Abstract
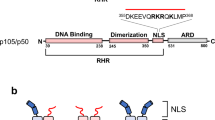
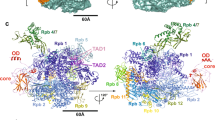
The NF-κB p50/p65 heterodimer is the classical member of the Relfamily of transcription factors which regulate diverse cellular functions such as immune response, cell growth, and development1,2,3. Other mammalian Rel family members, including theproteins p52, proto-oncoprotein c-Rel, and RelB, all have amino-terminal Rel-homology regions (RHRs)4,5,6,7. The RHR is responsible for the dimerization, DNA binding and cytosolic localization of these proteins by virtue of complex formation with inhibitor κB proteins8. Signal-induced removal of κB inhibitors allows translocation of dimers to the cell nucleus and transcriptional regulation of κB DNA-containing genes9. NF-κB specifically recognizes κB DNA elements1,10,11 with a consensus sequence of 5′-GGGRNYYYCC-3′ (R is an unspecified purine; Y is an unspecified pyrimidine; and N is any nucleotide). Here we report the crystal structure at 2.9 Å resolution of the p50/p65 heterodimer bound to the κB DNA of the intronic enhancer of the immunoglobulin light-chain gene. Our structure reveals a 5-base-pair 5′ subsite for p50, and a 4-base-pair 3′ subsite for p65. This structure indicates why the p50/p65 heterodimer interface is stronger than that of either homodimer. A comparison of this structure with those of other Rel dimers reveals that both subunits adopt variable conformations in a DNA-sequence-dependent manner. Our results explain the different behaviour of the p50/p65 heterodimer with heterologous promoters.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Sen, R. & Baltimore, D. Inducibility of κ immunoglobulin enhancer-binding protein NF-κB by a post-translational mechanism. Cell 47, 921–928 (1986).
Baldwin, A. S. J The NF-κB and I κB proteins: new discoveries and insights. Annu. Rev. Immunol. 14, 649–683 (1996).
Siebenlist, U., Franzoso, G. & Brown, K. Structure, regulation and function of NF-κB. Annu. Rev. Cell. Biol. 10, 405–455 (1994).
Ghosh, S.et al. Cloning of the p50 DNA binding subunit of NF-κB: homology to rel and dorsal. Cell 62, 1019–1029 (1990).
Kieran, M.et al. The DNA binding subunit of NF-κB is identical to factor KBF1 and homologous to the rel oncogene product. Cell 62, 1007–1018 (1990).
Nolan, G. P., Ghosh, S., Liou, H. C., Tempst, P. & Baltimore, D. DNA binding and I κB inhibition of the cloned p65 subunit of NF-κB, a rel-related polypeptide. Cell 64, 961–969 (1991).
Ruben, S. M.et al. Isolation of a rel-related human cDNA that potentially encodes the 65-kD subunit of NF-κB. Science 251, 1490–1493 (1991).
Baeuerle, P. A. & Baltimore, D. I. κB: a specific inhibitor of the NF-κB transcription factor. Science 242, 540–546 (1988).
Chen, Z.et al. Signal-induced site-specific phosphorylation targets I κB alpha to the ubiquitin-proteasome pathway. Genes Dev. 9, 1586–1597 (1995).
Zabel, U., Schreck, R. & Baeuerle, P. A. DNA binding of purified transcription factor NF-κB. Affinity, specificity, Zn2+ dependence, and differential half-site recognition. J. Biol. Chem. 266, 252–260 (1991).
Lenardo, M. J. & Baltimore, D. NF-κB: a pleiotropic mediator of inducible and tissue-specific gene control. Cell 58, 227–229 (1989).
Ghosh, G., van Duyne, G., Ghosh, S. & Sigler, P. B. Structure of NF-κB p50 homodimer bound to a κB site. Nature 373, 303–310 (1995).
Müller, C. W., Rey, F. A., Sodeoka, M., Verdine, G. L. & Harrison, S. C. Structure of the NF-κB p50 homodimer bound to DNA. Nature 373, 311–317 (1995).
Urban, M. B., Schreck, R. & Baeuerle, P. A. NF-κB contacts DNA by a heterodimer of the p50 and p65 subunit. EMBO J. 10, 1817–1825 (1991).
Lavery, R. & Sklenar, H. The definition of generalized helocoidal parameters and of axis curvature for irregular nucleic acids. J. Biomol. Struct. Dyn. 6, 63–91 (1988).
Falvo, J. V., Thanos, D. & Maniatis, T. Reversal of intrinsic DNA bends in the IFN-β gene enhancer by transcription factors and the architectural protein HMB I(Y). Cell 83, 1101–1111 (1995).
Toledano, M. B., Ghosh, D., Trinh, F. & Leonard, W. J. N-terminal DNA-binding domains contribute to differential DNA-binding specificities of NF-κB p50 and p65. Mol. Cell. Biol. 13, 852–860 (1993).
Thanos, D. & Maniatis, T. The high mobility group protein HMB I(Y) is required for NF-κB-dependent virus induction of the human IFN-β gene. Cell 71, 777–789 (1992).
Urban, M. B. & Baeuerle, P. A. The 65-kD subunit of NF-κB is a receptor for I κB and a modulator of DNA-binding specificity. Genes Dev. 4, 1975–1984 (1990).
Klemm, J. D., Rould, M. A., Aurora, R., Herr, W. & Pabo, C. O. Crystal structure of the Oct-1 POU domain bound to an octamer site: DNA recognition with tethered DNA-binding modules. Cell 77, 21–32 (1994).
Whitley, M. Z., Thanos, D., Read, M. A., Maniatis, T. & Collins, T. Astriking similarity in the organization of the E-selectin and β-interferon gene promoters. Mol. Cell. Biol. 14, 6464–6475 (1994).
Hansen, S. K., Guerrini, L. & Blasi, F. Differential DNA sequence specificity and regulation of HIV-1 enhancer activity by cRel-RelA transcription factor. J. Biol. Chem. 269, 22230–22237 (1944).
Cross, S. L., Halden, N. F., Lenardo, M. J. & Leonard, W. J. Functionally distinct NF-κB binding sites in the immunoglobulin κ and IL-2 receptor α chain genes. Science 244, 466–469 (1989).
Otwinowski, Z. in Proceedings of the CCP4 Study Weekend: Data Collection and Processing (eds Sawyer, L., Isaacs, N. & Bayley, S.) 56–62 (SERC Daresbury Laboratory, Warrington, UK, 1993).
Brünger, A. T. X-PLOR Version 3.1: A System for X-ray Crystallography and NMR 1–382 (Yale University Press, New Haven, 1992).
Cambillau, C. & Horjales, E. TOM: a FRODO subpackage for protein ligand fitting with interactive energy minimization. J. Mol. Graph. 5, 174–177 (1987).
Jones, T. A., Zou, Y., Cowan, S. W. & Kjeldgaard. Improved methods for binding protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Engh, R. A. & Huber, R. Accurate bond angle parameters for X-ray protein structure refinement. Acta Crystallogr. A 47, 392–400 (1991).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structure coordinates. J. Appl. Crystallogr. A 42, 140–149 (1993).
Evans, S. V. SETOR: hardware-lighted three-dimensional solid model representations of macromolecules. J. Mol. Graph. 11, 134–138 (1993).
Chen, Y. Q., Ghosh, S. & Ghosh, G. Anovel DNA recognition mode by NF-κB p65 homodimer. Nature Struct. Biol. (in the press).
Acknowledgements
We thank S. Kempiak for help with protein purification and T. Huxford and S.Malek for critically reading the manuscript. This work was supported by fellowships from the NSF and the Lucille P. Markey Charitable Trust to F.C. Y.-Q.C. is a fellow of the Irvington Institute of Medical Research. G.G. is a recipient of a young investigator award from the university wide AIDS research program. This work is supported by a NIH research grant to G.G.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chen, F., Huang, DB., Chen, YQ. et al. Crystal structure of p50/p65 heterodimer of transcription factor NF-κB bound to DNA. Nature 391, 410–413 (1998). https://doi.org/10.1038/34956
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/34956
This article is cited by
-
Semaphorin 7A interacts with nuclear factor NF-kappa-B p105 via integrin β1 and mediates inflammation
Cell Communication and Signaling (2023)
-
Explaining the interaction of mangiferin with MMP-9 and NF-ƙβ: a computational study
Journal of Molecular Modeling (2022)
-
IκBα targeting promotes oxidative stress-dependent cell death
Journal of Experimental & Clinical Cancer Research (2021)
-
An Integrated Approach of the Potential Underlying Molecular Mechanistic Paradigms of SARS-CoV-2-Mediated Coagulopathy
Indian Journal of Clinical Biochemistry (2021)
-
Hepatoprotective effects of phytochemicals berberine and umbelliferone against methotrexate-induced hepatic intoxication: experimental studies and in silico evidence
Environmental Science and Pollution Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.