Abstract
Roughly 25% of cadaveric, but rarely living donor renal transplant recipients, develop postischemic acute renal failure, which is a main risk factor for reduced long-term allograft survival. An accurate prediction of recipients at risk for ARF is not possible on the basis of donor kidney morphology or donor/recipient demographics. We determined the genome-wide gene-expression pattern using cDNA microarrays in three groups of 36 donor kidney wedge biopsies: living donor kidneys with primary function, cadaveric donor kidneys with primary function and cadaveric donor kidneys with biopsy proven acute renal failure. The descriptive genes were characterized in gene ontology terms to determine their functional role. The validation of microarray experiments was performed by real-time PCR. We retrieved 132 genes after maxT adjustment for multiple testing that significantly separated living from cadaveric kidneys, and 48 genes that classified the donor kidneys according to their post-transplant course. The main functional roles of these genes are cell communication, apoptosis and inflammation. In particular, members of the complement cascade were activated in cadaveric, but not in living donor kidneys. Thus, suppression of inflammation in the cadaveric donor might be a cheap and promising intervention for postischemic acute renal failure.
Similar content being viewed by others
Main
Kidney transplantation is the treatment of choice for end-stage renal disease. Despite a reduction in the number of early acute rejection episodes by new immunosuppressive protocols, the incidence of postischemic acute renal failure (ARF) of cadaveric donor organs remained virtually unchanged over the past two decades. Kidneys from live donors hardly ever exhibit ARF and display longer allograft survival compared to cadaveric organs.1 The rate of ARF was reported to average 25% in 1980 and was still in the same range in 2001. ARF is a key risk factor for reduced long-term allograft survival and acute rejection occurs more frequently in grafts with ARF.2 ARF and early rejection episodes exert an additive negative effect on allograft survival. An accurate prediction of donor kidneys at risk for subsequent ARF would be of major advantage in the postoperative management, but cannot be performed on the basis of clinical or demographic donor and recipient parameters. Therefore, investigators focused on the evaluation of certain pathways in the donor organ, which have been reported to be essential regulators of the ischemia–reperfusion injury. Among these groups are apoptosis regulatory genes and adhesion molecules.3, 4 Furthermore, it has been shown that the number of apoptotic tubule cells was higher in donor kidneys with subsequent ARF compared to primarily functioning grafts.5 All these studies used a deductive approach in that the investigators picked certain regulators that have been shown to be involved in the process of ischemia–reperfusion injury. It is, however, very likely that other yet unknown or not investigated molecules contribute additionally to the regulation of ARF. The large number of genes potentially involved in the regulation of cell death and resolution of ARF emphasizes the importance of studying multiple gene expression alterations in concert. Systematic investigation of expression patterns of thousands of genes in donor kidney biopsies with and without subsequent ARF might provide the basis for a better understanding of the pathophysiology of ARF and may ultimately lead to rational prophylactic and therapeutic interventions.
We therefore applied an inductive approach to genome-wide gene-expression analysis using cDNA microarrays, together with descriptive statistics of donor and recipient parameters, to identify key regulatory pathways of ARF. We, furthermore, evaluated the differences in genome-wide gene-expression between kidneys from living and cadaveric organ donors and identified the main functional roles of these genes.
Material and methods
Donor and Recipient Characteristics
An average of 200 renal transplantations are performed annually at the University of Vienna Medical Center. Between April 2001 and January 2002, 102 cadaveric donor renal transplantations and 20 live donor renal transplantations were performed. Only the 82 cadaveric donor organs from heart-beating donors with well-documented ICU stay were used. In total, 15 of the live donors could be included, the other five refused to participate in the study. In order to study a homogeneous group of recipients, only first transplant, nonsensitized recipients with cyclosporine-based standard triple immunosuppression were included. These criteria were met by 14 recipients of live donations and by 32 recipients of cadaveric organs. None of the living donor organ recipients, except 15 out of 32 selected recipients of cadaveric organs, developed ARF. ARF was defined as dialysis dependency of at least 7 days post-transplant in the absence of biopsy-confirmed rejection. An early rejection within the first week after transplantation was diagnosed by biopsy in one out of the 14 living donor organ recipients and three out of the 17 cadaveric organ recipients with primary graft function. Thus, 13 living donor organs and recipients with primary function (LIV-PF), 14 recipients of cadaveric donor organs and with primary function (CAD-PF) and 15 cadaveric donor organ recipients with ARF (CAD-ARF) were available. The RNA of donor kidney biopsy was of insufficient quality or amount in one LIV-PF, two CAD-PF and three CAD-ARF. Thus, 12 LIV-PF, 12 CAD-PF and 12 CAD-ARF were studied. Nine key attributes of the 66 obtained donor and recipient characteristics are listed in Table 1.
The study was approved by the local Ethical Committee (# EK-415/01; to be found at http://ohrp.cit.nih.gov/search/asearch.asp).
Donor Kidney Biopsy Specimen, RNA Isolation and Amplification
All cadaveric and living donor kidneys were perfused with the histidine–tryptophan–ketoglutarate (HTK) cold preservation solution at 4°C during organ procurement.6 Wedge biopsies were obtained under sterile conditions at the end of the cold ischemia time immediately before transplantation. The biopsies were instantly submerged in RNAlater™ (Ambion, Austin, TX, USA) and homogenized.7 Total RNA was isolated and purified with RNeasy columns (Qiagen, Hilden, Germany). The RNA yield and quality was checked with the Agilent 2100 Bioanalyzer and RNA6000 LabChip® kit (Agilent, Palo Alto, CA, USA). Stratagene universal human reference RNA was used as standard (Stratagene, La Jolla, CA, USA).
Owing to the small amounts of isolated total RNA (8–15 μg per biopsy), a T7 RNA amplification step using the RiboAmp RNA amplification kit (Arcturus, Mountain View, CA, USA) was necessary as described and evaluated previously by us (M. Rudnicki, personal communication) and others.8, 9 The amplification factor was generally between 500 and 700 fold and the amplification profile was checked on an ethidium bromide stained 1% agarose gel and on the Agilent 2100 Bioanalyzer.
Microarray Analysis
cDNA microarrays holding 26 338 genes and 14 783 ESTs were obtained from the Stanford University Functional Genomics core facility. All microarray experiments were performed as described previously.10 Detailed protocols are available at http://genome-www.stanford.edu/. In brief, 800 ng of sample and standard aRNA were labeled with CyScribe cDNA postlabeling kit (Amersham Pharmacia Biotech, Buckinghamshire, UK) in a two-step procedure.
Samples were loaded on arrays and incubated for 16 h in a 65°C water bath. After two washing steps, fluorescent images of hybridized microarrays were obtained by using a GenePix 4000B scanner (Axon Instruments, Union City, CA, USA). Image griding and calculation of spot intensity was performed with GenePix Pro 4.0 software. The primary data tables and the images files were stored in the Stanford microarray database (http://genome-www4.stanford.edu/MicroArray/SMD/)11. Information on materials, experimental setup and procedures is in accordance with the MIAME microarray guidelines,12 and can be found at http://www.akh-wien.ac.at/user/rainer.oberbauer. The arrays were processed in random order and renumbered sequentially afterwards in order to facilitate orientation in the manuscript.
Validation of Microarray Results by Real-Time PCR
An ABI Prism 7700 Sequence Detector thermal cycler was used for all PCR experiments (Applied Biosystems, Foster City, CA, USA). Quantitative real-time PCR was performed and analyzed as described previously; 18S RNA served as internal standard.13 The validation of microarray experiments was performed in eight LIV and nine CAD kidneys for five genes of different main functional groups—complement system, general metabolism, protein modification, cell cycle and miscellaneous functions (Table 2). The PCR primer and TaqMan probes for these genes were purchased from Applied Biosystems. The same universal human RNA from Stratagene that was used in the microarray experiments served as reference in the PCR analysis. The delta Ct method was used to calculate the relative expression of each of the five genes in the samples compared to the Stratagene RNA.13 The differences in gene expression between the investigated CAD and LIV samples were evaluated by an unpaired t-test.
Statistical Analysis
The full data set consisting of 41 121 genes and ESTs was used for data preprocessing. First the data set was cleared from missing values using an 80% filter. The remaining missing values were substituted applying a k-nearest-neighbor algorithm, where the number of neighbors, k, was set to 10.14 Finally, another filter was applied on the remaining data set selecting all genes with a standard deviation above 0.9. A total of 1081 genes passed the data preprocessing and were used for further analysis. Two different batches of microarrays were used in all three kidney groups. To account for a putative bias of the two different batches, we applied singular-value decomposition analysis as described elsewhere.15
A hierarchical cluster algorithm within Statistica 6 (Statsoft Inc., Tulsa, OK, USA) was used for inductive clustering of arrays.16 Complete linkage with Pearson R linkage distance was computed. Statistical significance of genes separating arrays in distinct clusters was performed by calculating P-values, which were corrected for multiple testing using a maxT step-down procedure.17 This significance analysis was performed in the Bioconductor module of the R software package (http://www.bioconductor.org/).18 To reveal the sensitivity of the significant genes on putative outliers, a Jackknife procedure was performed.19 The descriptive genes were finally characterized according to their biological function using GenMAPP and HAPI.20, 21
Results
Classification of Donor Biopsies by Hierarchical Clustering of Gene Expression Patterns
Live versus cadaveric donor samples
The hierarchical clustering of 32 kidneys is displayed in Figure 1a. Four of the 36 kidneys had to be censored because of known post-transplant reason for ARF (#CAD-ARF9-12): ureteric leakage and drainage of urine into the abdominal cavity via a peritoneal perforation (#CAD-ARF9), early acute rejection in the revised histopathological diagnosis of the first-week biopsy (#CAD-ARF10), transplant artery stenosis that was successfully treated by PTA in week 2 (#CAD-ARF11) and hemorrhagic shock of the recipient on the first day due to severe anastomotic bleeding (#CAD-ARF12).
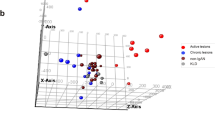
Hierarchical clustering of LIV-PF, CAD-PGF and CAD-ARF. Hierarchical clustering of the full 32-sample data set (a). Live donor kidneys clearly separate from the cadaveric donor renal samples. After adjustment for multiple comparisons, 132 genes could be identified as being significantly differentially expressed between these groups. The cadaveric samples did not clearly separate into CAD-PF and CAD-ARF groups. Especially CAD-PF #6–12, indicated by the gray underscore, clustered into the ARF group. Of note, left and right kidney from the same cadaveric donor are #CAD-ARF1 and CAD-PF5; CAD-PF8 and 9; CAD-PF 11 and 12, indicated by the gray rectangular background. Hierarchical clustering of the processed data set holding 24 samples (b). After maxT correction for multiple comparisons, 48 genes remained significantly differentially regulated between the CAD-PF and CAD-ARF groups. Genetic profiling correctly predicted early allograft function in all 12 LIV-PF kidneys, five of 12 CAD-PF kidneys and seven of eight CAD-ARF kidneys (exception CAD-ARF1). If ARF was defined less stringent, for example, no optimal function within the first 3 days, additional four CAD-PF would become CAD-ARF kidneys and thus would have been correctly ranked.
The gene expression profile of kidneys from live donors was clearly different from those of the cadaveric organs. We identified 132 genes and ESTs that, after maxT adjustment for multiple comparisons and Jackknife sensitivity analysis, clearly separated the two groups. Among the 132 sequences, 90 could be assigned to gene ontology terms (http://www.geneontology.org); the remaining 40 sequences were ESTs, hypothetical proteins and ORFs. The main functional roles of the identified 90 genes are cell communication,22 cell cycle/growth,23 immune response/complement,11 and metabolism.9 The remaining 25 genes had other functions. A total of 65% of the isolated genes were upregulated in the CAD samples, in particular, the genes involved in cell communication and immune response. These experimentally derived functional roles around cell communication and immune response fit well to the clinical observation of severe systemic inflammation in cadaveric organ donors, which is caused by an autonomous storm after brain death.
Our data set included three pairs of left and right kidney from the same cadaveric donor (#CAD-ARF1&CAD-PF5; CAD-PF8&9; CAD-PF11&12). These pairs clustered exceptionally well, as illustrated by the gray underlay in Figure 1a. Of note is the overall short linkage distance within groups, suggesting similar expression profiles within clusters.
Evaluation of intragroup gene expression pattern homogeneity
The LIV-PF kidneys (living donor kidneys with primary function) exhibited a homogenous gene expression profile (Figure 2a). The 1-Pearson R linkage distance was below 0.2, thus representing an overall good correlation.
Group–internal clustering of LIV-PF, CAD-PF and CAD-ARF. Group-internal clustering of kidneys from living donors (LIV-PF), cadaveric donor kidneys with primary function (CAD-PF) and ARF (CAD-ARF). The linkage distances, given as 1–Pearson R are denoted. The LIV-PF group is homogenous (a), whereas the CAD-PF group shows a clear split into two subsets, represented as PF groups 1 and 2 (b). Four of the seven PF group 2 members exhibited delayed graft function, although not meeting the strict ARF criteria (a). Those PF group 2 kidneys exhibited similar gene expression pattern as the CAD-ARF kidneys (Figure 1a). The CAD-ARF group represents a homogenous cluster, with the exception of CAD-ARF1.
The CAD-PF kidneys (cadaveric donor kidneys with primary function) were split into two clusters, based on their gene expression profiles (Figure 2b): PF group 1 holding five kidneys, and PF group 2 consisting of seven kidneys. Four out of these seven PF group 2 kidneys (#CAD-PF8, 9, 11, 12) exhibited delayed graft function, as expressed by the high creatinine levels in the first week, although not meeting the strict criteria of ARF. This is illustrated in Figure 3a, where the individual post-transplant course of serum creatinine concentrations is illustrated. These four kidneys clustered together with the other CAD-ARF kidneys (Figure 1a).
Post-transplant course of serum creatinine. Course of recipients who were classified as CAD-PF, not meeting the strict, arbitrary definition of CAD-ARF (a). However, four (#8, 9 and 11, 12) of the five CAD-PF subjects (#8–12), who clustered into the CAD-ARF group in Figure 1a, exhibited delayed allograft function indicated by slowly decreasing serum creatinine values within the first few days after engraftment. Creatinine values of CAD-ARF subjects (b). They all were dialysis dependent for at least 1 week after transplantation.
With the exception of CAD-ARF1, all CAD-ARF kidneys exhibit a homogenous gene expression profile (Figure 2c). A very likely clinical explanation for the CAD-ARF1 outlier, which clustered to the CAD-PF group in Figure 1a, is the long anastomosis time of 60 min. Furthermore, CAD-ARF1 and CAD-PF5 were left and right kidney from the same donor (Figure 1a).
Classification of cadaveric donor kidneys
In order to identify gene expression patterns that were able to separate the two CAD groups clearly, we censored the kidneys of intermediate clinical function (CAD-PF#8, 9, 11 and 12) and the CAD-ARF1 kidney (Figure 1b). The unexplained CAD-PF kidneys #6, 7 and 10 were not considered in the analysis, because their inclusion would have precluded the identification of separator sequences. Although we did not have a solid explanation for their omission, it is of note that these kidneys experienced the shortest anastomosis times among the PF group samples.
After these adjustments, 48 genes and ESTs were identified, which significantly separated ARF from PF kidneys. Among these 48 genes, 20 were robust separators for all array substitutions, as evaluated by Jackknife sensitivity analysis. The 48 genes of interest could be classified into the main gene ontology terms of cell cycle regulation,4 cell growth/metabolism,10 and communication/signal transduction.9 Three had other functions, six were ESTs and for 16 no defined gene ontology could be assigned. However, using this gene expression profiling in our data set yielded correct post-transplant group prediction in seven of eight clear CAD-ARF kidneys and in five of eight clear CAD-PF kidneys. Interestingly, all 48 genes were upregulated in the CAD-ARF cases compared to CAD-PF samples. Comparing LIV and CAD already showed this tendency of gene upregulation for CAD samples, and this finding is even more prominent in the CAD-ARF situation. The functional roles involved in the group segregation clearly reflect the peculiar biochemical status of cadaveric samples, where CAD-ARF shows additional upregulation of metabolic cascades.
Metabolic Networks Separating LIV-PF from CAD, and CAD-PF from CAD-ARF
Genes significantly separating LIV-PF and CAD, as well as CAD-PF and CAD-ARF, mainly belong to the functional roles cell communication, cell cycle/growth, immune response/complement and metabolism (Table 3). One predominant pathway, comparing LIV-PF and CAD, is the classical complement pathway: The C1r and C1s subunits of the C1 complex are strongly upregulated in the CAD situation, triggering the cleavage of C4 into C4a and C4b. Furthermore, the factor C2 is upregulated in the CAD samples, also cleaved by C1r and C1s. These products are key elements towards the formation of the membrane attack complex. Another typical differentiator in the CAD-PF to CAD-ARF situation is casein kinase II, blocking dephosphorylation of c-JUN, which further hinders the formation of the AP-1 complex. The transcription factor AP1 is, among other sites, involved in the transcription initiation of several detoxifying genes.
Post-Transplant Clinical Course
Figure 3a illustrates the individual course of the serum creatinine concentrations in recipients of CAD-PF kidneys, and Figure 3b shows the individual course of CAD-ARF recipients. In order to obtain useful information, we categorized the continuous variable ‘excretory renal function’ into two groups, PF and ARF. As mentioned above, recipients of kidneys #CAD-PF8, 9, 11 and 12 exhibit a protracted post-transplant decline of serum creatinine and clustered together with the CAD-ARF kidneys (Figure 1a).
The similarity of selected recipients precluded the evaluation of clinical factors for early allograft function, which was not the aim of this study.
Validation of Microarray Experiments
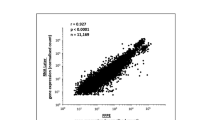
The expression of the five selected genes determined by real-time PCR in CAD and LIV kidneys relative to the universal human RNA is displayed in Table 2. PCR analysis confirmed the differences in the expression of these genes between LIV and CAD kidneys that were identified by the microarray studies before.
Discussion
We reported the genome-wide gene-expression profiles of 32 donor kidney biopsies using cDNA microarrays. The complete data set is available in a searchable format on the Stanford microarray database at http://genome-www4.stanford.edu/MicroArray/SMD/.
In these kidney biopsies, we identified a set of 132 genes that clearly separated living donor from cadaveric donor organs. Many of these genes play a major role in cell communication, growth/survival, inflammation, and metabolic pathways. The experimental data on gene expression nicely reflect the clinical observation of severe systemic inflammation in brain-dead organ donors.22, 23 We previously identified key members of the inflammation cascade, namely the adhesion molecules, and mitochondrial controllers of apoptosis as being strongly differentially regulated in cadaveric donor organs compared to kidneys from live donors.3 Cadaveric renal transplant donors often experience hemodynamic instability due to central diabetes insipitus and hormonal misregulation of circulation and massive cytokine release. Therefore, the majority of cadaveric donors receive vasopressor substances during their ICU stay. All these factors contribute to renal hypoperfusion and tissue ischemia.24, 25 Living donors on the other hand receive thoroughly diagnostic investigations before an organ donation is considered. Systemic inflammatory disease or arterial hypotension are contraindications for donation. Recipients of living donor kidneys usually do not exhibit postischemic ARF and have a significantly longer graft half-life than those of cadaveric grafts.1 The obviously shorter duration of cold ischemia of living donor kidneys is unlikely to contribute considerably to the better post-transplant function, since cell metabolism virtually ceases during organ preservation at 4°C. Genome-wide gene-expression patterns are not different in sequential donor kidney biopsies obtained at explantation and after 24 h of cold ischemia (personal communication). Furthermore, locally shared cadaveric organs with equally short ischemia time as living donor organs exhibit a higher incidence of ARF and reduced graft survival.26
We also identified 48 genes in cadaveric donor kidney biopsies that separated organs with primary function from those with subsequent acute renal failure. The majority of these genes could be classified as members of signal transduction pathways, cell cycle regulation, cell growth/metabolism and others. Based on these gene-expression patterns, a predictive classification of donor kidneys at risk for post-transplant ARF might be possible. The prediction of postischemic ARF would considerably alter the early postoperative management of the transplant recipients. Furthermore, the identification of the novel pathways in the regulation of ischemic injury might offer the opportunity for rational prophylactic and therapeutic approaches to ARF.
It is of major importance to avert ARF, because it is the key risk factor for reduced allograft half-life.27 The data of over 86 000 patients from the United Network of Organ Sharing registry showed that the half-life of a cadaveric donor kidney with ARF averaged 8 years compared to 14 years for primary functioning grafts.28
The obtained expression pattern of 48 genes could correctly classify seven of eight ARF and five of eight PF kidneys. This suggests that other, nondonor organ factors such as warm ischemia or anastomosis time, contribute to early allograft function.2 This issue was addressed by Suri and Meyer,29 who studied the influence of donor factors of early allograft function in 61 donors who provided kidneys that were transplanted locally into two separate recipients.29 Pairwise ANOVA showed that donor factors accounted for 45% of the variation in recipient creatinine clearance up to week 2 after engraftment. It should be emphasized that errors in the precise assessment of graft function caused an underestimation of the importance of donor factors. Therefore, the donor effect on early allograft function may have been underestimated by these authors and in our study, since we categorized early transplant function into PF and ARF. A correlation of the gene-expression data with early renal function given as a continuous variable was, however, not feasible with the limited number of samples.
The current study revealed a large aggregate effect of donor factors mirrored by a distinct gene-expression profile and some individual donor and recipient factors on early graft function. We identified two distinct gene-expression patterns, which could clearly separate living from cadaveric donors and primary graft function from postischemic acute renal failure.
References
Hariharan S, Johnson CP, Bresnahan BA, et al. Improved graft survival after renal transplantation in the United States, 1988 to 1996. N Engl J Med 2000;342:605–612.
Ojo AO, Wolfe RA, Held PJ, et al. Delayed graft function: risk factors and implications for renal allograft survival. Transplantation 1997;63:968–974.
Schwarz C, Hauser P, Steininger R, et al. Failure of BCL-2 up-regulation in proximal tubular epithelial cells of donor kidney biopsy specimens is associated with apoptosis and delayed graft function. Lab Invest 2002;82:941–948.
Schwarz C, Regele H, Steininger R, et al. The contribution of adhesion molecule expression in donor kidney biopsies to early allograft dysfunction. Transplantation 2001;71:1666–1670.
Oberbauer R, Rohrmoser M, Regele H, et al. Apoptosis of tubular epithelial cells in donor kidney biopsies predicts early renal allograft function. J Am Soc Nephrol 1999;10:2006–2013.
Groenewoud AF, Thorogood J . Current status of the Eurotransplant randomized multicenter study comparing kidney graft preservation with histidine–tryptophan–ketoglutarate, University of Wisconsin, and Euro-Collins solutions. The HTK Study Group. Transplant Proc 1993;25:1582–1585.
Wang WH, McNatt LG, Shepard AR, et al. Optimal procedure for extracting RNA from human ocular tissues and expression profiling of the congenital glaucoma gene FOXC1 using quantitative RT–PCR. Mol Vis 2001;7:89–94.
Luo L, Salunga RC, Guo H, et al. Gene expression profiles of laser-captured adjacent neuronal subtypes. Nat Med 1999;5:117–122.
Scheidl SJ, Nilsson S, Kalen M, et al. mRNA expression profiling of laser microbeam microdissected cells from slender embryonic structures. Am J Pathol 2002;160:801–813.
Perou CM, Sorlie T, Eisen MB, et al. Molecular portraits of human breast tumours. Nature 2000;406:747–752.
Sherlock G, Hernandez-Boussard T, Kasarskis A, et al. The Stanford Microarray Database. Nucleic Acids Res 2001;29:152–155.
Brazma A, Hingamp P, Quackenbush J, et al. Minimum information about a microarray experiment (MIAME)—toward standards for microarray data. Nat Genet 2001;29:365–371.
Mitterbauer C, Schwarz C, Hauser P, et al. Impaired tubulointerstitial expression of ET-1 and NOS isoforms in donor kidney biopsies with postischemic acute renal failure. Transplantation 2003;76:715–720.
Troyanskaya O, Cantor M, Sherlock G, et al. Missing value estimation methods for DNA microarrays. Bioinformatics 2001;17:520–525.
Alter O, Brown PO, Botstein D . Singular value decomposition for genome-wide expression data processing and modeling. Proc Natl Acad Sci USA 2000;97:10101–10106.
Eisen MB, Spellman PT, Brown PO, et al. Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci U S A 1998;95:14863–14868.
Westfall PH, Young SS . Resampling-based Multiple Testing: Examples and Methods for P-Value Adjustment. Wiley Series in Probability and Mathematical Statistics. Wiley: New York, 1993.
Ihaka R, Gentleman R . R: A language for data analysis and graphics. J Comput Graphical Stat 1996;5:299–314.
Efron B, Tibshirani R . The bootstrap method for standard errors, confidence intervals, and other measures of statistical accuracy. Stat Sci 1986;1:1–35.
Dahlquist KD, Salomonis N, Vranizan K, et al. GenMAPP, a new tool for viewing and analyzing microarray data on biological pathways. Nat Genet 2002;31:19–20.
Masys DR, Welsh JB, Lynn Fink J, et al. Use of keyword hierarchies to interpret gene expression patterns. Bioinformatics 2001;17:319–326.
Van der Hoeven JA, Lindell S, van Schilfgaarde R, et al. Donor brain death reduces survival after transplantation in rat livers preserved for 20 hr. Transplantation 2001;72:1632–1636.
van der Hoeven JA, Ploeg RJ, Postema F, et al. Induction of organ dysfunction and up-regulation of inflammatory markers in the liver and kidneys of hypotensive brain dead rats: a model to study marginal organ donors. Transplantation 1999;68:1884–1890.
Solez K, Racusen LC, Abdulkareem F, et al. Adhesion molecules and rejection of renal allografts. Kidney Int 1997;51:1476–1480.
Lu CY, Penfield JG, Kielar ML, et al. Hypothesis: is renal allograft rejection initiated by the response to injury sustained during the transplant process? Kidney Int 1999;55:2157–2168.
Terasaki PI, Cecka JM, Gjertson DW, et al. High survival rates of kidney transplants from spousal and living unrelated donors. N Engl J Med 1995;333:333–336.
Ojo AO, Hanson JA, Meier-Kriesche H, et al. Survival in recipients of marginal cadaveric donor kidneys compared with other recipients and wait-listed transplant candidates. J Am Soc Nephrol 2001;12:589–597.
Gjertson DW . Impact of delayed graft function and acute rejection on kidney graft survival. Clin Transpl 2000, 467–480.
Suri D, Meyer TW . Influence of donor factors on early function of graft kidneys. J Am Soc Nephrol 1999;10:1317–1323.
Acknowledgements
This work was supported by Austrian Science Fund FWF, Grant No. P15679 (to RO), and the NIH grant (R21-DK63127 to TWM).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hauser, P., Schwarz, C., Mitterbauer, C. et al. Genome-wide gene-expression patterns of donor kidney biopsies distinguish primary allograft function. Lab Invest 84, 353–361 (2004). https://doi.org/10.1038/labinvest.3700037
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700037
Keywords
This article is cited by
-
Validation of systems biology derived molecular markers of renal donor organ status associated with long term allograft function
Scientific Reports (2018)
-
Transplant biopsy beyond light microscopy
BMC Nephrology (2015)
-
Omics und Systembiologie
Der Nephrologe (2012)
-
Assessment of kidney organ quality and prediction of outcome at time of transplantation
Seminars in Immunopathology (2011)
-
Genomics of chronic allograft injury
Kidney International (2010)