm) according to standard procedures. In brief, after ethanol fixation (70%) and methylene blue staining for 15 s, tumor cells were microdissected manually or by laser microdissection (PALM,49) to obtain at least 80% tumor cells as described previously.8 Fibrous tissue or areas of the lamina muscularis propria were microdissected from the same specimens for normal DNA. DNA was extracted by proteinase K digestion as described previously.50 DNA from cultured cells was extracted using the High Pure PCR Template Preparation Kit (Roche Diagnostics GmbH, Mannheim, Germany). For FISH analysis, the microdissected urothelial cells were incubated in CT100 (citric acid/0.5% Tween 20) for approximately 90 min at room temperature until the cytoplasm of the cells was dissolved. The cells were pelleted on silanized glass slides by standard microcentrifugation, fixed in freshly prepared methanol/acetic acid (3:1), air-dried and stored at −20°C.
DNA (CITB Libraries, clone no. RS549L18; Research Genetics) was isolated with alkaline lysis and labeled with digoxigenin-11-dUTP using standard nick translation protocols (Roche, Mannheim) as described previously.51
LOH analysis
The microsatellite markers used in this study and their location (Figure 1b) were taken from the Genome Database (http://www.gdb.org). All primers were obtained from GENSET SA (Paris, France). Matched normal/tumor DNA samples were amplified by PCR in a 25 μl volume containing 0.2 mmol/l dNTP, 0.3 μmol/l primers, 1.5 mmol/l MgCl2, 0.5 U Taq polymerase (Roche Diagnostics GmbH, Mannheim, Germany), using 1 μl isolated DNA as template. The reaction mixture was subjected to 3 min of denaturing at 95°C and 35 cycles of 95°C for 1 min, specific annealing temperature for 1 min and 72°C for 1 min, followed by a final extension step at 72°C for 10 min. PCR conditions were optimized by gradient PCR and were carried out in an MJ Research Thermocycler (PTC 100). Primer sequences and annealing temperatures are given in Table 1. The amplification products were visualized by polyacrylamide gel electrophoresis and silver staining as described.52 The silver-stained gels were assessed visually by two independent observers (RS and AH), and informative cases were scored as allelic loss when intensity of the signal for the tumor allele was decreased to at least 50% relative to the matched normal allele. To avoid errors due to preferential amplification of one allele during the PCR reaction, all LOH analyses were run in duplicates following independent PCR reactions.
Histomorphology and LOH and FISH analyses of chromosome 8 in papillary bladder tumors. (a) Poorly differentiated papillary urothelial carcinoma (patient 30, HE, × 50 and × 200). There was a large, predominantly moderately differentiated papillary urothelial carcinoma with focal poor differentiation with nuclear polymorphism, hyperchromasia and frequent mitosis throughout the urothelium (left) and multifocal stromal invasion (right). (b) Chromosomal localization of the microsatellite markers for LOH analysis and the BAC clone for FISH analysis. (c) Representative examples of allelic deletions (LOH) for microsatellite markers D8S1145 and D8S591 on chromosome arm 8p in bladder cancers. All analyses were run in duplicate to verify the results and to avoid false-positive results due to preferential PCR amplification. Arrows indicate loss of one allele. (d) FISH analysis of two bladder tumors with LOH on chromosome 8p. Red signal: centromer chromosome 8; green signal: locus-specific BAC clone 8p11.21; blue color: DAPI staining of the nucleus. In both tumors aneuploidy of chromosome 8 and deletion of 8p11.21 was found.
Fluorescence In Situ Hybridization
FISH was performed as described previously.53 Briefly, cells on slides were denatured in 70% formamide/2 × SSC, pH 7.0 at 75°C for 2.2 min. After dehydration in graded ethanol (70, 80 and 100% for 2 min each), samples were treated with proteinase K (Sigma) for 7 min at 37°C, followed again by ethanol dehydration. Proteinase K concentration varied between 0.4 and 0.8 μg/ml, dependent on tissue preservation and duration of slide storage. The hybridization mixture was denatured for 5 min at 75°C and subsequently reannealed for 40 min at 37°C. A measure of 10 μl of hybridization mixture (20–30 ng gene-specific probe, 1 μl direct labeled centromeric probe for chromosome 8 (CEP® 8 Spectrum Orange™, Vysis), 5–10 ng unlabeled, sonicated human placental DNA (Sigma) and 10 μl hybridization buffer (Vysis)) was applied to each cytospin. Hybridization was overnight at 37°C. For every hybridization cytospins of cultured urothelial cell line HCV29 without any alteration at the investigated gene locus were included to assure an estimation of hybridization efficiency. The gene-specific probe was visualized by immunostaining in two steps (10 μg/ml sheep anti-digoxigenein (Roche, Mannheim)/0.3 μg/ml FITC-conjugated anti-sheep IgG (Sigma)). Counterstaining was performed with DAPI in Vectashield mounting medium (Vector).
Scoring of FISH Signals
Cells were selected for scoring with DAPI staining according to morphological criteria. Clearly distinguishable small lymphocytes were disregarded, all other cells were scored. Slides were analyzed if more than 75% of cells were interpretable. Copy numbers for centromers and specific gene region were counted in at least 100 cells. Only cells with nonoverlapping and intact nuclei were counted. Cells without any signal were disregarded. All hybridizations were evaluated independently by two investigators (RS, AH) and the mean of both counts was used. As a measure of deletion, the percentage of cells containing either one copy of chromosome 8 and one gene-specific signal or fewer gene-specific signals than centromeric signals was calculated for each hybridization (defined as the percentage of deletion). The average percentage of deletion of 10 hybridizations of a urothelial cell line (HCV29) and six hybridizations of dissected normal urothelium from patients without bladder cancer was less than 15±5% for the used probes. A tumor was considered deleted for the specific chromosomal locus if the percentage of deletion was >40% (equivalent 2 × mean±s.d.) as a conservative evaluation of the results.
RNA Preparation
The microdissected tissue samples were collected in GTC buffer and poly-A+-RNA was prepared with the Poly-A-tract 1000 kit (Promega, Heidelberg, Germany), according to the manufacturer's recommendations. cDNA synthesis was initiated using a Oligo-dT-T7 primer promoter combination and in vitro transcription was performed to amplify the mRNA using the protocol of van Gelder et al54 with modifications using an ABI Prism Sequence Detection System 5700 (Applied Biosystems, Foster City, CA, USA). cDNA quantity and integrity was evaluated after each round of amplification by quantitative real-time PCR.
Quantitative RT-PCR
For quantitative RT-PCR, aRNA was reverse transcribed to cDNA using random hexamer primers. The amplification conditions consisted of an initial 10 min denaturation step at 95°C, followed by 40 cycles of denaturation at 95°C for 15 s and annealing and extension for 1 min at 60°C. Quantitative PCR was performed in a total reaction volume of 25 μl. The reaction mixture contained the TaqMan Universal PCR Master Mix (Applied Biosystems, Foster City, CA, USA), 300 nM Primers, 100 nM fluorophoric probe and 1 ng reverse transcribed aRNA. The oligonucleotides used for real-time RT-PCR are given in Table 1. cDNAs were quantified by the comparative CT method normalizing the CT-values to GAPDH and calculating the relative expression values of normal to cancerous tissue.55
Northern Blot Hybridization
Expression of sFRP1 in various human tissues and urothelial cancers was tested using commercial multiple tissue (Clontech, BD Biosciences, Erembodegem, Belgium) and bladder tumor blots (Invitrogen, Karlsruhe, Germany). Bladder tumor blot contained 20 μg total RNA from bladder tumors of different stages and grades. Hybridization was performed using a sFRP1-probe (Image Clone no. W21306, position: 240–2720; length: 2500 bp), which was verified by sequence analysis. 32P-labeling of the probe was achieved using the Megaprime™ DNA Labeling System (Amersham Biosciences, Freiburg, Germany). Hybridizations with a 32P-labeled DNA probe were performed according to the manufacturer's recommendation and analyzed using the Phosphoimager 595 (Molecular Dynamics, Sunnyvale, USA).
PCR Amplification and Mutation Analysis of sFRP1
All exons (1)–(3) and splice sites of the sFRP1 gene were examined for mutations by genomic sequencing analysis. Genomic DNA was amplified by PCR with six sets of primers. All primers were obtained from GIBCO Invitrogen Corporation (Karlsruhe, Germany). Primer sequences and annealing temperatures are given in Table 1. Each target sequence was amplified in a 25 μl reaction volume containing 50 ng DNA, 0.2 mmol/l dNTPs, 1.5 mmol/l MgCl2 (except exon 2 and 3′ region of exon 3: 2.5 mmol/l MgCl2), 2 pmol/l primers and 1.25 U Taq polymerase. PCR amplification consisted of 35 cycles (95°C for 30 s, specific annealing temperature for 30 s and 72°C for 30 s) after an initial preheating (95°C for 10 min). A final elongation step of 72°C for 10 min was performed. PCR products were purified before sequencing using the PCR Purification Kit (Qiagen, Hilden, Germany) according to the manufacturer's instructions. Sequencing was performed using ABI PRISM BigDyeTM Terminator Cycle Sequencing Ready Reaction Kit (Applied Biosystems, Foster City, CA, USA). PCR conditions were as follows: initial preheating (95°C for 5 min) and 35 cycles consisted of 95°C for 30 s, 50°C for 10 s and 60°C for 4 min. DNA (50 ng) were used for each reaction.
Bladder Cancer tissue microarray (TMA) and Clinical Follow-up Data
The construction and composition of the bladder cancer TMA containing 776 formalin-fixed, paraffin-embedded tissues were described previously.56, 57 All slides of all tumors were reviewed by one pathologist (GS) and tumor stage and grade were defined according to the UICC classification. Stage pT1 was defined by both unequivocal tumor invasion of the suburothelial stroma and tumor-free fragments of the muscular bladder wall. Cases with stromal invasion but no muscular bladder wall in the biopsy were classified as at least pT1 (pT1−). Growth pattern was determined for all invasive tumors. Papillary growth was defined if there was a papillary tumor component (≥20%) with identical grade as the invasive tumor. All other tumors were considered to have a solid growth pattern. These cases were considered in the statistical analysis with regard to histological type, grading and overall survival, but not in that with regard to stage or stage/grade. Patient survival time ranged from 1 to 236 months with a median follow-up period of 42 months in these patients.
sFRP1 Immunohistochemistry (IHC)
An avidin-biotin peroxidase method with diaminobenzidine was used on formalin-fixed, paraffin-embedded material after incubation and microwave antigen retrieval. Frozen sections (5 μm) were air-dried, fixed in acetone (−20°C for 5 min) and IHC was performed using ChemMate® Immunostaining Kit (Dako A/S, Denmark) following the manufacturer's instructions. IHC on formalin-fixed tissue was performed in an NEXES® Immuno Stainer (Ventana, Tucson, AZ, USA) following the manufacturer's instructions. The sFRP1 antibody (polyclonal rabbit anti-human sFRP1, Eurogentec, Seraing, Belgium, 1:50 dilution) was characterized by transfection experiments followed by Western blotting and immunofluorescence as described in detail elsewhere (Klopocki et al., submitted for publication) Sections were counterstained with hematoxylin and embedded in Entellan® mounting medium (EMS Diatome, Fort Washington, PA, USA). Immunohistochemical analysis was performed without the knowledge of histopathological or molecular data.
sFRP1 Immunofluorescence
Cultured cells were grown on standard glass slides. After acetone fixation (−20°C, 20 min), a standard indirect immune fluorescence labeling was performed. In brief, cells were incubated with 100 μl primary antibody (polyclonal rabbit anti-human sFRP1, amino acids 285–299, Eurogentec, Seraing, Belgium) (Klopocki et al., 2002, submitted for publication) in a dilution of 1:50 for 1 h at 37°C. Slides were washed with PBS several times, followed by incubation with 100 μl of a second antibody (swine anti-rabbit FITC-conjugated antibody; DAKO A/S, Denmark) in a dilution of 1:20 for 1 h 37°C. After several washes, slides were mounted with VECTASHIELD® Mounting Medium containing DAPI for nuclear staining, and assessed on a Axiovert S100 microscope (Zeiss, Germany).
Methylation Analysis of the sFRP1 Gene Promoter
The methylation analysis of the promoter region of the sFRP1 gene was performed using methylation-specific PCR (MSP) as described in Suzuki et al.41
Statistical Analysis
The χ2 test was used to determine whether there was an association among deletions on chromosome 8p, sFRP1 expression and histopathological characteristics of the tumors. P-values <0.05 were considered as significant. For simple multiple testing procedures P-values were adjusted using Bonferroni–Holm method.58
Contingency table analysis and χ2 test were used to study the relationship between histologic type, grade, stage and sFRP1 expression on the TMA. Survival curves were plotted according to Kaplan and Meier. A log-rank test was applied to examine the relationship between grade, stage of sFRP1 expression loss and clinical outcome. Only the first biopsy in patients with multiple tumors on the TMA was used for further statistical analyses. For the analysis of recurrence or progression, patients were censored at the time of their last clinical control showing no evidence of disease or at the date when cystectomy was performed. For survival analyses, patients were censored at the time of their last clinical control or at the time of death if they died from causes not related to their tumors. In all, 25 patients with unclear cause of death were excluded from survival analysis.
Results
Chromosome 8p Deletion Analysis by LOH and FISH Analysis
DNA from all 99 primary tumors and matched normal tissues was successfully investigated for allelic deletions. All investigated patients showed heterozygosity in at least 1/9 microsatellite markers, 97/99 (98%) patients showed heterozygosity in at least 3/9 markers. The marker D8S532 gave no interpretable results in any of the investigated patients. Allelic loss on chromosome 8p in at least one marker was found in 25/99 (25%) tumors (Figure 1c and Table 2). Of the 25 tumors, 11 showed LOH in more than three markers on chromosome 8p. In eight of the remaining 14 tumors more than three markers were not informative (NI) or not available (NA) and gave no information about the deletion status in these tumors. Of 25 (72%) tumors, 18 showed LOH at chromosome 8p12. In all, 11 tumors showed smaller deletions with a minimal region of deletion at 8p12–11.1 (D8S311, Table 2). There was a significant correlation of 8p LOH with infiltrating growth of superficial papillary tumors (pTa: 3/28 vs pT1: 17/37; P=0.016). Interestingly, the frequency of 8p deletion was lower in tumors with invasion of the muscularis propria (≥pT2) than in tumors with stromal invasion only (pT1) (5/24 vs 16/47; P=0.08). This result could be explained by the fact that the frequency of 8p LOH in infiltrating papillary tumors was much higher than in solid tumors without a papillary component (infiltrating papillary tumors: 20/48 vs solid tumors: 2/20; P=0.008). There was no significant association between 8p deletion and tumor grade. To adjust for multiple comparisons, the method of Bonferroni–Holm was used. Only one of the significant results did not reach the adjusted significance level (pTa vs pT1-4, P=0.048). The two other significant results (pTa vs pT1, P=0.016 and invasive vs papillary growth pattern, P=0.008) were still statistically significant.
Of 25 tumors with LOH at chromosome 8p, 16 could be successfully analyzed by FISH (Figure 1d, Table 2), using a centromeric probe for chromosome 8 and a locus-specific probe for 8p12–11.1 (sFRP1). Results from the LOH analysis could be confirmed in all 16 interpretable cases. There were no homozygous deletions in any of the tumors with LOH. In the nine noninformative cases either no tissue was available after microdissection and DNA isolation or FISH results were not interpretable (few cells, weak hybridization signals, tissue damage).
Northern Blot Analysis of sFRP1 Expression
RNA blot analysis of 15 different human normal tissues showed a homogeneous sFRP1 expression in all investigated organs (Figure 2a). Normal urothelial tissue showed clear sFRP1 expression, whereas all investigated tumor samples showed a complete loss of sFRP1 RNA (Figure 2b).
Expression analysis of sFRP1 in bladder cancer. (a) Northern blot analysis of sFRP1 in multiple human tissues. All specimens expressed the 4.4 kb transcript for the sFRP1 gene. Comparative loading and transfer of RNA were ascertained by beta-actin mRNA expression. (b) sFRP1 expression analysis in bladder tumors. Lanes 1, 3, 5, 7: RNA from bladder tumors (total RNA, 20 μg); lanes 2, 4, 6, 8: RNA from matched normal urothelium. Complete loss (lanes 1, 3) or strongly reduced (lanes 5, 7) sFRP1 expression in all tumors. (c) Promoter methylation analysis of the sFRP1 gene. All cases with weak or strong MSP signal are marked.
sFRP1 Sequence Analysis
The genomic sequencing analysis of the coding sequence of sFRP1 (exons and splice sites) was successfully performed in 18/25 tumors with allelic deletion on chromosome 8p. Neither mutations nor polymorphisms were found. A 3bp deletion (5′GCA3′ at codon 13) detected in tumor tissue of breast cancer patients (data not shown) was not found in this study. An additional 46 randomly selected snap-frozen bladder tumors with different staging and grading (ranging from pTaG1 to pT4aG3) were analyzed to avoid selection bias. No mutation or sequence variant was found in any of the investigated tumors.
Quantitative RT-PCR Analysis of sFRP1 Expression
Overall, 52 bladder tumors (ranging from pTaG1 and CIS to pT2G3) including the tumors from sequencing analyses, and 14 benign urothelial tissue samples (normal urothelium, urothelial hyperplasia and cystitis) were analyzed for sFRP1 expression by real-time quantitative PCR. RNA with a low level of degradation (defined as amplification of the housekeeping gene GAPDH within 27 cycles) could be extracted from 29/52 bladder tumors and 5/14 benign tissue samples. Of the 29 tumors, 11 (38%) but none of the five nonmalignant tissue samples showed loss of sFRP1 expression (defined as the lack of amplification within 35 cycles). The loss of sFRP1 expression was already detectable in 5/15 (33%) pTa tumors. There was no significant correlation between sFRP1 loss and histopathological features of the tumors.
sFRP1 Immunofluorescence
Four bladder cell lines were investigated for cellular distribution of sFRP1. The breast cancer cell line MCF7 that is known to lack sFRP1 expression38 was utilized as a control. In all urothelial cell lines, a perinuclear cytoplasmatic staining was seen, indicating a localization of sFRP1 in the Golgi apparatus as expected for a secreted protein. UROtsa, RT4 and J82 showed a strong staining, whereas in RT112 only a weak staining was found. MCF7 was negative (Figure 3a).
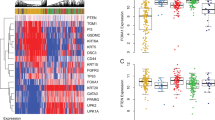
Expression of sFRP1 protein in urothelial cell lines and bladder cancers. (a) Immunofluorescence staining of urothelial cell lines J82 and RT112 for sFRP1 (green) protein. Strong perinuclear cytoplasmatic staining was found in J82 cells, RT112 cells showed a reduction of sFFP1 protein expression. FISH analysis of RT112 cells revealed chromosomal deletion of 8p11.21. MCF7 breast cancer cells were used as negative controls for sFRP1 protein expression. (b) Immunohistochemical staining of bladder cancers for sFRP1. I: papillary bladder cancer (pTaG2) with strong sFRP1 expression. Note the perinuclear cytoplasmatic localization of sFRP1 protein (× 400). II: papillary bladder cancer (pT1G2) with only faint staining for sFRP1. Note the positive staining of small vessels as internal control (× 400). (c) Immunohistochemical staining of Bladder cancer tissue microarray for sFRP1 representative examples of tumors showing strong, moderate and weak sFRP1 expression (× 50).
sFRP1 Immunohistochemistry
Immunohistochemical staining for the expression of sFRP1 was performed in 12/18 tumors with LOH on chromosome 8p12 (Figure 3b). No tissue was available from the remaining tumors. Nine tumors showed a strong sFRP1 staining, three tumors (25%) lacked sFRP1 protein expression (3x pT1G3). To avoid a bias due to preselection of the tumors resulting from the 8p deletion analysis, an additional 46 randomly selected, formalin-fixed, paraffin-embedded archival bladder tumors (16 pTaG1, 11 pTaG2, nine pT1G2, seven pT1G3, two pT2G2, one inverted papilloma) were investigated for sFRP1 expression in large tissue sections. In 12 (26%) of the 46 tumors no or only very weak sFRP1 protein expression was detectable (3 × pTaG1, 4 × pTaG2, 2 × pT1G2, 3 × pT1G3). There was no correlation between the lack of sFRP1 protein expression and the histopathological features in this small series. Investigation of a large series of bladder cancers (n=776, Figure 3c, Table 3) using the TMA technology59 revealed a loss of sFRP1 expression in 35% of cases (238/684) and a strong reduction in 31% of cases (215/684). Expression loss of sFRP1 was associated with higher tumor stage and grade and shorter overall survival (Figure 4a). In addition, loss of SFRP1 was an independent indicator of poor survival in papillary, but not solid muscle invasive bladder cancer (Figure 4b and c).
Kaplan–Meier survival curves using immunostaining for sFRP1 protein in tissue microarrays. (a) Expression loss of sFRP1 is strongly associated with a shorter overall survival of the patients. (b) SFRP1 is an independent predictor of the survival in patients with muscle invasive papillary bladder tumors. (c) No significant association was found between sFRP1 expression status and the survival of patients with solid muscle invasive tumors.
Methylation Analysis of The sFRP1 Promoter
The methylation status of CpG islands in the sFRP1 gene promoter was investigated in 42 bladder tumors (ranging from pTaG1 to pT4aG3) and six normal urothelial samples using MSP. We found a strong methylation in three tumors and weak methylation signals in nine tumors (Figure 2c). None of the normal urothelial samples showed a MSP signal. All three tumors with strong methylation signal also showed the loss of sFRP1 mRNA or sFRP1 protein. Only from 6/9 tumors with weak methylation additional data (quantitative RT-PCR or IHC) were available. Three tumors showed normal protein staining in the IHC analysis, three tumors lacked completely the sFRP1 protein or showed weak sFRP1 expression. In addition, 14 tumors with SFRP1 loss showed no MSP product.
Discussion
Most patients with papillary bladder cancer show an indolent clinical course with a low risk of progression. However, 20% of patients will eventually progress to invasive cancer. The search for molecular markers for risk assessment of papillary bladder cancer is an important focus of research in the last years.3
Deletions of chromosome 8p are found in many tumor types14, 15 and are associated with progression and metastasis of bladder, lung or breast cancer16, 19 Herein, we describe a deletion analysis using LOH analysis of chromosome 8p in 99 clinically and pathologically well-characterized bladder cancers of different stages and grades. There were deletions in 25% of all tumors. Interphase FISH analyses could confirm the deletions in all interpretable cases. There was a correlation of LOH at chromosome 8p with invasive tumor growth. Interestingly, this was due to a highly significant association of 8p deletions with progression of papillary bladder cancer (42%; P=0.008), whereas solid tumors without papillary component were only very infrequently deleted at this chromosome (9%). Previous LOH and FISH studies investigated the deletion frequency at chromosome 8p in bladder cancer. Simon et al19 showed a significant association between invasive tumor growth and 8p deletions detected by FISH. Takle et al20 found deletions of 8p in a very similar frequency to our results using LOH analyses. In contrast, Muscheck et al18 reported a much higher percentage of 8p deletions (75%) focusing on 8p23 and found a strong correlation with tumor grade (G3). However, all three studies did not correlate the growth architecture of the tumor with 8p deletions.
The correlation of LOH at chromosome 8p and progression of papillary tumors supports the development of invasive bladder cancer through two different pathways with stepwise accumulation of different genetic alterations and development of two distinct tumor entities (Figure 5, Cordon-Cardo et al.2 Low-risk noninvasive papillary tumors are characterized by early deletions of both arms of chromosome 94 and mutations of the FGFR3 receptor,5 and show very infrequent additional genetic alterations.10, 12 They frequently recurr but only rarely progress and could be called ‘papilloma’. In contrast flat, solid tumors (carcinoma in situ, CIS) show in addition to chromosome 9 deletions LOH at chromosome 17p13.1, frequent mutations in the p53 gene and many additional genetic alterations in early stages and develop muscle invasive disease very quickly. However, there seems to be a group of patients with initially papillary tumors and progression to aggressive bladder cancer (Figure 1a). Histopathological examination of cystectomy specimens with invasive bladder cancer frequently show coexisting poorly differentiated papillary tumors, CIS, and invasive cancer in the same patient, suggesting a possible overlap of both cancer pathways. Our data indicate that 8p deletions could be important in the transition of papillary tumors to invasive cancer. Comparative genomic hybridization studies by several groups comparing noninvasive papillary tumors (pTaG1–2) and invasive tumors (≥pT1)9, 10, 60, 61, 62 revealed additional genetic alterations associated with the invasive phenotype (eg deletions at 2q, 5q or 11p, see Figure 5), although an association with papillary growth architecture was not assessed in these studies.
Prospective studies using LOH and/or FISH analyses for progression-associated chromosomal regions that correlate the genetic data with the risk of recurrence and progression could reveal a marker panel useful in distinguishing harmless ‘papillomas’ from potentially dangerous papillary bladder cancers. Such marker panel could be utilized for more specific molecular urine screening with microsatellite analyses63 or interphase multicolor FISH.64, 65
The frequent chromosomal losses on 8p12–22 in many malignancies strongly suggest the presence of important tumor suppressor genes within this region. In bladder cancer, only three candidate genes have been investigated so far. The mutation analyses of POLB and PPP2CB did not find mutations in tumors with LOH at 8p, although a possible epigenetic inactivation was not investigated.27 Vecchione et al26 showed the downregulation or complete loss of FEZ1/LZTS1 in more than 60% of the investigated primary bladder tumors and a close correlation between loss or reduction of the protein and high-grade bladder tumors.
Gene expression studies using DNA microarrays are new and powerful tools to identify new tumor-related genes in chromosomal candidate regions. Both in silico analyses of differentially expressed genes in normal urothelium and bladder cancer searching different databases and expression analyses using a custom-made Affymetrix chip revealed a frequent loss of expression of the sFRP1 at chromosome 8p12–11.1 (data not shown).
sFRP1 is a member of the Wnt pathway, a multimember signal cascade implicated to play an important role in the development of many malignancies (see, for review, Morin34). The effects of activated Wnt signaling are tightly regulated. Soluble Wnt binding proteins (eg sFRP1, WIF-1, FrzB, Dkk) act as antagonists to the transmembrane Frizzled receptors modulating activation of the Wnt signal cascade.33 Cellular loss of one of these antagonists could result in an increased Wnt pathway activation with increased and uncontrolled proliferation leading to tumorigenesis.
The loss of sFRP1 expression detected in cDNA expression studies could be verified by quantitative real-time PCR and Northern blot in this study. Overall, 34% of the bladder tumors demonstrated the loss of sFRP1 expression. In immunohistochemical studies investigating large tissue sections using a polyclonal antibody, 25% of the tumors lacked sFRP1 expression (Figure 3b). In addition, most of the tumors demonstrated a weaker staining than the corresponding normal urothelium adjacent to the tumor. In 22 tumors investigated both with real-time PCR and immunohistochemistry, there was a very good correlation (73%) between both methods. Analysis of a large number of tumors on a TMA revealed SFRP1 loss as an independent indicator for poor survival in patients with invasive papillary tumors and a significant correlation with a shorter overall survival. These data demonstrate the power of TMAs allowing to investigate large patient cohorts.
The mechanism leading to downregulation of sFRP1 expression is not fully understood and remains unknown. We were not able to demonstrate sFRP1 mutations in sequencing analyses of 64 bladder tumors. FISH analyses in 16 tumors and four cell lines revealed no evidence for homozygous deletions. However, methylation analysis of the sFRP1 promoter showed a methylation signal in 29% of cases. Although promoter methylation is not as frequent than in colorectal and gastric cancer,41 our data are in accordance with the study from Suzuki et al41 and suggest that epigenetic gene silencing could be the reason for sFRP1 expression loss in bladder cancer. As we studied only one CpG site by MSP which was most frequently methylated in colorectal cancer,41 it is possible that we missed the methylation of CpG sites in the sFRP1 promoter in the remaining 14 cases with SFRP1 protein loss.
In conclusion, we could show a close correlation of chromosome 8p deletions and progression of papillary bladder tumors. Although the loss of the SFRP1 protein was not significantly associated with 8p deletion, it was associated with higher tumor stage and grade and poor overall survival especially in patients with muscle invasive papillary tumors. Promoter methylation was demonstrated as the likely reason of sFRP1 downregulation in a subset of tumors. These data suggest that sFRP1 could be one of the progression-related genes on chromosome 8p. The functional analysis of reduced sFRP1 expression in urothelial carcinogenesis awaits further analysis.
References
Stein JP, Grossfeld GD, Ginsberg DA, et al. Prognostic markers in bladder cancer: a contemporary review of the literature. J Urol 1998;160 (3 Part 1):645–659.
Cordon-Cardo C, Cote RJ, Sauter G . Genetic and molecular markers of urothelial premalignancy and malignancy. Scand J Urol Nephrol Suppl 2000;205:82–93.
Knowles M . What we could do now: molecular pathology of bladder cancer. J Clin Pathol: Mol Pathol 2001;54:215–221.
Simoneau M, Aboulkassim TO, LaRue H, et al. Four tumor suppressor loci on chromosome 9q in bladder cancer: evidence for two novel candidate regions at 9q22.3 and 9q31. Oncogene 1999;18:157–163.
Van Rhijn BW, Lurkin I, Radnanyi F, et al. The fibroblast growth factor receptor 3 (FGFR3) mutation is a strong indicator of superficial bladder cancer with low recurrence rate. Cancer Res 2001;61:1265–1268.
Spruck III CH, Ohneseit PF, Gonzalez-Zulueta M, et al. Two molecular pathways to transitional cell carcinoma of the bladder. Cancer Res 1994;54:784–788.
Stoehr R, Hartmann A, Hiendlmeyer E, et al. Oligoclonality in early lesions of the urothelium as determined by microdissection-supported genetic analysis. Pathobiology 2000;68:165–172.
Hartmann A, Schlake G, Zaak D, et al. Occurrence of chromosome 9 and p53 alterations in multifocal dysplasia and carcinoma in situ of human urinary bladder. Cancer Res 2002;62:809–818.
Richter J, Jiang F, Gorog JP, et al. Marked genetic differences between stage pTa and stage pT1 papillary bladder cancer detected by comparative genomic hybridization. Cancer Res 1997;57:2860–2864.
Richter J, Wagner U, Schraml P, et al. Chromosomal imbalances in early invasive (pT1) urinary bladder cancer. Cancer Res 1999;59:5687–5691.
Simon R, Burger H, Brinkschmidt C, et al. Chromosomal aberrations associated with invasion in papillary superficial bladder cancer. J Pathol 1998;185:345–351.
Voorter C, Joos S, Bringuier PP, et al. Detection of chromosomal imbalances in transitional cell carcinoma of the bladder by comparative genomic hybridization. Am J Pathol 1995;146:1341–1354.
Grossman HB . Biomarkers for transitional cell carcinoma—pro. Urology 2001;57:847–848.
Emi M, Fujiwara Y, Nakajima T, et al. Frequent loss of heterozygosity for loci on chromosome 8p in hepatocellular carcinoma, colorectal cancer, and lung cancer. Cancer Res 1992;52:5368–5372.
Vocke CD, Pozzatti RO, Bostwick DG, et al. Analysis of 99 microdissected prostate carcinomas reveals a high frequency of allelic loss on chromosome 8p12–21. Cancer Res 1996;56:2411–2416.
Chan KL, Lee JM, Guan XY, et al. High-density allelotyping of chromosome 8p in hepatocellular carcinoma and clinicopathologic correlation. Cancer 2002;94:3179–3185.
Schullerus D, von Knobloch R, Chudek J, et al. Microsatellite analysis reveals deletion of a large region at chromosome 8p in conventional renal cell carcinoma. Int J Cancer 1999;80:22–24.
Muscheck M, Sukosd F, Pesti T, et al. High density deletion mapping of bladder cancer localizes the putative tumor suppressor gene between loci D8S504 and D8S264 at chromosome 8p23.3. Lab Invest 2000;80:1089–1093.
Simon R, Richter J, Wagner U, et al. High-throughput tissue microarray analysis of 3p25 (RAF1) and 8p12 (FGFR1) copy number alterations in urinary bladder cancer. Cancer Res 2001;61:4514–4519.
Takle LA, Knowles MA . Deletion mapping implicates two tumor suppressor genes on chromosome 8p in the development of bladder cancer. Oncogene 1996;12:1083–1087.
Sunwoo JB, Sun PC, Gupta VK, et al. Localization of a putative tumor suppressor gene in the sub-telomeric region of chromosome 8p. Oncogene 1999;18:2651–2655.
Pribill I, Speiser P, Leary J, et al. High frequency of allelic imbalance at regions of chromosome arm 8p in ovarian carcinoma. Cancer Genet Cytogenet 2001;129:23–29.
Yu CE, Oshima J, Goddard KA, et al. Linkage disequilibrium and haplotype studies of chromosome 8p 11.1.21.1 markers and Werner syndrome. Am J Hum Genet 1994;55:356–364.
Walczak H, Degli-Esposti MA, Johnson RS, et al. TRAIL-R2: a novel apoptotis-mediating receptor for TRAIL. EMBOJ 1997;16:5386–5397.
Lutchman M, Pack S, Kim AC, et al. Loss of heterozygosity on 8p in prostate cancer implicates a role for dematin in tumor progression. Cancer Genet Cytogenet 1999;115:65–69.
Vecchione A, Ishii H, Baldassarre G, et al. FEZ1/LZTS1 is down-regulated in high-grade bladder cancer, and its restoration suppresses tumorigenicity in transitional cell carcinoma cells. Am J Pathol 2002;160:1345–1352.
Eydmann ME, Knowles MA . Mutation analysis of 8p genes POLB and PPP2CB in bladder cancer. Cancer Genet Cytogenet 1997;93:167–171.
Fich PW, He X, Kelly MJ, et al. Purification and molecular cloning of a secreted, Frizzled-related antagonist of Wnt action. Proc Natl Acad Sci USA 1997;94:6770–6775.
Eastman Q, Grosschedl R . Regulation of LEF-1/TCF transcription factors by Wnt and other signals. Curr Opin Cell Biol 1999;11:233–240.
Polakis P . Wnt signaling and cancer. Genes Dev 2000;14:1837–1851.
He TC, Sparks AB, Rago C, et al. Identification of c-myc as a target of the APC pathway. Science 1998;281:1509–1512.
Shtutman M, Zhurinsky J, Simcha I, et al. The cyclin D1 gene is a target of β-catenin/LEF pathway. Proc Natl Acad Sci USA 1999;96:5522–5527.
Huelsken J, Behrens J . The WNT signalling pathway. J Cell Sci 2000;113:3545.
Morin PJ . β-catenin signaling and cancer. Bio Essays 1999;21:1021–1030.
Freanhead NS, Britton MP, Bodmer WF . The ABC of APC. Hum Mol Gen 2000;10:721–733.
Bafico A, Gazit A, Pramila T, et al. Interaction of frizzled related protein (FRP) with Wnt ligands and the frizzled receptor suggests alternative mechanisms for FRP inhibition of Wnt signaling. J Biol Chem 1999;274:16180–16187.
Uren A, Reichsman F, Anest V, et al. Secreted frizzled-related protein-1 binds directly to Wingless and is a biphasic modulator of Wnt signaling. J Biol Chem 2000;275:4374–4382.
Melkonyan HS, Chang WC, Shapiro JP, et al. SARPs: a family of secreted apoptosis-related proteins. Proc Natl Acad Sci USA 1997;94:13636–13641.
Zhou Z, Wang J, Han X, et al. Up-regulation of human secreted frizzled homolog in apoptosis and its down-regulation in breast tumors. Int J Cancer 1998;78:95–99.
Ugolini F, Charafe-Jauffret E, Bardou VJ, et al. WNT pathway and mammary carcinogenesis: loss of expression of candidate tumor suppressor gene SFRP1 in most invasive carcinomas except the medullary type. Oncogene 2001;20:5810–5817.
Suzuki H, Gabrielson E, Chen W, et al. A genomic screen for genes upregulated by demethylation and histone deacetylase inhibition in human colorectal cancers. Nat Genet 2002;31:141–149.
Zaak D, Kriegmair M, Stepp H, et al. Endoscopic detection of transitional cell carcinoma with 5-aminolevulinic acid: results of 1012 fluorescence endoscopies. Urology 2001;57:690–694.
Mostofi FK, Davis CJJ, Sesterhenn IA . Histological typing of urinary bladder tumours. In: World Health Organization—International histological classification. Mosteti FW, Davis CJ, Sesterhenn IA (eds). Springer: Berlin, Heidelberg, New York, Tokyo, 1999.
Sobin LH, Wittekind CH . TNM classification of malignant tumours. In: Sobin LH, Wittekind CH (eds). Wiley-Liss: New York, 1997, pp 187–194.
Rigby CC, Franks LM . A human tissue culture cell line from a transitional cell tumour of the urinary bladder: growth, chromosome pattern and ultrastructure. Br J Cancer 1970;24:746–754.
Marshall CJ, Franks LM, Carbonell AM . Markers of neoplastic transformation in epithelial cell lines derived from human carcinomas. J Natl Cancer Inst 1977;58:1743–1751.
O’Toole C, Price TH, Ohnuki Y, et al. Ultrastructure, karyology and immunology of a cell line originated from a human transitional-cell carcinoma. Br J Cancer 1978;38:64–67.
Petzoldt JL, Leigh IM, Duffy PG, et al. Immortalisation of human urothelial cells. Urol Res 1995;23:3377–3380.
Schuetze K, Lahr G . Identification of expressed genes by laser-mediated manipulation of single cells. Nat Biotechnol 1998;16:737–742.
Hartmann A, Rosner U, Schlake G, et al. Clonality and genetic divergence in multifocal low-grade superficial urothelial carcinomas as determined by chromosome 9 and p53 deletion analysis. Lab Invest 2000;80:709–718.
Hartmannx A, Moser K, Kriegmair M, et al. Frequent genetic alterations in simple urothelial hyperplasias of the bladder in patients with papillary urothelial carcinoma. Am J Pathol 1999;154:721–727.
Schlegel J, Bocker T, Zirngibl H, et al. Detection of microsatellite instability in human colorectal carcinomas using a non-radioactive PCR-based screening technique. Virchows Arch 1995;426:223–227.
Sauter G, Deng G, Moch H, et al. Physical deletion of the p53 gene in bladder cancer: detection by fluorescence in situ hybridization. Am J Pathol 1994;144:756–766.
van Gelder RN, von Zastrow ME, et al. Amplified RNA synthesized from limited quantities of heterogeneous cDNA. Proc Natl Acad Sci USA 1990;87:1663–1667.
Fink L, Seeger W, Ermert L, et al. Real-time quantitative RT-PCR after laser-assisted cell picking. Nat Med 1998;4:1329–1333.
Nocito A, Bubendorf L, Tinner EM, et al. Microarrays of bladder cancer tissues are highly representative of proliferation index and histological grade. J Pathol 2001;194:349–357.
Simon R, Mirlacher M, Sauter G . Tissue microarray in cancer diagnosis. Expert Rev Mol Diagn 2003;3:421–430.
Ludbrook J . Multiple comparison procedures updated. Clin Exp Pharmacol Physiol 1998;25:1032–1037.
Kononen J, Bubendorf L, Kallioniemi A, et al. Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat Med 1998;4:844–847.
Richter J, Beffa L, Wagner U, et al. Patterns of chromosomal imbalances in advanced urinary bladder cancer detected by comparative genomic hybridization. Am J Pathol 1998;153:1615–1621.
Prat E, Bernues M, Caballin MR, et al. Detection of chromosomal imbalances in papillary bladder tumors by comparative genomic hybridization. Urology 2001;57:986–992.
Zhao J, Richter J, Wagner U, et al. Chromosomal imbalances in noninvasive papillary bladder neoplasms (pTa). Cancer Res 1999;59:4658–4661.
Van Rhijn BW, Lurkin I, Kirkels WJ, et al. Microsatellite analysis—DNA test in urine competes with cystoscopy in follow-up of superficial bladder carcinoma: a phase II trial. Cancer 2001;92:768–775.
Halling KC, King W, Sokolova IA, et al. A comparison of cytology and fluorescence in situ hybridization for the detection of urothelial carcinoma. J Urol 2000;164:1768–1775.
Bubendorf L, Grilli B, Sauter G, et al. Multiprobe FISH for enhanced detection of bladder cancer in voided urine specimens and bladder washings. Am J Clin Pathol 2001;116:79–86.
Acknowledgements
We thank Ms Andrea Schneider, Ms Monika Kerscher, Ms Martina Mirlader and Mr Rudolf Jung for excellent technical assistance in tissue preparation, sequence analysis and immunohistochemistry. This work was supported by the following grants: 10-1598/HA2 (Dr Mildred Scheel-Stiftung for Cancer Research); Kn 263/7-2 (German Research Foundation DFG); 01KW9940 (BMBF).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Stoehr, R., Wissmann, C., Suzuki, H. et al. Deletions of chromosome 8p and loss of sFRP1 expression are progression markers of papillary bladder cancer. Lab Invest 84, 465–478 (2004). https://doi.org/10.1038/labinvest.3700068
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700068
Keywords
This article is cited by
-
Functional analysis of the Frzb2 gene in Schistosoma japonicum
Veterinary Research (2019)
-
Urothelial organoids originating from Cd49fhigh mouse stem cells display Notch-dependent differentiation capacity
Nature Communications (2019)
-
Fluorescence in situ hybridization as prognostic predictor of tumor recurrence during treatment with Bacillus Calmette–Guérin therapy for intermediate- and high-risk non-muscle-invasive bladder cancer
Medical Oncology (2017)
-
Apoptosis associated with Wnt/β-catenin pathway leads to steroid-induced avascular necrosis of femoral head
BMC Musculoskeletal Disorders (2015)
-
Functional analyses and prognostic significance of SFRP1 expression in bladder cancer
Journal of Cancer Research and Clinical Oncology (2015)