Abstract
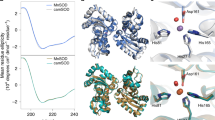
THE enzyme Cu, Zn superoxide dismutase (SOD) protects against oxidative damage by dismuting the superoxide radical O.−2 to molecular oxygen and hydrogen peroxide1–3 at the active-site Cuion4,5 in a reaction that is rate-limited by diffusion3,6 and enhanced by electrostatic guidance7–10. SOD has evolved to be one of the fastest enzymes known ( Vmax ~ 2 x 109 M−1 s−1)6,11. The new crystal structures of human SOD12 show that amino-acid site chains that are implicated in electrostatic guidance8 (Glu 132, Glu 133 and Lys 136) form a hydrogen-bonding network. Here we show that site-specific mutants that increase local positive charge while maintaining this orienting network (Glu→Gin) have faster reaction rates and increased ionic-strength dependence, matching brownian dynamics simulations incorporating electrostatic terms. Increased positive charge alone is insufficient: one charge reversal (Glu→Lys) mutant is slower than the equivalent charge neutralization (Glu→Gin) mutant, showing that the newly introduced positive charge disrupts the orienting network. Thus, electrostatically facilitated diffusion rates can be increased by design, provided the detailed structural integrity of the active-site electrostatic network is maintained.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Fridouich, I. in Advances in Enzymology Vol. 58 (ed. Meister, A.) 61–97 (Wiley, New York, 1986).
Halliwell, B. & Gutteridge, J. M. C. Free Radicals in Biology and Medicine 2nd edn (Clarendon, Oxford, 1989).
Tainer, J. A., Roberts, V. A., Fisher, C. L. Hallewell, R. A. & Getzoff, E. D. in A Study of Enzymes II. Mechanism of Enzyme Action (ed. Kuby, S. A.) 499–538 (CRC, Boca Raton, Florida, 1991).
Tainer, J. A., Getzoff, E. D., Beem, K. M., Richardson, J. S. & Richardson, D. C. J. molec. Biol. 160, 181–217 (1982).
Tainer, J. A., Getzoff, E. D., Richardson, J. S. & Richardson, D. C. Nature 306, 284–287 (1983).
Rotilio, G., Bray, R. C. & Fielden, E. M. Biochim. biophys. Acta 268, 605–609 (1972).
Koppenol, W. H. in Oxygen and Oxy-Radicals in Chemistry and Biology (eds Rodgers, M. A. J. & Powers, E. L.) 671–674 (Academic, New York, 1981).
Getzoff, E. D. et al. Nature 306, 287–290 (1983).
Sharp, K., Fine, R. & Honig, B. Science 236, 1460–1463 (1987).
Sines, J. J., Allison, S. A. & McCammon, J. A. Biochemistry 29, 9403–9412 (1990).
Klug, D., Rabani, J. & Fridovich, I. J. biol. Chem. 247, 4839–4842 (1972).
Parge, H. E., Hallewell, R. A. & Tainer, J. A. Proc. natn. Acad. Sci. U.S.A. (in the press).
Cudd, A. & Fridovich, I. J. biol. Chem. 257, 11443–11447 (1982).
Argese, E., Viglino, P., Rotilio, G., Scarpa, M. & Rigo, A. Biochemistry 26, 3224–3228 (1987).
Alberty, R. A. & Hammes, G. G. J. phys. Chem. 62, 154–159 (1958).
Allison, S. A., Bacquet, R. J. & McCammon, J. A. Biopolymers 27, 251–269 (1988).
Getzoff, E. D., Tainer, J. A., Stempien, M. M., Bell, G. I. & Hallewell, R. A. Proteins: Struct. Funct. Genet. 5, 322–336 (1989).
Banci, L. et al. Inorg. Chem. 29, 2398–2403 (1990).
Shen, J. & McCammon, J. A. J. chem. Phys. 158, 191–198 (1991).
Hallewell, R. A. et al. J. biol. Chem. 264, 5260–5268 (1989).
Lepock, J. R., Frey, H. E. & Hallewell, R. A. J. biol. Chem. 265, 21612–21618 (1990).
Hallewell, R. A. et al. Biochem. biophys. Res. Commun. 181, 474–480 (1991).
Weiner, S. J. et al. J. Am. chem. Soc. 106, 765–784 (1984).
Geysen, H. M. et al. Science 235, 1184–1190 (1987).
Schwarz, H. A. J. chem. Ed. 58, 101–105 (1981).
Steinman, H. J. biol. Chem. 262, 1882–1887 (1987).
Büchel, D. E., Gronenborn, B. & Müller-Hill, B. Nature 283, 541–545 (1980).
Hallewell, R. A. et al. Nucleic Acids Res. 13, 2017–2034 (1985).
Hanahan, D. in DNA Cloning Vol. 1 (ed. Glover, D. M.) 109–135 (IRL, Oxford, UK, 1988).
Koshland, D. & Botstein, D. Cell 20, 749–765 (1980).
Beyer, W. F. Jr, Fridovich, I., Mullenbach, G. T. & Hallewell, R. A. J. biol. Chem. 262, 11182–11187 (1987).
Davis, M. E. & McCammon, J. A. J. comp. Chem. 12, 909–912 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Getzoff, E., Cabelli, D., Fisher, C. et al. Faster superoxide dismutase mutants designed by enhancing electrostatic guidance. Nature 358, 347–351 (1992). https://doi.org/10.1038/358347a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/358347a0
This article is cited by
-
Geometric and defects engineering collaboration for enhanced cascade enzymatic nanoreactors
Nano Research (2024)
-
Nucleation and kinetics of SOD1 aggregation in human cells for ALS1
Molecular and Cellular Biochemistry (2020)
-
Computational Investigation on Electrostatic Loop Mutants Instigating Destabilization and Aggregation on Human SOD1 Protein Causing Amyotrophic Lateral Sclerosis
The Protein Journal (2019)
-
Scavenging of superoxide by a membrane-bound superoxide oxidase
Nature Chemical Biology (2018)
-
Computing disease-linked SOD1 mutations: deciphering protein stability and patient-phenotype relations
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.