Abstract
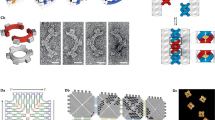
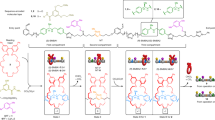
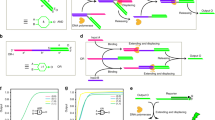
Devices that convert information from one form into another according to a definite procedure are known as automata. One such hypothetical device is the universal Turing machine1, which stimulated work leading to the development of modern computers. The Turing machine and its special cases2, including finite automata3, operate by scanning a data tape, whose striking analogy to information-encoding biopolymers inspired several designs for molecular DNA computers4,5,6,7,8. Laboratory-scale computing using DNA and human-assisted protocols has been demonstrated9,10,11,12,13,14,15, but the realization of computing devices operating autonomously on the molecular scale remains rare16,17,18,19,20. Here we describe a programmable finite automaton comprising DNA and DNA-manipulating enzymes that solves computational problems autonomously. The automaton's hardware consists of a restriction nuclease and ligase, the software and input are encoded by double-stranded DNA, and programming amounts to choosing appropriate software molecules. Upon mixing solutions containing these components, the automaton processes the input molecule via a cascade of restriction, hybridization and ligation cycles, producing a detectable output molecule that encodes the automaton's final state, and thus the computational result. In our implementation 1012 automata sharing the same software run independently and in parallel on inputs (which could, in principle, be distinct) in 120 μl solution at room temperature at a combined rate of 109 transitions per second with a transition fidelity greater than 99.8%, consuming less than 10-10 W.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Turing, A. M. On computable numbers, with an application to the Entcheidungproblem. Proc. Lond. Math. Soc. II Ser. 42, 230–265 (1936).
Hopcroft, J. E., Motwani, R. & Ullman, J. D. Introduction to Automata Theory, Languages and Computation 2nd edn (Addison-Wesley, Boston, Massachusetts, 2000).
McCulloch, W. S. & Pitts, W. A logical calculus immanent in nervous activity. Bull. Math. Biophys. 5, 115–133 (1943).
Bennet, C. H. The thermodynamics of computation—a review. Int. J. Theor. Phys. 21, 905–940 (1982).
Rothemund, P. W. K. in DNA Based Computers: Proceedings of the DIMACS Workshop, April 4, 1995, Princeton University (eds Lipton, R. J. & Baum, E. B.) 75–119 (American Mathematical Society, Providence, Rhode Island, 1996).
Smith, W. D. in DNA Based Computers: Proceedings of the DIMACS Workshop, April 4, 1995, Princeton University (eds Lipton, R. J. & Baum, E. B.) 121–185 (American Mathematical Society, Providence, Rhode Island, 1996).
Garzon, M. et al. in Automata Implementation: Lecture Notes in Computer Science 1436 (eds Wood, D. & Yu, S.) 56–74 (Springer, Berlin, 1998).
Shapiro, E. & Karunaratne, K. S. G. Method and system of computing similar to a Turing machine. US Patent 6,266,569 (2001).
Adelman, L. M. Molecular computation of solutions to combinatorial problems. Science 266, 1021–1024 (1994).
Lipton, R. J. DNA solution of hard computational problem. Science 268, 542–545 (1995).
Ouyang, Q., Kaplan, P. D., Liu, S. & Libchaber, A. DNA solution of the maximal clique problem. Science 278, 446–449 (1997).
Landweber, L. F., Lipton, R. J. & Rabin, M. O. in DNA Based Computers III: DIMACS Workshop, June 23-27, 1997, University of Pennsylvania (eds Rubin, H. & Wood, D. H.) 161–172 (American Mathematical Society, Providence, Rhode Island, 1997).
Liu, Q. et al. DNA computing on surfaces. Nature 403, 175–179 (2000).
Faulhammer, D., Cukras, A. R., Lipton, R. J. & Landweber, L. F. Molecular computation: RNA solutions to chess problems. Proc. Natl Acad. Sci. USA 97, 1385–1389 (2000).
Ruben, A. J. & Landweber, L. F. The past, present and future of molecular computing. Nature Rev. Mol. Cell Biol. 1, 69–72 (2000).
Winfree, E., Liu, F. R., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Sakamoto, K. et al. State transitions by molecules. Biosystems 52, 81–91 (1999).
Hartemink, A. J., Gifford, D. K. & Khodor, J. Automated constraint-based nucleotide sequence selection for DNA computation. Biosystems 52, 227–235 (1999).
Khodor, J. & Gifford, D. K. Design and implementation of computational systems based on programmed mutagenesis. Biosystems 52, 93–97 (1999).
Mao, C., LaBean, T. H., Reif, J. H. & Seeman, N. C. Logical computation using algorithmic self-assembly of DNA triple-crossover molecules. Nature 407, 493–496 (2000).
Chandrasegaran, S. & Smith, J. Chimeric restriction enzymes: What is next? Biol. Chem. 380, 841–848 (1999).
Acknowledgements
We thank I. Sagi and A. Yonath for use of their laboratory for our first set of experiments, and S. Shuping, A. Regev and N. Kessler for assistance and advice. E.K. is the incumbent of the Benno Gitter and Ilana Ben-Ami Chair of Biotechnology, Technion. Z.L. is the Incumbent of The Maxwell Ellis Professorial Chair in Biomedical Research. This work was supported by the Dolphi and Lola Ebner Center for Biomedical Research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Benenson, Y., Paz-Elizur, T., Adar, R. et al. Programmable and autonomous computing machine made of biomolecules. Nature 414, 430–434 (2001). https://doi.org/10.1038/35106533
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35106533
This article is cited by
-
DNA-based programmable gate arrays for general-purpose DNA computing
Nature (2023)
-
Trumpet is an operating system for simple and robust cell-free biocomputing
Nature Communications (2023)
-
A tape-reading molecular ratchet
Nature (2022)
-
Chemical-to-mechanical molecular computation using DNA-based motors with onboard logic
Nature Nanotechnology (2022)
-
Synthetic DNA applications in information technology
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.