Abstract
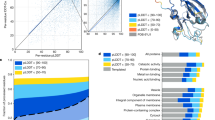
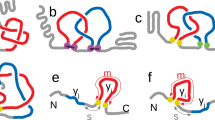
The polypeptide chains that make up proteins have thousands of atoms and hence millions of possible inter-atomic interactions. It might be supposed that the resulting complexity would make prediction of protein structure and protein-folding mechanisms nearly impossible. But the fundamental physics underlying folding may be much simpler than this complexity would lead us to expect: folding rates and mechanisms appear to be largely determined by the topology of the native (folded) state, and new methods have shown great promise in predicting protein-folding mechanisms and the three-dimensional structures of proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Anfinson,C. Principles that govern the folding of protein chains. Science 181, 223–227 (1973).
Baldwin,R. L. & Rose,G. D. Is protein folding hierarchic? II. Folding intermediates and transition states. Trends Biochem. Sci. 24, 26–33 ( 1999).
Jackson,S. E. How do small single-domain proteins fold? Fold. Des. 3, R81–91 (1998).
Fersht,A. Structure and Mechanism in Protein Science: A Guide to Enzyme Catalysis and Protein Folding (Freeman, New York, 1999).
Chan,H. S. & Dill,K. A. Protein folding in the landscape perspective: chevron plots and non- Arrhenius kinetics. Proteins 30, 2–33 (1998 ).
Bryngelson,J. D., Onuchic,J. N., Socci,N. D. & Wolynes,P. G. Funnels, pathways, and the energy landscape of protein folding: a synthesis. Proteins 21, 167–195 (1995).
Dobson,C. M. & Karplus,M. The fundamentals of protein folding: bringing together theory and experiment. Curr. Opin. Struct. Biol. 9, 92–101 ( 1999).
Horwich,A. L. Chaperone rings in protein folding and degradation. Proc. Natl Acad. Sci. USA 96, 11033–11040 (1999).
Shakhnovich,E. I. Folding nucleus: specific or multiple? Insights from lattice models and experiments. Fold. Des. 3, R108–111 (1998).
Pande,V. S., Grosberg,A. Y., Tanaka,T. & Rokhsar,D. Pathways for protein folding: is a new view needed? Curr. Opin. Struct. Biol. 8, 68–79 ( 1998).
Thirumalai,D. & Klimov,D. K. Fishing for folding nuclei in lattice models and proteins. Fold. Des. 3, R112– 118 (1998).
Alm,E. & Baker,D. Matching theory and experiment in protein folding. Curr. Opin. Struct. Biol. 9, 189 –196 (1999).
Riddle,D. S. et al. Functional rapidly folding proteins from simplified amino acid sequences. Nature Struct. Biol. 4, 805–809 (1997).
Kim,D. E., Gu,H. & Baker,D. The sequences of small proteins are not extensively optimized for rapid folding by natural selection. Proc. Natl Acad. Sci. USA 95, 4982–4986 (1998).
Perl,D. et al. Conservation of rapid two-state folding in mesophilic, thermophilic and hyperthermophilic cold shock proteins. Nature Struct. Biol. 5, 229–235 ( 1998).
Chiti,F. et al. Mutational analysis of acylphosphatase suggests the importance of topology and contact order in protein folding. Nature Struct. Biol. 6, 1005–1009 ( 1999).
Martinez,J. C. & Serrano,L. The folding transition state between SH3 domains is conformationally restricted and evolutionarily conserved. Nature Struct. Biol. 6, 1010– 1016 (1999).
Riddle,D. S. et al. Experiment and theory highlight role of native state topology in SH3 folding. Nature Struct. Biol. 6, 1016–1024 (1999).
Plaxco,K. W., Simons,K. T. & Baker, D. Contact order, transition state placement and the refolding rates of single domain proteins. J. Mol. Biol. 277, 985–994 (1998).
Shea J. E., Onuchic J. N. & Brooks,C. L. III. Exploring the origins of topological frustration: design of a minimally frustrated model of fragment B of protein A. Proc. Natl Acad. Sci. USA 96, 12512–12517 ( 1999).
Onuch,J. N., Nymeyer,H., Garcia,A. E., Chaine,J. & Socci,N. D. The energy landscape theory of protein folding: insights into folding mechanism and scenarios. Adv. Protein Chem. 53, 87–152 (2000).
Micheletti,C., Banavar,J. R., Maritan,A. & Seno,F. Protein structures and optimal folding from a geometrical variational principle. Phys. Rev. Lett. 82, 3372–3375 (1999).
Abkevich,V., Gutin,A. & Shakhnovich, E. Specific nucleus as the transition state for protein folding: evidence from the lattice model. Biochemistry 33, 10026–10036 (1994).
Munoz,V. & Eaton,W. A. A simple model for calculating the kinetics of protein folding from three-dimensional structures. Proc. Natl Acad. Sci. USA 96, 11311– 11316 (1999).
Galzitskaya,O. V. & Finkelstein,A. V. A theoretical search for folding/unfolding nuclei in three-dimensional protein structures. Proc. Natl Acad. Sci. USA 96, 11299– 11304 (1999).
Alm,E. & Baker,D. Prediction of protein-folding mechanisms from free-energy landscapes derived from native structures. Proc. Natl Acad. Sci. USA 96, 11305–11310 (1999).
Go,N. Theoretical studies of protein folding. Annu. Rev. Biophys. Bioeng. 12, 183–210 (1983).
Portman J. J., Takada,S. & Wolynes,P. G. Variational theory for site resolved protein folding free energy surfaces. Phys. Rev. Lett. 81, 5237–5240 (1998).
Debye,D & Goddard,W. A. First principles prediction of protein-folding rates. J. Mol. Biol. 294, 619–625 (1999).
Li,A. J. & Daggett,V. Identification and characterization of the unfolding transition state of chymotrypsin inhibitor 2 by molecular dynamics simulations. J. Mol. Biol. 257, 412–429 (1996).
Lazaridis,T. & Karplus,M. ‘New view’ of protein folding reconciled with the old through multiple unfolding simulations. Science 278, 1928–1931 ( 1997).
Sheinerman,F. B. & Brooks,C. L. III A molecular dynamics simulation study of segment B1 of protein G. Proteins 29, 193–202 ( 1997).
Burton,R. E., Myers,J. K. & Oas,T. G. Protein folding dynamics: quantitative comparison between theory and experiment. Biochemistry 37, 5337–5343 (1998).
Moult,J., Hubbard,T., Fidelis,K. & Pederson,J. T. Critical assessment of methods of protein structure prediction (CASP): round III. Proteins (Suppl.) 3, 2–6 (1999).
Orengo,C. A., Bray,J. E., Hubbard,T. LoConte,L. & Sillitoe, I. Analysis and assessment of ab initio three-dimensional prediction, secondary structure, and contacts prediction. Proteins (Suppl.) 3, 149–170 ( 1999).
Murzin,A. G. Structure classification-based assessment of CASP3 predictions for the fold recognition targets. Proteins (Suppl.) 3, 88–103 (1999).
Villegas,V., Martinez,J. C., Aviles,F. X. & Serrano,L. Structure of the transition state in the folding process of human procarboxypeptidase A2 activation domain. J. Mol. Biol. 283, 1027–1036 (1998).
Lopez-Hernandez,E. & Serrano,L. Structure of the transition state for folding of the 129 aa protein CheY resembles that of a smaller protein, CI-2. Fold. Des. 1, 43 –55 (1996).
Simons,K. T., Bonneau,R., Ruczinski,I. & Baker,D. Ab initio protein structure prediction of CASP III targets using ROSETTA. Proteins (Suppl.) 3, 171–176 ( 1999).
Samudrala,R. Xia,Y. Huang,E. & Levitt,M. Ab initio protein structure prediction using a combined hierarchical approach. Proteins (Suppl.) 3, 194–198 ( 1999).
Ortiz,A. R., Kolinski,A., Rotkiewicz,P., Ilkowski,B. & Skolnick,J. Ab initio folding of proteins using restraints derived from evolutionary information. Proteins (Suppl.) 3, 177–185 ( 1999).
Weigelt,J., Brown,S. E., Miles,C. S. & Dixon,N. E. NMR structure of the N-terminal domain of E. coli DnaB helicase: implications for structure rearrangements in the helicase hexamer. Structure 7, 681–690 (1999).
Slupsky,C. M. et al. Structure of the Ets-1 pointed domain and mitogen-activated protein kinase phosphorylation site. Proc. Natl Acad. Sci. USA 95, 12129–12134 ( 1998).
Rhee,S., Martin,R. G., Rosner,J. L. & Davies,D. R. A novel DNA-binding motif in MarA: the first structure for an AraC family transcriptional activator. Proc. Natl Acad. Sci. USA 95, 10413–10418 (1998).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Baker, D. A surprising simplicity to protein folding. Nature 405, 39–42 (2000). https://doi.org/10.1038/35011000
Issue Date:
DOI: https://doi.org/10.1038/35011000
This article is cited by
-
A tile model of circuit topology for self-entangled biopolymers
Scientific Reports (2023)
-
A topology framework for macromolecular complexes and condensates
Nano Research (2022)
-
StructureDistiller: Structural relevance scoring identifies the most informative entries of a contact map
Scientific Reports (2019)
-
Sequence and structural patterns detected in entangled proteins reveal the importance of co-translational folding
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.