Abstract
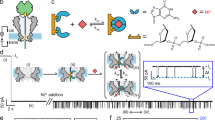
Sensory systems use a variety of membrane-bound receptors, including responsive ion channels, to discriminate between a multitude of stimuli. Here we describe how engineered membrane pores can be used to make rapid and sensitive biosensors with potential applications that range from the detection of biological warfare agents to pharmaceutical screening. Notably, use of the engineered pores in stochastic sensing, a single-molecule detection technology, reveals the identity of an analyte as well as its concentration.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Hladky, S. B. & Haydon, D. A. Discreteness of conductance change in biomolecular lipid membranes in the presence of certain antibiotics. Nature 225, 451–453 (1970).
Bayley, H., Braha, O. & Gu, L.-Q. Stochastic sensing with protein pores. Adv. Mater. 12, 139–142 (2000).
Bayley, H. & Martin, C. R. Resistive-pulse sensing: from microbes to molecules. Chem. Rev. 100, 2575–2594 (2000).
Hume, R. I., Role, L. W. & Fischbach, G. D. Acetylcholine release from growth cones detected with patches of acetylcholine receptor-rich membranes. Nature 305, 632–634 (1983).
Young, S. H. & Poo, M. Spontaneous release of transmitter from growth cones of embryonic cells. Nature 305, 634–637 (1983).
Braha, O. et al. Designed protein pores as components for biosensors. Chem. Biol. 4, 497–505 (1997).
Trivedi, B. & Kramer, R. H. Real-time patch cram detection of intracellular cGMP reveals long-term suppression of responses to NO and muscarinic agents. Neuron 21, 895–906 (1998).
Cheley, S., Braha, O., Lu, X., Conlan, S. & Bayley, H. A functional protein pore with a “retro” transmembrane domain. Protein Sci. 8, 1257–1267 (1999).
Braha, O. et al. Simultaneous stochastic sensing of divalent metal ions. Nature Biotechnol. 17, 1005–1007 (2000).
Gu, L.-Q., Braha, O., Conlan, S., Cheley, S. & Bayley, H. Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter. Nature 398, 686–690 (1999).
Sanchez-Quesada, J., Ghadiri, M. R., Bayley, H. & Braha, O. Cyclic peptides as molecular adapters for a pore-forming protein. J. Am. Chem. Soc. 122, 11758–11766 (2000).
Gu, L.-Q., Cheley, S. & Bayley, H. Capture of a single molecule in a nanocavity. Science 291, 636–640 (2001).
Kasianowicz, J. J., Brandin, E., Branton, D. & Deamer, D. W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl Acad. Sci. USA 93, 13770–13773 (1996).
Akeson, M., Branton, D., Kasianowicz, J. J., Brandin, E. & Deamer, D. W. Microsecond time-scale discrimination among polycytidylic acid, polyadenylic acid and polyuridylic acid as homopolymers or as segments within single RNA molecules. Biophys. J. 77, 3227–3233 (1999).
Meller, A., Nivon, L., Brandin, E., Golovchenko, J. & Branton, D. Rapid nanopore discrimination between single polynucleotide molecules. Proc. Natl Acad. Sci. USA 97, 1079–1084 (2000).
Henrickson, S. E., Misakian, M., Robertson, B. & Kasianowicz, J. J. Driven DNA transport into an asymmetric nanometer-scale pore. Phys. Rev. Lett. 85, 3057–3060 (2000).
Deamer, D. W. & Akeson, M. Nanopores and nucleic acids: prospects for ultrarapid sequencing. Trends Biotechnol. 18, 147–151 (2000).
Bezrukov, S. M. Ion channels as molecular Coulter counters to probe metabolite transport. J. Membr. Biol. 174, 1–13 (2000).
Vercoutere, W. et al. Rapid discrimination among individual DNA hairpin molecules at single-nucleotide resolution using an ion channel. Nature Biotechnol. 19, 248–252 (2001).
Howorka, S., Cheley, S. & Bayley, H. Sequence-specific detection of individual DNA strands using engineered nanopores. Nature Biotechnol. 19, 636–639 (2001).
Movileanu, L., Howorka, S., Braha, O. & Bayley, H. Detecting protein analytes that modulate transmembrane movement of a polymer chain within a single protein pore. Nature Biotechnol. 18, 1091–1095 (2000).
Hirn, R., Bayerl, T. M., Rädler, J. O. & Sackmann, E. Collective membrane motions of high and low amplitude, studied by dynamic light scattering and micro-interferometry. Faraday Disc. 111, 17–30 (1998).
Schmidt, C., Mayer, M. & Vogel, H. A chip-based biosensor for the functional analysis of single ion channels. Angew. Chem. Int. Edn Engl. 39, 3137–3140 (2000).
McGeoch, J. E. M., McGeoch, M. W., Carter, D. J. D., Shuman, R. F. & Guidotti, G. Biological-to-electronic interface with pores of ATP synthase subunit c in silicon nitride barrier. Med. Biol. Eng. Comput. 38, 113–119 (2000).
Sackmann, E. Supported membranes: scientific and practical applications. Science 271, 43–48 (1996).
Sackmann, E. & Tanaka, M. Supported membranes on soft polymer cushions: fabrication, characterization and applications. Trends Biotechnol. 18, 58–64 (2000).
Glazier, S. A. et al. Reconstitution of the pore-forming toxin α-hemolysin in phospholipid/18-octadecyl-1-thiahexa(ethylene oxide) and phospholipid/n-octadecanethiol supported bilayer membranes. Langmuir 16, 10428–10435 (2000).
Cornell, B. A. et al. A biosensor that uses ion-channel switches. Nature 387, 580–583 (1997).
Stora, T., Lakey, J. H. & Vogel, H. Ion-channel gating in transmembrane receptor proteins: functional activity in tethered lipid membranes. Angew. Chem. Int. Edn Engl. 38, 389–392 (1999).
Lucas, S. W. & Harding, M. M. Detection of DNA vnia an ion channel switch biosensor. Analyt. Biochem. 282, 70–79 (2000).
Straub, B., Meyer, E. & Fromherz, P. Recombinant maxi-K channels on transistor, a prototype of iono-electronic interfacing. Nature Biotechnol. 19, 121–124 (2001).
Wiegand, G., Neumaier, K. R. & Sackmann, E. Fast impedance spectroscopy: general aspects and performance study for single ion channel measurements. Rev. Sci. Instrum. 71, 2309–2320 (2000).
Tonucci, R. J., Justus, B. L., Campillo, A. J. & Ford, C. E. Nanochannel array glass. Science 258, 783–785 (1992).
Jirage, K. B., Hulteen, J. C. & Martin, C. R. Nanotubule-based molecular-filtration membranes. Science 278, 655–658 (1997).
Schuster, B., Pum, D., Braha, O., Bayley, H. & Sleytr, U. B. Self-assembled α-hemolysin pores in an S-layer supported lipid bilayer. Biochim. Biophys. Acta 1370, 280–288 (1998).
Schuster, B. et al. S-layer ultrafiltration membranes: a new support for stabilizing functionalized lipid membranes. Langmuir 17, 499–503 (2001).
Ide, T. & Yanagida, T. An artificial lipid bilayer formed on an agarose-coated glass for simultaneous electrical and optical measurement of single ion channels. Biochem. Biophys. Res. Commun. 265, 595–599 (1999).
Boxer, S. G. Molecular transport and organization in supported bilayer membranes. Curr. Opin. Chem. Biol. 4, 704–709 (2000).
Cremer, P. S. & Yang, T. Creating spatially addressed arrays of planar supported fluid phospholipid membranes. J. Am. Chem. Soc. 121, 8130–8131 (1999).
Yang, T., Simanek, E. E. & Cremer, P. S. Creating addressable aqueous microcompartments above solid supported bilayers using lithographically patterned PDMS molds. Analyt. Chem. 72, 2587–2589 (2000).
Figeys, D. & Pinto, D. Lab-on-a-chip: a revolution in biological and medical sciences. Analyt. Chem. 72, 330A–335A (2000).
Yang, T., Jung, S., Mao, H. & Cremer, P. S. Fabrication of phospholipid bilayer-coated microchannels for on-chip immunoassays. Analyt. Chem. 73, 165–169 (2001).
Dickinson, T. A., White, J., Kauer, J. S. & Walt, D. R. A chemical-detecting system based on a cross-reactive optical sensor array. Nature 382, 687–700 (1996).
Kodadek, T. Protein microarrays: prospects and problems. Chem. Biol. 8, 105–115 (2001).
Borman, S. Multivalency: strength in numbers. Chem. Eng. News 48–53 (9 October 2000).
Kramer, R. H. Patch cramming: monitoring intracellular messengers in intact cells with membrane patches containing detector ion channels. Neuron 4, 335–341 (1990).
Hulteen, J. C., Jirage, K. B. & Martin, C. R. Introducing chemical transport selectivity into gold nanotubule membranes. J. Am. Chem. Soc. 120, 6603–6604 (1998).
Sun, L. & Crooks, R. M. Single carbon nanotube membranes: a well-defined model for studying mass transport through nanoporous materials. J. Am. Chem. Soc. 122, 12340–12345 (2000).
Li, J. et al. Ion beam sculpting on the nanoscale. Nature 412, 166–169 (2001).
Weiss, S. Fluorescence spectroscopy of single biomolecules. Science 283, 1676–1683 (1999).
Xie, X. S. & Lu, H. P. Single-molecule enzymology. J. Biol. Chem. 274, 15967–15970 (1999).
Mehta, A. D., Rief, M., Spudich, J. A., Smith, D. A. & Simmons, R. M. Single-molecule biomechanics with optical methods. Science 283, 1689–1695 (1999).
Fisher, T. E. et al. Single molecule force spectroscopy of modular proteins in the nervous system. Neuron 27, 435–446 (2000).
Ball, P. Life's lessons in design. Nature 409, 413–416 (2001).
Acknowledgements
The investigation of stochastic sensors in the Bayley laboratory has been supported by DARPA, DOE, NIH, ONR and the Texas Advanced Technology Program. The development of membrane arrays in the Cremer laboratory has been supported by ONR, ARO, ACS-PRF, 3M Corporation and the Texas Advanced Technology Program. We thank the members of our laboratories for their energetic pursuit of these projects, and D. Deamer and J. Kauer for comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bayley, H., Cremer, P. Stochastic sensors inspired by biology. Nature 413, 226–230 (2001). https://doi.org/10.1038/35093038
Issue Date:
DOI: https://doi.org/10.1038/35093038
This article is cited by
-
High-resolution discrimination of homologous and isomeric proteinogenic amino acids in nanopore sensors with ultrashort single-walled carbon nanotubes
Nature Communications (2023)
-
Stochastic microsensors based on modified graphene for pattern recognition of maspin in biological samples
Analytical and Bioanalytical Chemistry (2022)
-
2D disposable stochastic sensors for molecular recognition and quantification of maspin in biological samples
Microchimica Acta (2022)
-
DNA sequencing: an overview of solid-state and biological nanopore-based methods
Biophysical Reviews (2022)
-
3D stochastic microsensors for molecular recognition and determination of heregulin-α in biological samples
Analytical and Bioanalytical Chemistry (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.