Abstract
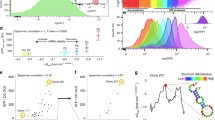
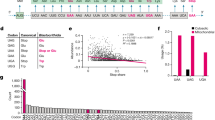
The two translational release factors of prokaryotes, RF1 and RF2, catalyse the termination of polypeptide synthesis at UAG/UAA and UGA/UAA stop codons, respectively1,2,3. However, how these polypeptide release factors read both non-identical and identical stop codons is puzzling4. Here we describe the basis of this recognition. Swaps of each of the conserved domains between RF1 and RF2 in an RF1–RF2 hybrid led to the identification of a domain that could switch recognition specificity. A genetic selection among clones encoding random variants of this domain showed that the tripeptides Pro-Ala-Thr and Ser-Pro-Phe determine release-factor specificity in vivo in RF1 and RF2, respectively. An in vitro release study of tripeptide variants indicated that the first and third amino acids independently discriminate the second and third purine bases, respectively. Analysis with stop codons containing base analogues indicated that the C2 amino group of purine may be the primary target of discrimination of G from A. These findings show that the discriminator tripeptide of bacterial release factors is functionally equivalent to that of the anticodon of transfer RNA, irrespective of the difference between protein and RNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Scolnick, E., Tompkins, R., Caskey, T. & Nirenberg,, M. Release factors differing in specificity for terminator codons. Proc. Natl Acad. Sci. USA 61, 768–774 ( 1968).
Nakamura, Y., Ito, K. & Isaksson, L. A. Emerging understanding of translation termination. Cell 87, 147–150 ( 1996).
Buckingham, R. H., Grentzmann, G. & Kisselev, L. Polypeptide chain release factors. Mol. Microbiol. 24, 449–456 ( 1997).
Nakamura, Y. & Ito, K. How protein reads the stop codon and terminates translation. Genes Cells 3, 263 –278 (1998).
Moffat, J. G. & Tate, W. P. A single proteolytic cleavage in release factor 2 stabilizes ribosome binding and abolishes peptidyl–tRNA hydrolysis activity. J. Biol. Chem. 269, 18899–18903 (1994).
Ito, K., Ebihara, K., Uno, M. & Nakamura, Y. Conserved motifs of prokaryotic and eukaryotic polypeptide release factors: tRNA–protein mimicry hypothesis. Proc. Natl Acad. Sci. USA 93, 5443–5448 (1996).
Ito, K., Uno, M. & Nakamura, Y. Single amino acid substitution in prokaryote polypeptide release factor 2 permits it to terminate translation at all three stop codons. Proc. Natl. Acad. Sci. USA 95, 8165– 8169 (1998).
Auld, D. S. & Schimmel, P. Switching recognition of two tRNA synthetases with an amino acid swap in a designed peptide. Science 267, 1994–1996 ( 1995).
Wakasugi, K., Quinn, C. L., Tao, N. & Schimmel, P. Genetic code in evolution: switching species–specific aminoacylation with a peptide transplant. EMBO J. 17, 297– 305 (1998).
Crick, F. H. C. Codon–anticodon pairing: the wobble hypothesis. J. Mol. Biol. 19, 548–555 ( 1966).
Soll, D. & RajBhandary,, U. L. Studies on polynucleotides. LXXVI. Specificity of tRNA for codon recognition as studied by amino acid incorporation. J. Mol. Biol. 29, 113– 124 (1967).
Murgola, E. J., Hijiazi, K. A., Goringer, H. U. & Dahlberg, A. E. Mutant 16S rRNA: a codon specific translational suppressor. Proc. Natl Acad. Sci. USA 85, 4162–4165 (1988).
Uno, M., Ito, K. & Nakamura, Y. Functional specificity of amino acid at position 246 in the tRNA mimicry domain of bacterial release factor 2. Biochimie 78, 935–943 ( 1996).
Yoshimura, K., Ito, K. & Nakamura, Y. Amber (UAG) suppressors affected in UGA/UAA–specific polypeptide release factor 2 of bacteria: genetic prediction of initial binding to ribosome preceding stop codon recognition. Genes Cells 4, 253–266 (1999).
Rydén, S. M. & Isaksson, L. A. A temperature–sensitive mutant of Escherichia coli that shows enhanced misreading of UAG/A and increased efficiency for some tRNA nonsense suppressors. Mol. Gen. Genet. 193, 38–45 ( 1984).
Kawakami, K., Inada, T. & Nakamura,, Y. Conditionally lethal and recessive UGA–suppressor mutations in the prfB gene encoding peptide chain release factor 2 of Escherichia coli. J. Bacteriol. 170, 5378–5381 (1988).
Acknowledgements
We thank C. Yanofsky and J. Atkins for critical reading of the manuscript and valuable comments. This work was supported in part by grants from The Ministry of Education, Science, Sports and Culture, Japan; the Human Frontier Science Program (awarded in 1993 and 1997); and the Basic Research for Innovation Biosciences Program of Bio-oriented Technology Research Advancement Institution (BRAIN).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ito, K., Uno, M. & Nakamura, Y. A tripeptide ‘anticodon’ deciphers stop codons in messenger RNA. Nature 403, 680–684 (2000). https://doi.org/10.1038/35001115
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35001115
This article is cited by
-
Human mtRF1 terminates COX1 translation and its ablation induces mitochondrial ribosome-associated quality control
Nature Communications (2022)
-
Release factor-dependent ribosome rescue by BrfA in the Gram-positive bacterium Bacillus subtilis
Nature Communications (2019)
-
Cultured bloodstream Trypanosoma brucei adapt to life without mitochondrial translation release factor 1
Scientific Reports (2018)
-
Structural basis for ArfA–RF2-mediated translation termination on mRNAs lacking stop codons
Nature (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.