Abstract
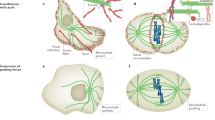
The mitotic spindle uses microtubule-based motor proteins to assemble itself and to segregate sister chromatids. It is becoming clear that motors invoke several distinct mechanisms to generate the forces that drive mitosis. Moreover, in carrying out its function, the spindle appears to pass through a series of transient steady-state structures, each established by a delicate balance of forces generated by multiple complementary and antagonistic motors. Transitions from one steady state to the next can occur when a change in the activity of a subset of mitotic motors tips the balance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hoyt, M. A. & Geiser, J. R. Genetic analysis of the mitotic spindle. Annu. Rev. Genet. 30, 7– 33 (1996).
McIntosh, J. R. & McDonald, K. L. The mitotic spindle. Sci. Am. 261, 48– 56 (1989).
Inoue, S. & Sato, H. Cell motility by labile association of molecules. The nature of mitotic spindle fibers and their role in chromosome movement. J. Gen. Physiol. 50 (Suppl.), 259–292 (1967).
Mitchison, T. & Kirschner, M. Dynamic instability of microtubule growth. Nature 312, 237– 242 (1984).
Inoue, S. & Salmon, E. D. Force generation by microtubule assembly/disassembly in mitosis and related movements. Mol. Biol. Cell 6, 1619–1640 ( 1995).
McIntosh, J. R., Hepler, P. K. & Van Wie, D. G. Model for mitosis. Nature 224, 659–663 (1969).
McDonald, K. L., Edwards, M. K. & McIntosh, J. R. Cross-sectional structure of the central mitotic spindle of Diatoma vulgare. Evidence for specific interactions between antiparallel microtubules. J. Cell Biol. 83, 443–461 (1979).
Vale, R. D. & Fletterick, R. J. The design plan of kinesin motors. Annu. Rev. Cell Dev. Biol. 13, 745 –777 (1997).
Holzbaur, E. L. & Vallee, R. B. DYNEINS: molecular structure and cellular function. Annu. Rev. Cell Biol. 10, 339–372 (1994).
Enos, A. P. & Morris, N. R. Mutation of a gene that encodes a kinesin-like protein blocks nuclear division in A. nidulans. Cell 60, 1019–1027 ( 1990).
Meluh, P. B. & Rose, M. D. KAR3, a kinesin-related gene required for yeast nuclear fusion. Cell 60, 1029– 1041 (1990); erratum ibid 61, 548.
Cole, D. G., Saxton, W. M., Sheehan, K. B. & Scholey, J. M. A “slow” homotetrameric kinesin-related motor protein purified from Drosophila embryos. J. Biol. Chem. 269, 22913–22916 (1994).
Kashina, A. S. et al. A bipolar kinesin. Nature 379, 270–272 (1996).
Gordon, D. M. & Roof, D. M. The kinesin-related protein Kip1p of Saccharomyces cerevisiae is bipolar. J. Biol. Chem. 274, 28779–28786 (1999).
Sharp, D. J. et al. The bipolar kinesin, KLP61F, cross-links microtubules within interpolar microtubule bundles of Drosophila embryonic mitotic spindles. J. Cell Biol. 144, 125–138 (1999).
Hagan, I. & Yanagida, M. Novel potential mitotic motor protein encoded by the fission yeast cut7+ gene. Nature 347 , 563-566 (1990).
Hoyt, M. A., He, L., Loo, K. K. & Saunders, W. S. Two Saccharomyces cerevisiae kinesin-related gene products required for mitotic spindle assembly. J. Cell Biol. 118, 109– 120 (1992).
Roof, D. M., Meluh, P. B. & Rose, M. D. Kinesin-related proteins required for assembly of the mitotic spindle. J. Cell Biol. 118, 95–108 (1992).
Sawin, K. E., LeGuellec, K., Philippe, M. & Mitchison, T. J. Mitotic spindle organization by a plus-end-directed microtubule motor. Nature 359, 540–543 ( 1992).
Heck, M. M. et al. The kinesin-like protein KLP61F is essential for mitosis in Drosophila. J. Cell Biol. 123, 665– 679 (1993).
Blangy, A. et al. Phosphorylation by p34cdc2 regulates spindle association of human Eg5, a kinesin-related motor essential for bipolar spindle formation in vivo. Cell 83, 1159– 1169 (1995).
Sharp, D. J., Yu, K. R., Sisson, J. C., Sullivan, W. & Scholey, J. M. Antagonistic microtubule-sliding motors position mitotic centrosomes in Drosophila early embryos. Nature Cell Biol. 1, 51–54 (1999 ).
Huxley, H. E. Sliding filaments and molecular motile systems. J. Biol. Chem. 265, 8347–8350 ( 1990).
Chandra, R., Salmon, E. D., Erickson, H. P., Lockhart, A. & Endow, S. A. Structural and functional domains of the Drosophila ncd microtubule motor protein. J. Biol. Chem. 268, 9005–9013 ( 1993).
Kuriyama, R. et al. Characterization of a minus end-directed kinesin-like motor protein from cultured mammalian cells. J. Cell Biol. 129, 1049–1059 (1995).
Pidoux, A. L., LeDizet, M. & Cande, W. Z. Fission yeast pkl1 is a kinesin-related protein involved in mitotic spindle function. Mol. Biol. Cell 7, 1639–1655 (1996).
Karabay, A. & Walker, R. A. Identification of microtubule binding sites in the Ncd tail domain. Biochemistry 38, 1838–1849 (1999).
Narasimhulu, S. B. & Reddy, A. S. Characterization of microtubule binding domains in the Arabidopsis kinesin-like calmodulin binding protein. Plant Cell 10, 957– 965 (1998).
Kuriyama, R. et al. Heterogeneity and microtubule interaction of the CHO1 antigen, a mitosis-specific kinesin-like protein. Analysis of subdomains expressed in insect sf9 cells. J. Cell Sci. 107, 3485 –3499 (1994).
Waterman-Storer, C. M., Karki, S. & Holzbaur, E. L. The p150Glued component of the dynactin complex binds to both microtubules and the actin-related protein centractin (Arp-1). Proc. Natl Acad. Sci. USA 92, 1634– 1638 (1995).
McDonald, H. B., Stewart, R. J. & Goldstein, L. S. The kinesin-like ncd protein of Drosophila is a minus end-directed microtubule motor. Cell 63, 1159–1165 (1990).
Nislow, C., Lombillo, V. A., Kuriyama, R. & McIntosh, J. R. A plus-end-directed motor enzyme that moves antiparallel microtubules in vitro localizes to the interzone of mitotic spindles. Nature 359, 543–547 (1992).
Verde, F., Berrez, J. M., Antony, C. & Karsenti, E. Taxol-induced microtubule asters in mitotic extracts of Xenopus eggs: requirement for phosphorylated factors and cytoplasmic dynein. J. Cell Biol. 112, 1177–1187 (1991).
Sharp, D. J. et al. Functional coordination of three mitotic motors in Drosophila embryos. Mol. Biol. Cell 11, 241– 253 (2000).
Adams, R. R., Tavares, A. A., Salzberg, A., Bellen, H. J. & Glover, D. M. pavarotti encodes a kinesin-like protein required to organize the central spindle and contractile ring for cytokinesis. Genes Dev. 12, 1483– 1494 (1998).
Raich, W. B., Moran, A. N., Rothman, J. H. & Hardin, J. Cytokinesis and midzone microtubule organization in Caenorhabditis elegans require the kinesin-like protein ZEN-4. Mol. Biol. Cell 9, 2037–2049 (1998).
Sharp, D. J., Rogers, G. C. & Scholey, J. M. Roles of motor proteins in building microtubule-based structures: a basic principle of cellular design. Biochim. Biophys. Acta 1496, 128–141 ( 2000).
Yeh, E., Skibbens, R. V., Cheng, J. W., Salmon, E. D. & Bloom, K. Spindle dynamics and cell cycle regulation of dynein in the budding yeast, Saccharomyces cerevisiae. J. Cell Biol. 130, 687–700 (1995).
Busson, S., Dujardin, D., Moreau, A., Dompierre, J. & De Mey, J. R. Dynein and dynactin are localized to astral microtubules and at cortical sites in mitotic epithelial cells. Curr. Biol. 8, 541–4 (1998 ).
O'Connell, C. B. & Wang, Y. Mammalian spindle orientation and position respond to changes in cell shape in a dynein-dependent fashion. Mol. Biol. Cell 11, 1765– 1764 (2000).
Holleran, E. A., Tokito, M. K., Karki, S. & Holzbaur, E. L. Centractin (ARP1) associates with spectrin revealing a potential mechanism to link dynactin to intracellular organelles. J. Cell Biol. 135, 1815–1829 (1996).
Merdes, A. & Cleveland, D. W. Pathways of spindle pole formation: different mechanisms; conserved components. J. Cell Biol. 138, 953–956 (1997).
Matthies, H. J., McDonald, H. B., Goldstein, L. S. & Theurkauf, W. E. Anastral meiotic spindle morphogenesis: role of the non-claret disjunctional kinesin-like protein. J. Cell Biol. 134, 455–464 (1996).
Heald, R. et al. Self-organization of microtubules into bipolar spindles around artificial chromosomes in Xenopus egg extracts. Nature 382, 420–425 (1996).
Rieder, C. L. & Salmon, E. D. The vertebrate cell kinetochore and its roles during mitosis. Trends Cell Biol. 8, 310–8 (1998).
Hyman, A. A. & Mitchison, T. J. Two different microtubule-based motor activities with opposite polarities in kinetochores. Nature 351, 206–211 ( 1991).
Yen, T. J., Li, G., Schaar, B. T., Szilak, I. & Cleveland, D. W. CENP-E is a putative kinetochore motor that accumulates just before mitosis. Nature 359, 536– 539 (1992).
Steuer, E. R., Wordeman, L., Schroer, T. A. & Sheetz, M. P. Localization of cytoplasmic dynein to mitotic spindles and kinetochores. Nature 345, 266–268 ( 1990).
Pfarr, C. M. et al. Cytoplasmic dynein is localized to kinetochores during mitosis. Nature 345, 263–265 (1990).
Wood, K. W., Sakowicz, R., Goldstein, L. S. & Cleveland, D. W. CENP-E is a plus end-directed kinetochore motor required for metaphase chromosome alignment. Cell 91, 357– 366 (1997).
Starr, D. A., Williams, B. C., Hays, T. S. & Goldberg, M. L. ZW10 helps recruit dynactin and dynein to the kinetochore. J. Cell Biol. 142, 763–774 ( 1998).
Bowman, A. B. et al. Drosophila roadblock and Chlamydomonas LC7: a conserved family of dynein-associated proteins involved in axonal transport, flagellar motility, and mitosis. J. Cell Biol. 146, 165–180 (1999).
Lee, S., Wisniewski, J. C., Dentler, W. L. & Asai, D. J. Gene knockouts reveal separate functions for two cytoplasmic dyneins in Tetrahymena thermophila. Mol. Biol. Cell 10, 771–784 (1999).
Rieder, C. L. & Salmon, E. D. Motile kinetochores and polar ejection forces dictate chromosome position on the vertebrate mitotic spindle. J. Cell Biol. 124, 223– 233 (1994).
Wang, S. Z. & Adler, R. Chromokinesin: a DNA-binding, kinesin-like nuclear protein. J. Cell Biol. 128, 761– 768 (1995).
Molina, I. et al. A chromatin-associated kinesin-related protein required for normal mitotic chromosome segregation in Drosophila. J. Cell Biol. 139, 1361–1371 (1997).
Ruden, D. M., Cui, W., Sollars, V. & Alterman, M. A Drosophila kinesin-like protein, Klp38B, functions during meiosis, mitosis, and segmentation. Dev. Biol. 191, 284– 296 (1997).
Alphey, L. et al. KLP38B: a mitotic kinesin-related protein that binds PP1. J. Cell Biol. 138, 395– 409 (1997).
Tokai, N. et al. Kid, a novel kinesin-like DNA binding protein, is localized to chromosomes and the mitotic spindle. Embo J 15, 457-467 (1996).
Lombillo, V. A., Nislow, C., Yen, T. J., Gelfand, V. I. & McIntosh, J. R. Antibodies to the kinesin motor domain and CENP-E inhibit microtubule depolymerization-dependent motion of chromosomes in vitro. J. Cell Biol. 128, 107– 115 (1995).
Walczak, C. E., Mitchison, T. J. & Desai, A. XKCM1: a Xenopus kinesin-related protein that regulates microtubule dynamics during mitotic spindle assembly. Cell 84, 37–47 ( 1996).
Desai, A., Verma, S., Mitchison, T. J. & Walczak, C. E. Kin I kinesins are microtubule-destabilizing enzymes. Cell 96, 69–78 (1999).
Wordeman, L. & Mitchison, T. J. Identification and partial characterization of mitotic centromere- associated kinesin, a kinesin-related protein that associates with centromeres during mitosis. J. Cell Biol. 128, 95–104 ( 1995).
Maney, T., Hunter, A. W., Wagenbach, M. & Wordeman, L. Mitotic centromere-associated kinesin is important for anaphase chromosome segregation. J. Cell Biol. 142, 787– 801 (1998).
Straight, A. F., Sedat, J. W. & Murray, A. W. Time-lapse microscopy reveals unique roles for kinesins during anaphase in budding yeast. J. Cell Biol. 143 , 687–694 (1998).
Saunders, W. S. & Hoyt, M. A. Kinesin-related proteins required for structural integrity of the mitotic spindle. Cell 70, 451–458 ( 1992).
O'Connell, M. J., Meluh, P. B., Rose, M. D. & Morris, N. R. Suppression of the bimC4 mitotic spindle defect by deletion of klpA, a gene encoding a KAR3-related kinesin-like protein in Aspergillus nidulans. J. Cell Biol. 120, 153– 162 (1993).
Mountain, V. et al. The kinesin-related protein, HSET, opposes the activity of Eg5 and cross-links microtubules in the mammalian mitotic spindle. J Cell Biol 147, 351-366 (1999 ).
Follette, P. J. & O'Farrell, P. H. Cdks and the Drosophila cell cycle. Curr. Opin. Genet. Dev. 7, 17–22 (1997).
Sawin, K. E. & Mitchison, T. J. Mutations in the kinesin-like protein Eg5 disrupting localization to the mitotic spindle. Proc. Natl Acad. Sci. USA 92, 4289–4293 (1995).
Lee, K. S., Yuan, Y. L., Kuriyama, R. & Erikson, R. L. Plk is an M-phase-specific protein kinase and interacts with a kinesin- like protein, CHO1/MKLP-1. Mol. Cell Biol. 15, 7143–7151 (1995).
Chan, G. K., Jablonski, S. A., Sudakin, V., Hittle, J. C. & Yen, T. J. Human BUBR1 is a mitotic checkpoint kinase that monitors CENP-E functions at kinetochores and binds the cyclosome/APC. J. Cell Biol. 146, 941– 954 (1999).
Giet, R., Uzbekov, R., Cubizolles, F., Le Guellec, K. & Prigent, C. The Xenopus laevis aurora-related protein kinase pEg2 associates with and phosphorylates the kinesin-related protein XlEg5. J. Biol. Chem. 274, 15005 –15013 (1999).
Mayer, T. U. et al. Small molecule inhibitor of mitotic spindle bipolarity identified in a phenotype-based screen. Science 286, 971–974 (1999).
Nicklas, R. B. The forces that move chromosomes in mitosis. Annu. Rev. Biophys. Biophys. Chem. 17, 431–449 (1988).
Heald, R. Motor function in the mitotic spindle. Cell 102, 399–402 (2000).
Funabiki, H. & Murray, A. W. The Xenopus Chromokinesin, XKid, is essential for metaphase chromosome alignment and must be degraded to allow anaphase chromosome movement. Cell 102, 411–424 (2000).
Antonio, C. et al. XKid, a chromokinesin required for chromosome alignment on the metaphase plate. Cell 102, 425– 435 (2000).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Rights and permissions
About this article
Cite this article
Sharp, D., Rogers, G. & Scholey, J. Microtubule motors in mitosis. Nature 407, 41–47 (2000). https://doi.org/10.1038/35024000
Issue Date:
DOI: https://doi.org/10.1038/35024000
This article is cited by
-
Bioinformatical analysis of the key differentially expressed genes for screening potential biomarkers in Wilms tumor
Scientific Reports (2023)
-
Anti-aging Effects of Alu Antisense RNA on Human Fibroblast Senescence Through the MEK-ERK Pathway Mediated by KIF15
Current Medical Science (2023)
-
Vesicle-mediated transport of ALIX and ESCRT-III to the intercellular bridge during cytokinesis
Cellular and Molecular Life Sciences (2023)
-
A gelation transition enables the self-organization of bipolar metaphase spindles
Nature Physics (2022)
-
Synthesis,characterization and biological activities of nitrogen-containing Combretastatin A-4 derivatives
Medicinal Chemistry Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.