Abstract
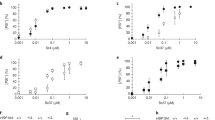
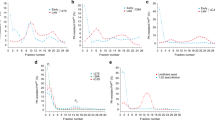
A perplexing feature of prion-based inheritance is that prions composed of the same polypeptide can evoke different phenotypes (such as distribution of brain lesions), even when propagated in genetically identical hosts1,2,3. The molecular basis of this strain diversity and the relationship between strains and barriers limiting transmission between species remain unclear. We have used the yeast prion phenomenon [PSI+]4 to investigate these issues and examine the role that conformational differences1,2,5 may have in prion strains. We have made a chimaeric fusion between the prion domains of two species (Saccharomyces cerevisae and Candida albicans) of Sup35, the protein responsible for [PSI+]4,6. Here we report that this chimaera forms alternate prion strains in vivo when initiated by transient overexpression of different Sup35 species. Similarly, in vitro the purified chimaera, when seeded with different species of Sup35 fibres, establishes and propagates distinct amyloid conformations. These fibre conformations dictate amyloid seeding specificity: a chimaera seeded by S. cerevisiae fibres efficiently catalyses conversion of S. cerevisiae Sup35 but not of C. albicans Sup35, and vice versa. These and other considerations1,2,5,7,8 argue that heritable prion strains result from self-propagating conformational differences within the prion protein itself. Moreover, these conformational differences seem to act in concert with the primary structure to determine a prion's propensity for transmission across a species barrier.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Prusiner, S. B. Prions. Proc. Natl Acad. Sci. USA 95, 13363–13383 (1998).
Bessen, R. A. et al. Non-genetic propagation of strain-specific properties of scrapie prion protein. Nature 375, 698–700 (1995).
Derkatch, I. L., Chernoff, Y. O., Kushnirov, V. V., Inge-Vechtomov, S. G. & Liebman, S. W. Genesis and variability of [PSI] prion factors in Saccharomyces cerevisiae. Genetics 144, 1375–1386 (1996).
Serio, T. R. & Lindquist, S. L. [PSI+]: an epigenetic modulator of translation termination efficiency. Annu. Rev. Cell. Dev. Biol. 15, 661–703 (1999).
Telling, G. C. et al. Evidence for the conformation of the pathologic isoform of the prion protein enciphering and propagating prion diversity. Science 274, 2079–2082 (1996).
Paushkin, S. V., Kushnirov, V. V., Smirnov, V. N. & Ter-Avanesyan, M. D. Propagation of the yeast prion-like [psi+] determinant is mediated by oligomerization of the SUP35-encoded polypeptide chain release factor. EMBO J. 15, 3127–3134 (1996).
Collinge, J., Sidle, K. C., Meads, J., Ironside, J. & Hill, A. F. Molecular analysis of prion strain variation and the aetiology of ‘new variant’ CJD. Nature 383, 685–690 (1996).
Collinge, J. Variant Creutzfeldt-Jakob disease. Lancet 354, 317–323 (1999).
Stansfield, I. et al. The products of the SUP45 (eRF1) and SUP35 genes interact to mediate translation termination in Saccharomyces cerevisiae. EMBO J 14, 4365-4373 (1995).
Chernoff, Y. O., Lindquist, S. L., Ono, B., Inge-Vechtomov, S. G. & Liebman, S. W. Role of the chaperone protein Hsp104 in propagation of the yeast prion-like factor [psi+]. Science 268, 880–884 (1995).
Ter-Avanesyan, M. D., Dagkesamanskaya, A. R., Kushnirov, V. V. & Smirnov, V. N. The SUP35 omnipotent suppressor gene is involved in the maintenance of the non-Mendelian determinant [psi+] in the yeast Saccharomyces cerevisiae. Genetics 137, 671–676 (1994).
DePace, A. H., Santoso, A., Hillner, P. & Weissman, J. S. A critical role for amino-terminal glutamine/asparagine repeats in the formation and propagation of a yeast prion. Cell 93, 1241–1252 (1998).
Patino, M. M., Liu, J. J., Glover, J. R. & Lindquist, S. Support for the prion hypothesis for inheritance of a phenotypic trait in yeast. Science 273, 622–626 (1996).
King, C. Y. et al. Prion-inducing domain 2-114 of yeast Sup35 protein transforms in vitro into amyloid-like filaments. Proc. Natl Acad. Sci. USA 94, 6618–6622 (1997).
Glover, J. R. et al. Self-seeded fibers formed by Sup35, the protein determinant of [PSI+], a heritable prion-like factor of S. cerevisiae. Cell 89, 811–819 (1997).
Kushnirov, V. V., Kochneva-Pervukhova, N. V., Chechenova, M. B., Frolova, N. S. & Ter-Avanesyan, M. D. Prion properties of the Sup35 protein of yeast Pichia methanolica. EMBO J. 19, 324–331 (2000).
Santoso, A., Chien, P., Osherovich, L. Z. & Weissman, J. S. Molecular basis of a yeast prion species barrier. Cell 100, 277–288 (2000).
Chernoff, Y. O. et al. Evolutionary conservation of prion-forming abilities of the yeast Sup35 protein. Mol. Microbiol. 35, 865–876 (2000).
Liu, J. J. & Lindquist, S. Oligopeptide-repeat expansions modulate ’protein-only’ inheritance in yeast. Nature 400, 573–576 (1999).
Wickner, R. B. [URE3] as an altered URE2 protein: evidence for a prion analog in Saccharomyces cerevisiae. Science 264, 566–569 (1994).
Sondheimer, N. & Lindquist, S. Rnq1: an epigenetic modifier of protein function in yeast. Mol. Cell 5, 163–172 (2000).
Tuite, M. F., Mundy, C. R. & Cox, B. S. Agents that cause a high frequency of genetic change from [psi+] to [psi-] in Saccharomyces cerevisiae. Genetics 98, 691–711 (1981).
Zhou, P. et al. The yeast non-Mendelian factor [ETA+] is a variant of [PSI+], a prion-like form of release factor eRF3. EMBO J. 18, 1182–1191 (1999).
Serio, T. R. et al. Nucleated conformational conversion and the replication of conformational information by a prion determinant. Science 289, 1317–1321 (2000).
Paushkin, S. V., Kushnirov, V. V., Smirnov, V. N. & Ter-Avanesyan, M. D. In vitro propagation of the prion-like state of yeast Sup35 protein. Science 277, 381–383 (1997).
Kochneva-Pervukhova, N. V. et al. Mechanism of inhibition of Psi+ prion determinant propagation by a mutation of the N-terminus of the yeast Sup35 protein. EMBO J. 17, 5805–5810 (1998).
Appel, T. R., Dumpitak, C., Matthiesen, U. & Riesner, D. Prion rods contain an inert polysaccharide scaffold. Biol. Chem. 380, 1295–1306 (1999).
Klein, T. R., Kirsch, D., Kaufmann, R. & Riesner, D. Prion rods contain small amounts of two host sphingolipids as revealed by thin-layer chromatography and mass spectrometry. Biol. Chem. 379, 655–666 (1998).
Kocisko, D. A. et al. Species specificity in the cell-free conversion of prion protein to protease-resistant forms: a model for the scrapie species barrier. Proc. Natl Acad. Sci. USA 92, 3923–3927 (1995).
Wilesmith, J. W., Ryan, J. B. & Atkinson, M. J. Bovine spongiform encephalopathy—Epidemiologic studies on the origin.. Vet. Rec. 128, 199–203 (1991).
Acknowledgements
We thank H. Bourne, H. Field, D. Julius, B. Panning, S. Lindquist, J. Reddy, S. Ribich, M. Scott and members of the Weissman and Lim lab for discussion and comments. This work was supported by the NIH, the Searle Scholars Program, the David and Lucile Packard Foundation, the Howard Hughes Medical Institute and a National Science Foundation Graduate Fellowship (P.C.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chien, P., Weissman, J. Conformational diversity in a yeast prion dictates its seeding specificity. Nature 410, 223–227 (2001). https://doi.org/10.1038/35065632
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35065632
This article is cited by
-
Evidence of distinct α-synuclein strains underlying disease heterogeneity
Acta Neuropathologica (2021)
-
Environment-transformable sequence–structure relationship: a general mechanism for proteotoxicity
Biophysical Reviews (2018)
-
Discovering putative prion sequences in complete proteomes using probabilistic representations of Q/N-rich domains
BMC Genomics (2013)
-
Case for an RNA–prion world: a hypothesis based on conformational diversity
Journal of Biological Physics (2011)
-
Unraveling infectious structures, strain variants and species barriers for the yeast prion [PSI+]
Nature Structural & Molecular Biology (2009)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.