Abstract
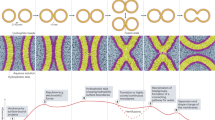
Lipid bilayer fusion is mediated by SNAREs (soluble N-ethylmaleimide-sensitive factor attachment protein receptors) located on the vesicle membrane (v-SNAREs) and the target membrane (t-SNAREs)1,2. The assembled v-SNARE/t-SNARE complex consists of a bundle of four helices, of which one is supplied by the v-SNARE and the other three by the t-SNARE3. For t-SNAREs on the plasma membrane, the protein syntaxin4 supplies one helix and a SNAP-25 protein5 contributes the other two. Although there are numerous homologues of syntaxin on intracellular membranes6, there are only two SNAP-25-related proteins in yeast, Sec9 and Spo20, both of which are localized to the plasma membrane and function in secretion7 and sporulation8, respectively. What replaces SNAP-25 in t-SNAREs of intracellular membranes? Here we show that an intracellular t-SNARE is built from a ‘heavy chain’ homologous to syntaxin and two separate non-syntaxin ‘light chains’. SNAP-25 may thus be the exception rather than the rule, having been derived from genes that encoded separate light chains that fused during evolution to produce a single gene encoding one protein with two helices.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Söllner, T. et al. SNAP receptors implicated in vesicle targeting and fusion. Nature 362, 318–324 (1993).
Weber, T. et al. SNAREpins: minimal machinery for membrane fusion. Cell 92, 759–772 ( 1998).
Sutton, R. B., Fasshauer, D., Jahn, R. & Brunger, A. T. Crystal structure of a SNARE complex involved in synaptic exocytosis at 2.4 Å resolution. Nature 395, 347–353 (1998).
Bennett, M. K., Calakos, N. & Scheller, R. H. Syntaxin: a synaptic protein implicated in docking of synaptic vesicles at presynaptic active zones. Science 257, 255–259 (1992).
Oyler, G. A. et al. The identification of a novel synaptosomal-associated protein, SNAP-25, differentially expressed by neuronal subpopulations. J. Cell Biol. 109, 3039–3052 (1989).
Pelham, H. R. SNAREs and the secretory pathway-lessons from yeast. Exp. Cell Res. 247, 1–8 ( 1999).
Rossi, G., Salminen, A., Rice, L. M., Brunger, A. T. & Brennwald, P. Analysis of a yeast SNARE complex reveals remarkable similarity to the neuronal SNARE complex and a novel function for the C terminus of the SNAP-25 homolog, Sec9. J. Biol. Chem. 272, 16610–16617 ( 1997).
Neiman, A. M. Prospore membrane formation defines a developmentally regulated branch of the secretory pathway in yeast. J. Cell Biol. 140, 29–37 (1998).
Nichols, B. J., Ungermann, C., Pelham, H. R., Wickner, W. T. & Haas, A. Homotypic vacuolar fusion mediated by t- and v-SNAREs. Nature 387, 199– 202 (1997).
Ungermann, C. & Wickner, W. Vam7p, a vacuolar SNAP-25 homolog, is required for SNARE complex integrity and vacuole docking and fusion. EMBO J. 17, 3269–3276 ( 1998).
Ungermann, C. et al. Three v-SNAREs and two t-SNAREs, present in a pentameric cis-SNARE complex on isolated vacuoles, are essential for homotypic fusion. J. Cell Biol. 145, 1435– 1442 (1999).
Sato, T. K., Darsow, T. & Emr, S. D. Vam7p, a SNAP-25-like molecule, and Vam3p, a syntaxin homolog, function together in yeast vacuolar protein trafficking. Mol. Cell. Biol. 18, 5308–5319 (1998).
Nichols, B. J. & Pelham, H. R. SNAREs and membrane fusion in the Golgi apparatus. Biochim. Biophys. Acta 1404, 9–31 (1998).
Fischer von Mollard, G., Nothwehr, S. F. & Stevens, T. H. The yeast v-SNARE Vti1p mediates two vesicle transport pathways through interactions with the t-SNAREs Sed5p and Pep12p. J. Cell Biol. 137, 1511–1524 (1997).
Struck, D. K., Hoekstra, D. & Pagano, R. E. Use of resonance energy transfer to monitor membrane fusion. Biochemistry 20, 4093– 4099 (1981).
Parlati, F. et al. Rapid and efficient fusion of phospholipid vesicles by the alpha-helical core of a SNARE complex in the absence of an N-terminal regulatory domain. Proc. Natl Acad. Sci. USA 96, 12565 –12570 (1999).
McNew, J. A. et al. Compartmental specificity of cellular membrane fusion encoded in SNARE proteins. Nature 407, 153– 159 (2000).
Peters, C. & Mayer, A. Ca2+/calmodulin signals the completion of docking and triggers a late step of vacuole fusion. Nature 396, 575–580 ( 1998).
Weimbs, T., Mostov, K., Low, S. H. & Hofmann, K. A model for structural similarity between different SNARE complexes based on sequence relationships. Trends Cell Biol. 8, 260– 262 (1998).
Fischer von Mollard, G. & Stevens, T. H. The Saccharomyces cerevisiae v-SNARE Vti1p is required for multiple membrane transport pathways to the vacuole. Mol. Biol. Cell 10, 1719 –1732 (1999).
Hay, J. C., Chao, D. S., Kuo, C. S. & Scheller, R. H. Protein interactions regulating vesicle transport between the endoplasmic reticulum and Golgi apparatus in mammalian cells. Cell 89, 149– 158 (1997).
Ungermann, C., Sato, K. & Wickner, W. Defining the functions of trans-SNARE pairs. Nature 396, 543–548 ( 1998).
Weber, T. et al. SNAREpins are functionally resistant to disruption by NSF and alpha-SNAP. J. Cell Biol. (in the press).
Peters, C. et al. Control of the terminal step of intracellular membrane fusion by protein phosphatase 1. Science 285, 1084 –1087 (1999).
Weisman, L. S., Bacallao, R. & Wickner, W. Multiple methods of visualizing the yeast vacuole permit evaluation of its morphology and inheritance during the cell cycle. J. Cell Biol. 105, 1539–1547 (1987).
Foster, L. J. et al. Binary interactions of the SNARE proteins syntaxin-4, SNAP23, and VAMP-2 and their regulation by phosphorylation. Biochemistry 37, 11089–11096 ( 1998).
Risinger, C. & Bennett, M. K. Differential phosphorylation of syntaxin and synaptosome-associated protein of 25 kDa (SNAP–25) isoforms. J. Neurochem. 72, 614– 624 (1999).
McNew, J. A. et al. Ykt6p, a prenylated SNARE essential for endoplasmic reticulum–Golgi transport. J. Biol. Chem. 272, 17776 –17783 (1997).
Schneiter, R. et al. Electrospray ionization tandem mass spectrometry (ESI-MS/MS) analysis of the lipid molecular species composition of yeast subcellular membranes reveals acyl chain-based sorting/remodeling of distinct molecular species en route to the plasma membrane. J. Cell Biol. 146, 741–754 (1999).
Zinser, E. & Daum, G. Isolation and biochemical characterization of organelles from the yeast, Saccharomyces cerevisiae. Yeast 11, 493–536 ( 1995).
Acknowledgements
We wish to thank B. Brügger for preparation of the vacuolar lipid mixture and T. Wolfe for help with the manuscript. This work was supported by an NIH grant (to J.E.R.) and postdoctoral fellowships of the Japan Society for the Promotion of Science (to R.F.), the Human Frontiers Science Program Organization (to W.N.), the Deutsche Forschungsgemeinschaft (to T.E.), the Medical Research Council of Canada (to F.P.), the National Institutes of Health (to J.A.M.), the Swiss National Science Foundation (to T.W.) and the European Molecular Biology Organization (to T.W.).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Fukuda, R., McNew, J., Weber, T. et al. Functional architecture of an intracellular membrane t-SNARE. Nature 407, 198–202 (2000). https://doi.org/10.1038/35025084
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35025084
This article is cited by
-
Physiological lipid composition is vital for homotypic ER membrane fusion mediated by the dynamin-related GTPase Sey1p
Scientific Reports (2016)
-
OsSNAP32, a SNAP25-type SNARE protein-encoding gene from rice, enhanced resistance to blast fungus
Plant Growth Regulation (2016)
-
Multiple and distinct strategies of yeast SNAREs to confer the specificity of membrane fusion
Scientific Reports (2014)
-
Dynamin−SNARE interactions control trans-SNARE formation in intracellular membrane fusion
Nature Communications (2013)
-
The impact of bacterial infection on mast cell degranulation
Immunologic Research (2011)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.