Key Points
-
Classical yeast genetics offers unique tools for discovering gene function in two evolutionarily diverse unicellular organisms: the budding yeast Saccharomyces cerevisiae and the fission yeast Schizosaccharomyces pombe.
-
The tools of classical genetics provide complementary strategies to take advantage of completed genome sequences.
-
Genetic approaches begin with isolation of mutants that affect the process of interest.
-
Synthetic enhancement, including suppression and synthetic lethality screens, allow definition of genetic networks starting from a single mutant allele.
-
Genetic interactions also lead to testable predictions about physical interactions.
-
Plasmid-based screens and functional tests facilitate the characterization of previously cloned genes and the isolation of novel alleles.
Abstract
Understanding the biology of complex systems is facilitated by comparing them with simpler organisms. Budding and fission yeasts provide ideal model systems for eukaryotic cell biology. Although they differ from one another in terms of a range of features, these yeasts share powerful genetic and genomic tools. Classical yeast genetics remains an essential element in discovering and characterizing the genes that make up a eukaryotic cell.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Goffeau, A. et al. Life with 6000 genes. Science 274, 546–567 (1996).
Beggs, J. D. Transformation of yeast by a replicating hybrid plasmid. Nature 275, 104–109 (1978).
Rothstein, R. One step gene disruption in yeast. Methods Enzymol. 101, 202–211 (1983).
Oliver, S. G. From gene to screen with yeast. Curr. Opin. Genet. Dev. 7, 405–409 (1997).
Oliver, S. G., Winson, M. K., Kell, D. B. & Banganz, F. Systematic functional analysis of the yeast genome. Trends Biotechnol. 16, 373–378 (1998).
Sipiczki, M. Phylogenesis of fission yeasts — contradictions surrounding the origin of a century old genus. Antonie Van Leeuwenhoek 68, 119–149 (1995).
Paquin, B. et al. The fungal mitochondrial genome project: evolution of fungal mitochondrial genomes and their gene expression. Curr. Genet. 31, 380–395 (1997).
Berbee, M. L. & Taylor, J. W. Dating the evolutionary radiations of the true fungi. Can. J. Bot. 71, 1114–1127 (1993).
Keogh, R. S., Seoighe, C. & Wolfe, K. H. Evolution of gene order and chromosome number in Saccharomyces, Kluyveromyces and related fungi. Yeast 14, 443–457 (1998).
Forsburg, S. L. The best yeast. Trends Genet. 15, 340–344 (1999).Summarizes some differences in the biology of the two yeast species.
Guthrie, C. & Fink, G. R. (eds) Guide to yeast genetics and molecular biology. Methods Enzymol. 194, 1–863 (1991).Describes more specific methods and protocols for both yeast species.
Moreno, S., Klar, A. & Nurse, P. Molecular genetic analysis of the fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194, 795–823 (1991).
Kumar, A. & Snyder, M. Emerging technologies in yeast genomics. Nature Rev. Genet. 2, 302–312 (2001).The genomics revolution complements the classical genetics approach.
Hoffman, C. S. & Welton, R. Mutagenesis and gene cloning in Schizosaccharomyces pombe using nonhomologous plasmid integration and rescue. Biotechniques 28, 532–539 (2000).
Chua, G., Taricani, L., Strangle, W. & Young, P. G. Insertional mutagenesis based on illegitimate recombination in Schizosaccharomyces pombe. Nucleic Acids Res. 28, E53 (2000).
Grallert, B., Nurse, P. & Patterson, T. E. A study of integrative transformation in Schizosaccharomyces pombe. Mol. Gen. Genet. 238, 26–32 (1993).
Hughes, T. R. et al. Widespread aneuploidy revealed by DNA microarray expression profiling. Nature Genet. 25, 333–337 (2000).
Wolfe, K. H. & Shields, D. C. Molecular evidence for an ancient duplication of the entire yeast genome. Nature 387, 708–713 (1997).
Giaever, G. et al. Genomic profiling of drug sensitivities via induced haploinsufficiency. Nature Genet. 21, 278–283 (1999).
Hartwell, L., Culotti, J. & Reid, B. Genetic control of the cell division cycle in yeast. I. Detection of mutants. Proc. Natl Acad. Sci. USA 66, 352–359 (1970).
Nurse, P. Genetic control of cell size at cell division in yeast. Nature 256, 547–551 (1975).
Nurse, P. Universal control mechanism regulating onset of M phase. Nature 344, 503–508 (1990).
Enoch, T., Carr, A. M. & Nurse, P. Fission yeast genes involved in coupling mitosis to completion of DNA replication. Genes Dev. 6, 2035–2046 (1992).
Li, R. & Murray, A. W. Feedback control of mitosis in budding yeast. Cell 66, 519–532 (1991).
Hoyt, M. A., Totis, L. & Roberts, B. T. S. cerevisiae genes required for cell cycle arrest in response to loss of microtubule function. Cell 66, 507–517 (1991).
Koshland, D., Kent, J. C. & Hartwell, L. H. Genetic analysis of the mitotic transmission of minichromosomes. Cell 40, 393–403 (1985).
Hieter, P., Mann, C., Snyder, M. & Davis, R. W. Mitotic stability of yeast chromosomes: a colony color assay that measures nondisjunction and chromosome loss. Cell 40, 381–382 (1985).
Rine, J. Gene overexpression in studies of Saccharomyces cerevisiae. Methods Enzymol. 194, 239–251 (1991).
Stevenson, L. F., Kennedy, B. K. & Harlow, E. A large-scale overexpression screen in Saccharomyces cerevisiae identifies previously uncharacterized cell cycle genes. Proc. Natl Acad. Sci. USA 98, 3946–3951 (2001).
Herskowitz, I. Functional inactivation of genes by dominant negative mutations. Nature 329, 219–222 (1987).
Prelich, G. Suppression mechanisms: themes from variations. Trends Genet. 15, 261–266 (1999).
Guarente, L. Synthetic enhancement in gene interaction: a genetic tool come of age. Trends Genet. 9, 362–366 (1993).References 28 and 30 are classic reviews describing the main features of the screens covered in this article and providing some theoretical background.
Booher, R. & Beach, D. Interaction between cdc13+ and cdc2+ in the control of mitosis in fission yeast: dissociation of the G1 and G2 roles of the cdc2+ protein kinase. EMBO J. 6, 3441–3447 (1987).
Sandrock, T. M., O'Dell, J. L. & Adams, A. E. Allele-specific suppression by formation of new protein–protein interactions in yeast. Genetics 147, 1635–1642 (1997).
Tye, B. K. & Sawyer, S. The hexameric eukaryotic MCM helicase: building symmetry from nonidentical parts. J. Biol. Chem. 275, 34833–34836 (2000).
Hardy, C. F. J., Dryga, O., Seematter, S., Pahl, P. M. B. & Sclafani, R. A. mcm5/cdc46-bob1 bypasses the requirement for the S phase activator Cdc7p. Proc. Natl Acad. Sci. USA 94, 3151–3155 (1997).
Lee, M. G. & Nurse, P. Complementation used to clone a human homologue of the fission yeast cell cycle control gene cdc2. Nature 327, 31–35 (1987).
Grallert, B. & Nurse, P. An approach to identify functional homologues and suppressors of genes in fission yeast. Curr. Genet. 32, 27–31 (1997).
Wittenberg, C., Sugimoto, K. & Reed, S. I. G1-specific cyclins of S. cerevisiae: cell cycle periodicity, regulation by mating pheromone and association with the p34CDC28 protein kinase. Cell 62, 225–237 (1990).
Hennessy, K. M., Lee, A., Chen, E. & Botstein, D. A group of interacting yeast DNA replication genes. Genes Dev. 5, 958–969 (1991).
Kroll, E. S., Hyland, K. M., Hieter, P. & Li, J. J. Establishing genetic interactions by a synthetic dosage lethality phenotype. Genetics 143, 95–102 (1996).
Pasion, S. G. & Forsburg, S. L. Nuclear localization of fission yeast Mcm2/Cdc19p requires MCM complex formation. Mol. Biol. Cell 10, 4043–4057 (1999).
Clarke, L. Centromeres: proteins, protein complexes, and repeated domains at centromeres of simple eukaryotes. Curr. Opin. Genet. Dev. 8, 212–218 (1998).
Sikorski, R. S. & Boeke, J. D. In vitro mutagenesis and plasmid shuffling: from cloned gene to mutant yeast. Methods Enzymol. 194, 302–318 (1991).
Liang, D. T. & Forsburg, S. L. Characterization of S. pombe mcm7+ and cdc23+ (MCM10) and interactions with replication checkpoints. Genetics (in the press).
Johnston, M. The yeast genome: on the road to the Golden Age. Curr. Opin. Genet. Dev. 10, 617–623 (2000).
Mullen, J. R., Kaliaraman, V., Ibrahim, S. S. & Brill, S. J. Requirement for three novel protein complexes in the absence of the Sgs1 DNA helicase in Saccharomyces cerevisiae. Genetics 157, 103–118 (2001).An excellent example of a synthetic lethal screen and plasmid shuffle analysis.
Acknowledgements
I thank L. Pillus for helpful comments on the manuscript. I am a scholar of the Leukemia Society of America. Support in my lab comes from the National Institutes of Health, the National Science Foundation and the American Cancer Society.
Author information
Authors and Affiliations
Related links
Related links
DATABASE LINKS
YEAST PHYLOGENY
GENOME DATABASES
Saccharomyces Genome Database (SGD)
Sanger Centre S. pombe database
PROTOCOLS
OTHER YEAST RESOURCES
Saccharomyces genome deletion project
LABS AND INVESTIGATORS
COMMUNITY INFORMATION
Glossary
- ASCOMYCETE
-
Free-living fungus that reproduces sexually through the formation of spores packaged in a sac called an ascus. Some taxonomists include non-sexually reproducing fungi with DNA sequences, which indicates a close degree of relatedness.
- COMPLEMENTATION TEST
-
Determines whether two recessive mutations are in the same functional unit or gene. Two recessive mutant strains, a1 and a2, crossed together complement each other if the resulting diploid has a wild-type phenotype; as each provides the function missing in the other, they are assumed to affect independent genes. If, instead, the diploid has the mutant phenotype, then a1 and a2 do not complement and are assumed to affect the same gene.
- RECOMBINATION
-
Any process in a diploid or partially diploid cell that generates new gene or chromosomal combinations not found in that cell or in its progenitors. At meiosis, recombination (or crossing over) is the process of reciprocal exchange between homologous chromosomal segments that generates a haploid product genotypically distinct from the two haploid genotypes of the original meiotic diploid.
- EPISTASIS
-
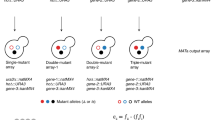
The phenotype caused by a mutation in one gene is masked by a mutation in another gene. Epistatic analysis requires that two mutants have distinguishable phenotypes. It can be used to determine the order of gene function by testing whether the phenotype of the double mutant ab is similar to that of mutant a, or mutant b.
- HETEROLOGOUS RECOMBINATION
-
Recombination between DNA molecules with significantly different sequences, for example when a transgenic construct integrates randomly in the genome.
- HAPLOINSUFFICIENCY
-
A gene dosage effect that occurs when a diploid requires both functional copies of a gene for a wild-type phenotype. An organism that is heterozygous for a haploinsufficient locus does not have a wild-type phenotype.
- REPLICA PLATING
-
A classic method to duplicate the colonies on an agar plate by stamping them on sterile velvets or filters, and then applying these copies to new (replica) plates. The replica plates can then be used to test the colonies for growth on different nutrient media or at different temperatures.
- HIGH-COPY LIBRARY
-
Most plasmid episomes in yeast are present at greater than one copy per cell, leading to an increased dosage of any gene(s) carried by the plasmid. The high copy dosage effect can be enhanced if the cells are transformed with a library that contains cDNAs expressed by strong promoters.
- EPISOME
-
An independent DNA element, such as a plasmid, that can replicate extrachromosomally or that can be maintained by integrating into the genome of the host.
- DOMINANT NEGATIVE
-
A mutant allele that interferes with the function of its wild-type version.
- SYNTHETIC INTERACTIONS
-
These occur when a double mutant has a phenotype different from either single mutant parent. For suppressors (synthetic viable), the double mutant is viable when at least one of the single mutants is not. For synthetic lethal mutants, the double mutant is inviable under conditions in which both parents are viable.
Rights and permissions
About this article
Cite this article
Forsburg, S. The art and design of genetic screens: yeast. Nat Rev Genet 2, 659–668 (2001). https://doi.org/10.1038/35088500
Issue Date:
DOI: https://doi.org/10.1038/35088500
This article is cited by
-
From beer to breadboards: yeast as a force for biological innovation
Genome Biology (2024)
-
Engineered cell differentiation and sexual reproduction in probiotic and mating yeasts
Nature Communications (2022)
-
Trehalose accumulation and radiation resistance due to prior heat stress in Saccharomyces cerevisiae
Archives of Microbiology (2022)
-
High-throughput synthetic rescue for exhaustive characterization of suppressor mutations in human genes
Cellular and Molecular Life Sciences (2020)
-
Current advances in haploid stem cells
Protein & Cell (2020)