Abstract
WE previously reported that roughly 1% of the short peptides encoded by Escherichia coli genomic DNA fragments act as transcriptional activating regions in yeast when fused to GAL4(1–147), a DNA-binding portion of the yeast transcriptional activator GAL4 (ref. 1). Struhl questioned the conclusion that we had identified new transcriptional activating sequences that function in the absence of yeast transcriptional activating sequences2. His criticism was based on two considerations: first, GAL4(1–147) contains an acidic segment (and subsequent experiments have shown that this region contains a weak activating region in vitro3); second, attempts to isolate new activating regions failed when the DNA-binding domain of a bacterial represser, LexA(l–87), was used as the DNA-binding unit2. We report here a repeat of our original experiment using the complete Lex A molecule LexA(1–202) as the DNA-binding region, instead of GAL4(1–147) or LexA(1–87). We find that, as in the original experiment, about 1% of the short peptides encoded by E. coli genomic fragments act as transcriptional activating regions when fused to intact LexA. All of the new activating regions whose sequences we determined bore an excess of acidic amino acids (see Table 1).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Ma, J. & Ptashne, M. Cell 51, 113–119 (1987).
Struhl, K. Nature 332, 649–650 (1988).
Lin, Y.-S., Carey, M. F., Ptashne, M. & Green, M. R. Cell 54, 659–664 (1988).
Carey, M., Kakidani, H., Leatherwood, J., Mostashari, F. & Ptashne, M. J. molec. Biol. 209, 423–432 (1990).
Little, J. W. & Mount, D. W. Cell 29, 11–22 (1982).
Kunkel, T. A. Proc. natn. Acad. Sci. U.S.A. 82, 488–492 (1985).
Brent, R. & Ptashne, M. Cell 43, 729–736 (1985).
Himmelfarb, H., Pearlberg, J., Last, D. & Ptashne, M. Cell 63, 1299–1309 (1990).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ruden, D., Ma, J., Li, Y. et al. Generating yeast transcriptional activators containing no yeast protein sequences. Nature 350, 250–252 (1991). https://doi.org/10.1038/350250a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/350250a0
This article is cited by
-
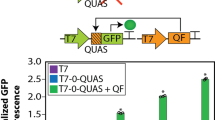
Engineering a two-gene system to operate as a highly sensitive biosensor or a sharp switch upon induction with β-estradiol
Scientific Reports (2022)
-
A novel reverse two-hybrid method for the identification of missense mutations that disrupt protein–protein binding
Scientific Reports (2020)
-
Identification of the Trans-Activation Domain and the Nuclear Location Signals of Human Zinc Finger Protein HZF1 (ZNF16)
Molecular Biotechnology (2010)
-
Genetic selection of peptide aptamers that recognize and inhibit cyclin-dependent kinase 2
Nature (1996)
-
The maize transcription factor Opaque-2 activates a wheat glutenin promoter in plant and yeast cells
Plant Molecular Biology (1995)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.