Abstract
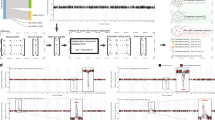
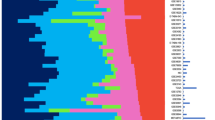
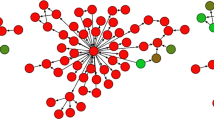
Human breast tumours are diverse in their natural history and in their responsiveness to treatments1. Variation in transcriptional programs accounts for much of the biological diversity of human cells and tumours. In each cell, signal transduction and regulatory systems transduce information from the cell's identity to its environmental status, thereby controlling the level of expression of every gene in the genome. Here we have characterized variation in gene expression patterns in a set of 65 surgical specimens of human breast tumours from 42 different individuals, using complementary DNA microarrays representing 8,102 human genes. These patterns provided a distinctive molecular portrait of each tumour. Twenty of the tumours were sampled twice, before and after a 16-week course of doxorubicin chemotherapy, and two tumours were paired with a lymph node metastasis from the same patient. Gene expression patterns in two tumour samples from the same individual were almost always more similar to each other than either was to any other sample. Sets of co-expressed genes were identified for which variation in messenger RNA levels could be related to specific features of physiological variation. The tumours could be classified into subtypes distinguished by pervasive differences in their gene expression patterns.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Tavassoli, F. A. & Schnitt, S. J. Pathology of the Breast (Elsevier, New York, 1992).
Eisen, M. B. & Brown, P. O. DNA arrays for analysis of gene expression. Methods Enzymol. 303, 179– 205 (1999).
Ross, D. T. et al. Systematic variation in gene expression patterns in human cancer cell lines. Nature Genet. 24, 227 –235 (2000).
Aas, T. et al. Specific P53 mutations are associated with de novo resistance to doxorubicin in breast cancer patients. Nature Med. 2, 811–814 (1996).
Eisen, M. B., Spellman, P. T., Brown, P. O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863– 14868 (1998).
Perou, C. M. et al. Distinctive gene expression patterns in human mammary epithelial cells and breast cancers. Proc. Natl Acad. Sci. USA 96, 9212–9217 (1999).
Yang, G. P., Ross, D. T., Kuang, W. W., Brown, P. O. & Weigel, R. J. Combining SSH and cDNA microarrays for rapid identification of differentially expressed genes. Nucleic Acids Res. 27, 1517–1523 (1999).
Hoch, R. V., Thompson, D. A., Baker, R. J. & Weigel, R. J. GATA-3 is expressed in association with estrogen receptor in breast cancer. Int. J. Cancer 84, 122– 128 (1999).
Pauletti, G., Godolphin, W., Press, M. F. & Slamon, D. J. Detection and quantitation of HER-2/neu gene amplification in human breast cancer archival material using fluorescence in situ hybridization. Oncogene 13, 63–72 ( 1996).
Pollack, J. R. et al. Genome-wide analysis of DNA copy-number changes using cDNA microarrays. Nature Genet. 23, 41– 46 (1999).
Ronnov-Jessen, L., Petersen, O. W. & Bissell, M. J. Cellular changes involved in conversion of normal to malignant breast: importance of the stromal reaction. Physiol. Rev. 76, 69–125 ( 1996).
Taylor-Papadimitriou, J. et al. Keratin expression in human mammary epithelial cells cultured from normal and malignant tissue: relation to in vivo phenotypes and influence of medium. J. Cell Sci. 94, 403– 413 (1989).
Golub, T. R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286 , 531–537 (1999).
Dairkee, S. H., Mayall, B. H., Smith, H. S. & Hackett, A. J. Monoclonal marker that predicts early recurrence of breast cancer. Lancet 1, 514 (1987).
Dairkee, S. H., Puett, L. & Hackett, A. J. Expression of basal and luminal epithelium-specific keratins in normal, benign, and malignant breast tissue. J. Natl Cancer Inst. 80, 691–695 (1988).
Malzahn, K., Mitze, M., Thoenes, M. & Moll, R. Biological and prognostic significance of stratified epithelial cytokeratins in infiltrating ductal breast carcinomas. Virchows Arch. 433, 119 –129 (1998).
Guelstein, V. I. et al. Monoclonal antibody mapping of keratins 8 and 17 and of vimentin in normal human mammary gland, benign tumors, dysplasias and breast cancer. Int. J. Cancer 42, 147– 153 (1988).
Gusterson, B. A. et al. Distribution of myoepithelial cells and basement membrane proteins in the normal breast and in benign and malignant breast diseases. Cancer Res. 42, 4763–4770 (1982).
Nagle, R. B. et al. Characterization of breast carcinomas by two monoclonal antibodies distinguishing myoepithelial from luminal epithelial cells. J. Histochem. Cytochem. 34, 869–881 (1986).
Berns, E. M. et al. Prevalence of amplification of the oncogenes c-myc, HER2/neu, and int-2 in one thousand human breast tumors: correlation with steroid receptors. Eur. J. Cancer 28, 697– 700 (1992).
Heintz, N. H., Leslie, K. O., Rogers, L. A. & Howard, P. L. Amplification of the c-erb B-2 oncogene and prognosis of breast adenocarcinoma. Arch. Pathol. Lab. Med. 114, 160– 163 (1990).
DeRisi, J. L., Iyer, V. R. & Brown, P. O. Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686 (1997).
Alizadeh, A. A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
Acknowledgements
We thank W. Gerald and L. Norton for the three New York tumour specimens; M. Stampfer and P. Yaswen for the 184 sample mRNAs; and members of the P. O. Brown, D. Botstein and A.-L. Børresen-Dale labs for discussions. We are grateful to the NCI and the Howard Hughes Medical Institute who provided support for this research. C.M.P. is a SmithKline Beecham Pharmaceuticals Fellow of the Life Sciences Research Foundation. T.S. is a research fellow of the Norwegian Cancer Society. M.B.E. is an Alfred P. Sloan Foundation Postdoctoral Fellow in Computational Molecular Biology. D.T.R. is a Walter and Idun Berry Fellow. P.O.B. is an Associate Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Perou, C., Sørlie, T., Eisen, M. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000). https://doi.org/10.1038/35021093
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/35021093
This article is cited by
-
Clustering of HR + /HER2− breast cancer in an Asian cohort is driven by immune phenotypes
Breast Cancer Research (2024)
-
Ensemble methods of rank-based trees for single sample classification with gene expression profiles
Journal of Translational Medicine (2024)
-
Identification of a Notch transcriptomic signature for breast cancer
Breast Cancer Research (2024)
-
Targeting HER2-positive breast cancer cells by a combination of dasatinib and BMS-202: Insight into the molecular pathways
Cancer Cell International (2024)
-
The androgen receptor interacts with GATA3 to transcriptionally regulate a luminal epithelial cell phenotype in breast cancer
Genome Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.