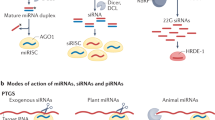
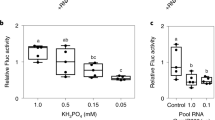
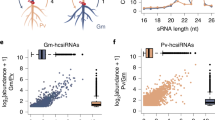
Abstract
A role for RNA as a non-cell-autonomous information macromolecule is emerging as a new model in biology. Studies on higher plants have shown the operation of cell-to-cell and long-distance communication networks that mediate the selective transport of RNA. The evolution and function of these systems are discussed in terms of an RNA-based signalling network that potentiates control over gene expression at the whole-plant level.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lehmann, R. Cell–cell signalling, microtubules, and the loss of symmetry in the Drosophila oocyte. Cell 83, 353–356 (1995).
Grünert, S. & St Johnston, D. RNA localization and the development of asymmetry during Drosophila oogenesis. Curr. Opin. Genet. Dev. 6, 395–402 (1996).
Schnapp, B. J., Arn, E. A., Deshler, J. O. & Highett, M. I. RNA localization in Xenopus oocytes. Semin. Cell Dev. Biol. 8, 529–540 (1997).
Jorgensen, R. A., Atkinson, R. G., Forster, R. L. S. & Lucas, W. J. An RNA-based information superhighway in plants. Science 279, 1486–1487 (1998).
Waterhouse, P., Wang, M.-B. & Lough, T. Gene silencing as an adaptive defence against viruses. Nature 411, 834–842 (2001).
Falke, J. J., Bass, R. B., Butler, S. L., Chervitz, S. A. & Danielson, M. A. The two-component signalling pathway of bacterial chemotaxis: a molecular view of signal transduction by receptors, kinases, and adaptation enzymes. Annu. Rev. Cell Dev. Biol. 13, 457–512 (1997).
Goudreau, P. N. & Stock, A. M. Signal transduction in bacteria: molecular mechanisms of stimulus-response coupling. Curr. Opin. Microbiol. 1, 160–169 (1998).
Stock, A. M., Robinson, V. L. & Goudreau, P. N. Two-component signal transduction. Annu. Rev. Biochem. 69, 183–215 (2000).
Artavanis-Tsakonas, S., Rand, M. D. & Lake, R. J. Notch signalling: cell fate control and signal integration in development. Science 284, 770–776 (1999).
Gurdon, J. B., Dyson, S. & St Johnson, D. Cells' perception of position in a concentration gradient. Cell 95, 159–162 (1998).
Clark, S. E. Cell signalling at the shoot meristem. Nature Reviews Mol. Cell Biol. 2, 276–284 (2001).
Goodenough, D. A., Goliger, J. A. & Paul, D. L. Connexins, connexons, and intercellular communication. Annu. Rev. Biochem. 65, 475–502 (1996).
Lucas, W. J., Ding, B. & van der Schoot, C. Plasmodesmata and the supracellular nature of plants. New Phytol. 125, 435–476 (1993).
Lucas, W. J. Plasmodesmata: intercellular channels for macromolecular transport in plants. Curr. Opin. Cell Biol. 7, 673–680 (1995).
Robards, A. W. & Lucas, W. J. Plasmodesmata. Annu. Rev. Plant Physiol. Plant Mol. Biol. 41, 369–419 (1990).
Lee, J.-Y., Yoo, B.-C. & Lucas, W. J. Parallels between nuclear-pore and plasmodesmal trafficking of information molecules. Planta 210, 177–187 (2000).
Jackson, D. & Hake, S. Morphogenesis on the move: cell-to-cell trafficking of plant regulatory proteins. Curr. Opin. Genet. Dev. 7, 495–500 (1997).
Zambryski, P. & Crawford, K. Plasmodesmata: gatekeepers for cell-to-cell transport of developmental signals in plants. Annu. Rev. Cell Dev. Biol. 16, 393–421 (2000).
Fujiwara, T., Giesman-Cookmeyer, D., Ding, B., Lommel, S. A. & Lucas, W. J. Cell-to-cell trafficking of macromolecules through plasmodesmata potentiated by the red clover necrotic mosaic virus movement protein. Plant Cell 5, 1783–1794 (1993).
Noueiry, A. O., Lucas, W. J. & Gilbertson, R. L. Two proteins of a plant DNA virus coordinate nuclear and plasmodesmal transport. Cell 76, 925–932 (1994).
Waigmann, E., Lucas, W. J., Citovsky, V. & Zambryski, P. Direct functional assay for tobacco mosaic virus cell-to-cell movement protein and identification of a domain involved in increasing plasmodesmal permeability. Proc. Natl Acad. Sci. USA 91, 1433–1437 (1994).
Fukuda, H. Tracheary element differentiation. Plant Cell 9, 1147–1156 (1997).
Schulz, A. Phloem: structure related to function. Prog. Bot. 59, 429–475 (1998).
Sjölund, R. T. The phloem sieve element: a river runs through it. Plant Cell 9, 1137–1146 (1997).
van Bel, A. J. E. & Knoblauch, M. Sieve element and companion cell: the story of the comatose patient and the hyperactive nurse. Aust. J. Plant Physiol. 27, 477–487 (2000).
Knoblauch, M., & van Bel, A. J. E. Sieve tubes in action. Plant Cell 10, 35–50 (1998).
Turgeon, R. Phloem loading and plasmodesmata. Trends Plant Sci. 1, 418–423 (1996).
Ziegler, H. Encyclopedia of Plant Physiology Vol. 1 (eds Zimmermann, M. H. & Milburn, J. A.) 59–100 (Springer, Berlin, 1975).
Kollmann, R. & Kleinig, H. Protein filaments — structural components of the phloem exudate. Planta 95, 86–94 (1970).
Kühn, C., Franceschi, V. R., Schulz, A., Lemoine, R. & Frommer, W. B. Macromolecular trafficking indicated by localization and turnover of sucrose transporters in enucleate sieve elements. Science 275, 1298–1300 (1997).
Ruiz-Medrano, R., Xoconostle-Cazares, B. & Lucas, W. J. Phloem long-distance transport of CmNACP mRNA: implications for supracellular regulation in plants. Development 126, 4405–4419 (1999).
Sasaki, T., Chino, M., Hayashi, H. & Fujiwara, T. Detection of several mRNA species in rice phloem sap. Plant Cell Physiol. 39, 895–897 (1998).
Xoconostle-Cazares, B. et al. Plant paralog to viral movement protein that potentiates transport of mRNA into the phloem. Science 283, 94–98 (1999).
Kim, M., Canio, W., Kessler, S. & Sinha, N. Developmental changes due to long-distance movement of a homeobox fusion transcript in tomato. Science 293, 287–289 (2001).
Ruiz-Medrano, R., Xoconostle-Cazares, B. & Lucas, W. J. The phloem as a conduit for inter-organ communication. Curr. Opin. Plant Biol. 4, 202–209 (2001).
Jorgensen, R. A. Cosuppression, flower color patterns, and metastable gene expression states. Science 268, 686–691 (1995).
Palauqui, J. C., et al. Frequencies, timing, and spatial patterns of co-suppression of nitrate reductase and nitrite reductase in transgenic tobacco plants. Plant Physiol. 112, 1447–1456 (1996).
Palauqui, J. C., Elmayan, T., Pollien, J. M. & Vaucheret, H. Systemic acquired silencing: transgene-specific post-transcriptional silencing is transmitted by grafting from silenced stocks to non-silenced scions. EMBO J. 16, 4738–4745 (1997).
Voinnet, O. & Baulcombe, D. C. Systemic signalling in gene silencing. Nature 389, 553 (1997).
Voinnet, O., Vain, P., Angell, S. & Baulcombe, D. C. Systemic spread of sequence-specific transgene RNA degradation in plants is initiated by localized introduction of ectopic promoterless DNA. Cell 95, 177–187 (1998).
Hull, R. The movement of viruses in plants. Annu. Rev. Phytopathol. 24, 213–240 (1989).
Gilbertson, R. L. & Lucas, W. J. How do viruses traffic on the vascular highway? Trends Plant Sci. 1, 260–268 (1996).
Carrington, J. C., Kasschau, K. D., Mahajan, S. K. & Schaad, M. C. Cell-to-cell and long-distance transport of viruses in plants. Plant Cell 8, 1669–1681 (1996).
Dolja, V. V., McBride, H. J. & Carrington, J. C. Tagging of plant potyvirus replication and movement by insertion of β-glucuronidase into the viral polyprotein. Proc. Natl Acad. Sci. USA 89, 10208–10212 (1992).
Oparka, K. J., Boevink, P. & Santa Cruz, S. Studying the movement of plant viruses using green fluorescent protein. Trends Plant Sci. 1, 412–418 (1996).
Roberts, A. G. et al. Phloem unloading in sink leaves of Nicotiana benthamiana: comparison of a fluorescent solute with a fluorescent virus. Plant Cell 9, 1381–1396 (1997).
Wang, H. L., Sudarshana, M. R., Gilbertson, R. L. & Lucas, W. J. Analysis of cell-to-cell and long-distance movement of a bean dwarf mosaic geminivirus-green fluorescent protein reporter in host and non-host species: identification of sites of resistance. Mol. Plant Microbe Interact. 12, 345–355 (1999).
Lindbo, J. A., Silva-Rosales, L., Proebsting, W. M. & Dougherty, W. G. Induction of a highly specific antiviral state in transgenic plants: implications for regulation of gene expression and virus resistance. Plant Cell 5, 1749–1759 (1993).
Ratcliff, F., Harrison, B. D. & Baulcombe, D. C. A similarity between viral defence and gene silencing in plants. Science 276, 1558–1560 (1997).
Jones, A. L., Thomas, C. L. & Maule, A. J. De novo methylation and co-suppression induced by a cytoplasmically replicating plant RNA virus. EMBO J. 17, 6385–6393 (1998).
Brigneti, G. et al. Viral pathogenicity determinants are suppressors of transgene silencing in Nicotiana benthamiana. EMBO J. 17, 6739–6746 (1998).
Beclin, C., Berthome, R., Palauqui, J. C., Tepfer, M. & Vaucheret, H. Infection of tobacco or Arabidopsis plants by CMV counteracts systemic post-transcriptional silencing of non-viral (trans)genes. Virology 252, 313–317 (1998).
Anandalakshmi, R. et al. A viral suppressor of gene silencing in plants. Proc. Natl Acad. Sci. USA 95, 13079–13084 (1998).
Voinnet, O., Lederer, C. & Baulcombe, D. C. A viral movement protein prevents spread of the gene silencing signal in Nicotiana benthamiana. Cell 103, 157–167 (2000).
Lucy, A. P., Guo, H.-S., Li, W.-X. & Ding, S.-W. Suppression of post-transcriptional gene silencing by a plant viral protein localized in the nucleus. EMBO J. 19, 1672–1680 (2000).
Mallory, A. C. et al. HC-Pro suppression of transgene silencing eliminates the small RNAs but not transgene methylation or the mobile signal. Plant Cell 13, 571–583 (2001).
Lachaud, S., Catesson, A. M. & Bonnemain, J. L. Structure and functions of the vascular cambium. C. R. Acad. Sci. III 322, 633–650 (1999).
Chaffey, N. Cambium: old challenges-new opportunities. Trees Struct. Funct. 13, 138–151 (1999).
Jansen, R.-P. mRNA localization: messages on the move. Nature Rev Mol. Cell Biol. 2, 247–256 (2001).
Bassell, G. J., Oleynikov, Y. & Singer, R. H. The travels of mRNAs through all cells large and small. FASEB J. 13, 447–454 (1999).
Schnorrer, F., Bohmann, K. & Nusslein-Volhard, C. The molecular motor dynein is involved in targeting Swallow and bicoid RNA to the anterior pole of Drosophila oocytes. Nature Cell Biol. 2, 185–190 (2000).
Heinlein, M., Epel, B. L., Padgett, H. S. & Beachy, R. N. Interaction of tobamovirus movement proteins with the plant cytoskeleton. Science 270, 1983–1985 (1995).
McLean, B. G., Zupan, J. & Zambryski, P. C. Tobacco mosaic virus movement protein associates with the cytoskeleton in tobacco cells. Plant Cell 7, 2101–2114 (1995).
Boyko, V., Ferralli, J., Ashby, J., Schellenbaum, P. & Heinlein, M. Function of microtubules in intercellular transport of plant virus RNA. Nature Cell Biol. 2, 826–832 (2000).
Oparka, K. J. & Santa Cruz, S. The great escape: phloem transport and unloading of macromolecules. Annu. Rev. Plant Physiol. Plant Mol. Biol. 51, 323–347 (2000).
Fisher, D. B., Wu, Y. & Ku, M. S. B. Turnover of soluble proteins in the wheat sieve tube. Plant Physiol. 100, 1433–1441 (1992).
Golecki, B., Schulz, A., Carstens-Behrens, U. & Kollmann, R. Evidence for graft transmission of structural phloem proteins or their precursors in heterografts of Cucurbitaceae. Planta 206, 630–640 (1998).
Golecki, B., Schulz, A. & Thompson, G. A. Translocation of structural P proteins in the phloem. Plant Cell 11, 127–140 (1999).
Kende, H. & Zeevaart, J. A. D. The five “classical” plant hormones. Plant Cell 9, 1197–1210 (1997).
Ryan, C. A. & Pearce, G. Polypeptide hormones. Plant Physiol. 125, 65–68 (2001).
Coruzzi, G. M. & Zhou, L. Carbon and nitrogen sensing and signalling in plants: emerging 'matrix effects'. Curr. Opin. Plant Biol. 4, 247–253 (2001).
Jang, J. C. & Sheen, J. Sugar sensing in higher plants. Trends Plant Sci. 2, 208–214 (1997).
Matzke, M. A., Matzke, A. J., Pruss, G. J. & Vance, V. B. RNA-based silencing strategies in plants. Curr. Opin. Genet. Dev. 11, 221–227 (2001).
Meins, F. Jr. RNA degradation and models for post-transcriptional gene-silencing. Plant Mol. Biol. 43, 261–273 (2000).
Baulcombe, D. C. Gene silencing: RNA makes RNA makes no protein. Curr. Biol. 9, R599–R601 (1999).
Vaucheret, H. & Fagard, M. Transcriptional gene silencing in plants: targets, inducers and regulators. Trends Genet. 17, 29–35 (2001).
Fire, A. RNA-triggered gene silencing. Trends Genet. 15, 358–363 (1999).
Eddy, S. R. Noncoding RNA genes. Curr. Opin. Genet. Dev. 9, 695–699 (1999).
Pasquinelli, A. E. et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 408, 86–89 (2000).
Slack, F. J. et al. The lin-41 RBCC gene acts in the C. elegans heterochronic pathway between the let-7 regulatory RNA and the LIN-29 transcription factor. Mol. Cell 5, 659–669 (2000).
Reinhart, B. J. et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403, 901–906 (2000).
Rinne, P. L. H. & van der Schoot, C. Symplasmic fields in the tunica of the shoot apical meristem coordinate morphogenetic events. Development 125, 1477–1485 (1998).
van der Schoot, C. & Rinne, P. L. Networks for shoot design. Trends Plant Sci. 4, 31–37 (1999).
Lucas, W. J. et al. Selective trafficking of KNOTTED1 homeodomain protein and its mRNA through plasmodesmata. Science 270, 1980–1983 (1995).
Carpenter, R. & Coen, E. S. Transposon induced chimeras show that FLORICAULA, a meristem identity gene, acts non-autonomously between cell-layers. Development 121, 19–26 (1995).
Hantke, S. S., Carpenter, R. & Coen, E. S. Expression of FLORICAULA in single-cell layers of periclinal chimeras activates downstream homeotic genes in all layers of floral meristems. Development 121, 27–35 (1995).
Perbal, M. C., Haughn, G., Saedler, H. & Schwarz-Sommer, Z. Non-cell-autonomous function of the Antirrhinum floral homeotic proteins DEFICIENS and GLOBOSA is exerted by their polar cell-to-cell trafficking. Development 122, 3433–3441 (1996).
Kragler, F., Monzer, J., Xoconostle-Cazares, B. & Lucas, W. J. Peptide antagonists of the plasmodesmal macromolecular trafficking pathway. EMBO J. 19, 2856–2868 (2000).
Kragler, F., Monzer, J., Shash, K., Xoconostle-Cazares, B. & Lucas, W. J. Cell-to-cell transport of proteins: requirement for unfolding and characterization of binding to a putative plasmodesmal receptor. Plant J. 15, 367–381 (1998).
Sessions, A., Yanofsky, M. F. & Weigel, D. Cell–cell signalling and movement by the floral transcription factors LEAFY and APETALA1. Science 289, 779–781 (2000).
Zeevaart, J. A. D. Physiology of flowering. Science 137, 723–731 (1962).
Colasanti, J. & Sundaresan, V. 'Florigen' enters the molecular age: long-distance signals that cause plants to flower. Trends Biochem. Sci. 25, 236–240 (2000).
Aukerman, M. J. & Amasino, R. M. Floral induction and florigen. Cell 93, 491–494 (1998).
Mezitt, L. A. & Lucas, W. J. Plasmodesmal cell-to-cell transport of proteins and nucleic acids. Plant Mol. Biol. 32, 251–273 (1996).
Lake, J. A., Quick, W. P., Beerling, D. J. & Woodward, F. I. Signals from mature to new leaves. Nature 411, 154 (2001).
Mette, M. F., Aufsatz, W., van der Winden, J., Matzke, M. A. & Matzke A. J. M. Transcriptional silencing and promoter methylation triggered by double-stranded RNA. EMBO. J. 19, 5194–5201 (2000).
Jones, L., Ratcliff, F. & Baulcombe, D. C. RNA-directed transcriptional gene silencing in plants can be inherited independently of the RNA trigger and requires Met1 for maintenance. Curr. Biol. 11, 747–757 (2001).
Amedeo, P., Habu, Y., Afsar, K., Mittelsten Scheid, O. & Paszkowski, J. Disruption of the plant gene MOM releases transcriptional silencing of methylated genes. Nature 405, 203–206 (2000).
Matzke, M., Matzke, A. & Kooter, J. M. RNA: guiding gene silencing. Science 293, 1080–1083 (2001).
Wolf, S., Deom, C. M., Beachy, R. N. & Lucas, W. J. Movement protein of tobacco mosaic virus modifies plasmodesmatal size exclusion limit. Science 246, 377–379 (1989).
Acknowledgements
We apologize to all of our colleagues whose work was not properly discussed due to space limitations. Work in our laboratory on plasmodesmata and the supracellular nature of plants was supported by grants from the National Science Foundation and the Department of Energy Office of Basic Energy Sciences.
Author information
Authors and Affiliations
Corresponding author
Related links
Glossary
- ALLOPOLYPLOIDY
-
The fusion of two distinct parental species to form a hybrid, the genome of which is the sum of the two parental genomes.
- ANGIOSPERMS
-
Any flowering vascular plant with seeds that are covered and protected by a carpel. Non-angiosperms include gymnosperms (for example, pine and cycads), ferns and mosses.
- CAMBIUM
-
A meristem that gives rise to radial rows of cells.
- COMPANION CELLS
-
These are specialized parenchymal cells that are associated with sieve-tube elements in angiosperm phloem. These cells arise from the same mother cell as the sieve-tube elements.
- ENDOREDUPLICATION
-
Repeated rounds of DNA replication in the absence of intervening mitoses, which lead to polyploidy.
- EPIGENETIC PROCESS
-
Any heritable influence (in the progeny of cells or of individuals) on gene activity, which is unaccompanied by a change in DNA sequence.
- GAP JUNCTION
-
A communicating junction (permeant to molecules up to 1 kDa) between adjacent animal cells, which is composed of 12 connexin protein subunits, six of which form a connexon or hemichannel contributed by each of the coupled cells.
- NON-CELL-AUTONOMOUS
-
A gene whose product (either RNA or protein) affects cells that are distant from the site of production.
- RIBONUCLEOPROTEIN
-
A conjugated protein that contains RNA as the non-protein component.
- SIEVE ELEMENTS
-
The cells of the phloem that are involved in the long-distance transport of nutrients.
- TRACHEID
-
An elongated, thick-walled cell of xylem, which conducts and supports, and is found in most vascular plants. In contrast to vessel elements, tracheids have tapering ends and pitted walls without perforations.
- VASCULAR MERISTEM
-
A cylindrical sheath of undifferentiated cells, which divide to produce secondary phloem and secondary xylem.
- VEGETATIVE STATE
-
The state relating to, or involving propagation by, asexual processes.
- VESSEL MEMBER
-
A constituent cell of a vessel.
Rights and permissions
About this article
Cite this article
Lucas, W., Yoo, BC. & Kragler, F. RNA as a long-distance information macromolecule in plants. Nat Rev Mol Cell Biol 2, 849–857 (2001). https://doi.org/10.1038/35099096
Issue Date:
DOI: https://doi.org/10.1038/35099096
This article is cited by
-
Grafting-induced transcriptome changes and long-distance mRNA movement in the potato/Datura stramonium heterograft system
Horticulture, Environment, and Biotechnology (2022)
-
Rhizobial infection triggers systemic transport of endogenous RNAs between shoots and roots in soybean
Science China Life Sciences (2020)
-
Transcriptomic analysis of interstock-induced dwarfism in Sweet Persimmon (Diospyros kaki Thunb.)
Horticulture Research (2019)
-
Ιntra-species grafting induces epigenetic and metabolic changes accompanied by alterations in fruit size and shape of Cucurbita pepo L.
Plant Growth Regulation (2019)
-
Vascular-mediated signalling involved in early phosphate stress response in plants
Nature Plants (2016)