Abstract
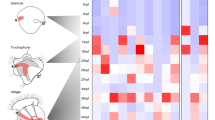
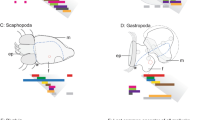
Bilateria are subdivided into Protostomia and Deuterostomia1,2. Indirect development through primary, ciliary larvae occurs in both of these branches; however, the closing blastopore develops into mouth and anus in Protostomia and into anus only in Deuterostomia. Because of this important difference in larval gut ontogeny, the tube-shaped guts in protostome and deuterostome primary larvae are thought to have evolved independently2,3. To test this hypothesis, we have analysed the expression of brachyury, otx and goosecoid homologues in the polychaete Platynereis dumerilii4, which develops by means of a trochophora larva—the primary, ciliary larva prototypic for Protostomia2. Here we show that brachyury expression in the ventral portion of the developing foregut in Platynereis and also otx expression along ciliated bands in the mouth region of the trochophora larva parallels expression in primary larvae in Deuterostomia5,6,7,8,9. In addition, goosecoid expression in the foregut of Platynereis mirrors the function in higher Deuterostomia10. We present molecular evidence for the evolutionary conservation of larval foreguts and mouth regions of Protostomia and Deuterostomia. Our data indicate that Urbilateria, the common bilaterian ancestors, developed through a primary, ciliary larva that already possessed a tripartite tube-shaped gut.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Willmer, P. Invertebrate Relationships (Cambridge Univ. Press, Cambridge, 1990).
Nielsen, C. Animal Evolution: Interrelationships of the Living Phyla (Oxford Univ. Press, Oxford, 1995).
Grobben, K. Die systematische Einteilung des Tierreichs. Verh. Zool. Bot. Ges. Wien 58, 491–511 (1908).
Dorresteijn, A. W. C., O'Grady, B., Fischer, A., Porchet-Henere, E. & Boilly-Marer, Y. Molecular specification of cell lines in the embryo of Platynereis (Annelida). Roux's Arch. Dev. Biol. 202, 264–273 (1993).
Shoguchi, E., Satoh, N. & Maruyama, Y. K. Pattern of Brachyury gene expression in starfish embryos resembles that of hemichordate embryos but not of sea urchin embryos. Mech. Dev. 82, 185–189 (1999).
Peterson, K. J., Cameron, R. A., Tagawa, K., Satoh, N. & Davidson, E. H. A comparative molecular approach to mesodermal patterning in basal deuterostomes: the expression pattern of Brachyury in the enteropneust hemichordate Ptychodera flava. Development 126, 85–95 (1999).
Tagawa, K., Humphreys, T. & Satoh, N. Novel pattern of Brachyury gene expression in hemichordate embryos. Mech. Dev. 75, 139–143 (1998).
Shoguchi, E., Harada, Y., Numakunai, T. & Satoh, N. Expression of the Otx gene in the ciliary bands during sea cucumber embryogenesis. Genesis 27, 58–63 (2000).
Harada, Y. et al. Developmental expression of the hemichordate otx ortholog. Mech. Dev. 91, 337–339 (2000).
Steinbeisser, H. & De Robertis, E. M. Xenopus goosecoid: a gene expressed in the prechordal plate that has dorsalizing activity. C. R. Acad. Sci. 316, 959–971 (1993).
Kispert, A., Herrmann, B. G., Leptin, M. & Reuter, R. Homologs of the mouse Brachyury gene are involved in the specification of posterior terminal structures in Drosophila, Tribolium, and Locusta. Genes Dev. 8, 2137–2150 (1994).
Woollard, A. & Hodgkin, J. The Caenorhabditis elegans fate-determining gene mab-9 encodes a T-box protein required to pattern the posterior hindgut. Genes Dev. 14, 596–603 (2000).
Kispert, A. & Herrmann, B. G. The Brachyury gene encodes a novel DNA binding protein. EMBO J. 12, 3211–3220 (1993).
Peterson, K. J., Cameron, R. A. & Davidson, E. H. Bilaterian origins: significance of new experimental observations. Dev. Biol. 219, 1–17 (2000).
Kusch, T. & Reuter, R. Functions for Drosophila brachyenteron and forkhead in mesoderm specification and cell signalling. Development 126, 3991–4003 (1999).
Smith, J. Brachyury and the T-box genes. Curr. Opin. Genet. Dev. 7, 474–480 (1997).
Wilson, E. B. The cell-lineage of Nereis. A contribution to the cytogeny of the annelid body. J. Morphol. 6, 361–480 (1892).
Wu, L. H. & Lengyel, J. A. Role of caudal in hindgut specification and gastrulation suggests homology between Drosophila amnioproctodeal invagination and vertebrate blastopore. Development 125, 2433–2442 (1998).
De Robertis, E. M. & Sasai, Y. A common plan for dorsoventral patterning in Bilateria. Nature 380, 37–40 (1996).
Arendt, D. & Nübler-Jung, K. Dorsal or ventral: similarities in fate maps and gastrulation patterns in annelids, arthropods and chordates. Mech. Dev. 61, 7–21 (1997).
Hirth, F. & Reichert, H. Conserved genetic programs in insect and mammalian brain development. BioEssays 21, 677–684 (1999).
Hahn, M. & Jäckle, H. Drosophila goosecoid participates in neural development but not in body axis formation. EMBO J. 15, 3077–3084 (1996).
Goriely, A. et al. A functional homologue of goosecoid in Drosophila. Development 122, 1641–1650 (1996).
Yasuo, H. & Satoh, N. Conservation of the developmental role of Brachyury in notochord formation in a urochordate, the ascidian Halocynthia roretzi. Dev. Biol. 200, 158–170 (1998).
Holland, P. W., Koschorz, B., Holland, L. Z. & Herrmann, B. G. Conservation of Brachyury (T) genes in amphioxus and vertebrates: developmental and evolutionary implications. Development 121, 4283–4291 (1995).
Haeckel, E. in Systematische Phylogenie. 2.Teil: Systematische Phylogenie der wirbellosen Thiere (Invertebrata). 259–347 (Georg Reimer, Berlin, 1896).
De Robertis, E. M. Evolutionary biology. The ancestry of segmentation. Nature 387, 25–26 (1997).
Arendt, D. & Nübler-Jung, K. Comparison of early nerve cord development in insects and vertebrates. Development 126, 2309–2325 (1999).
Strimmer, K. & Von Haeseler, A. Likelihood-mapping: A simple method to visualize phylogenetic content of a sequence alignment. Proc. Natl Acad. Sci. USA 94, 6815–6819 (1997).
Loosli, F., Köster, R. W., Carl, M., Krone, A. & Wittbrodt, J. Six3, a medaka homologue of the Drosophila homeobox gene sine oculis is expressed in the anterior embryonic shield and the developing eye. Mech. Dev. 74, 159–164 (1998).
Acknowledgements
We thank A. A. W. Dorresteijn, F. Loosli and R. Rieger for discussions; S. Cohen, B. Hobmayer, T. Holstein, T.-E. Rusten and L. Teixeira for comments on the manuscript; and members of the Wittbrodt laboratory for support. cDNA libraries were provided by C. Heimann, University of Mainz. This work was supported by a fellowship from the European Molecular Biology Organisation (EMBO) (D.A.), and by grants from the Deutsche Forschungsgemeinschaft (DFG) Schwerpunkt “Evolution entwicklungsbiologischer Prozesse” (U.T. and J.W.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Arendt, D., Technau, U. & Wittbrodt, J. Evolution of the bilaterian larval foregut. Nature 409, 81–85 (2001). https://doi.org/10.1038/35051075
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35051075
This article is cited by
-
An ancestral Wnt–Brachyury feedback loop in axial patterning and recruitment of mesoderm-determining target genes
Nature Ecology & Evolution (2022)
-
The Nereid on the rise: Platynereis as a model system
EvoDevo (2021)
-
Extensive gene rearrangements in the mitogenomes of congeneric annelid species and insights on the evolutionary history of the genus Ophryotrocha
BMC Genomics (2020)
-
Evolutionary transcriptomics of metazoan biphasic life cycle supports a single intercalation origin of metazoan larvae
Nature Ecology & Evolution (2020)
-
Gene expression analysis of three homeobox genes throughout early and late development of a feather star Anneissia japonica
Development Genes and Evolution (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.