Abstract
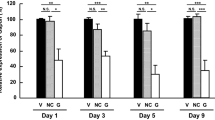
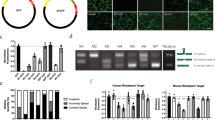
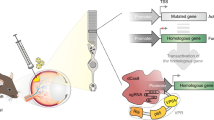
More than one hundred different mutations in the gene encoding rhodopsin are associated with a group of retinal degenerations including retinitis pigmentosa, congenital stationary night blindness and retinitis punctata albescens. Given this large heterogeneity of mutations, it would be ideal to develop mutation-independent therapies for these diseases. We describe use of RNA interference (RNAi) and specifically short hairpin RNAs (shRNAs) expressed from DNA templates to silence both normal and mutant (P23H) human rhodopsin alleles by 94.34±2.17 and 94.9±1.9%, respectively, in human embryonic retinoblasts. Degeneracy of the genetic code was used to engineer a codon-exchanged mRNA (cmRNA) that demonstrated complete resistance to silencing by the shRNA. Simulation of autosomal dominant retinitis pigmentosa in cell culture through triple transfection of DNAs expressing a cmRNA, a P23H mRNA and an shRNA revealed shRNA-mediated silencing, specifically of P23H rhodopsin by 90.64±5.19% and no loss of rhodopsin translation from the cmRNA in those cells. In addition, we present data on two alternative shRNA sequences targeting human rhodopsin. Our results have implications for the treatment of a very large variety of retinal degenerations in a mutation-independent manner.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Baylor D . How photons start vision. Proc Natl Acad Sci USA 1996; 93: 560–565.
Dryja TP, Berson EL, Rao VR, Oprian DD . Heterozygous missense mutation in the rhodopsin gene as a cause of congenital stationary night blindness. Nat Genet 1993; 4: 280–283.
Souied E et al. Retinitis punctata albescens associated with the Arg135Trp mutation in the rhodopsin gene. Am J Ophthalmol 1996; 121: 19–25.
Montgomery RA, Dietz HC . Inhibition of fibrillin 1 expression using U1 shRNA as a vehicle for the presentation of antisense targeting sequence. Hum Mol Genet 1997; 6: 519–525.
Millington-Ward S et al. Strategems in vitro for gene therapies directed to dominant mutations. Hum Mol Genet 1997; 6: 1415–1426.
Baulcombe D . RNA silencing. Curr Biol 2002; 12: R82–R84.
Kumar-Singh R, Binkley E, Cashman S . Mutation independent knock down of rhodopsin by short hairpin RNAs (shRNA) and replacement by a shRNA-resistant rhodopsin transgene. Invest Ophthalmol Vis Sci 2004; 45: 3482.
Brummelkamp TR, Bernards R, Agami R . A system for stable expression of short interfering RNAs in mammalian cells. Science 2002; 296: 550–553.
Xia H, Mao Q, Paulson HL, Davidson BL . siRNA-mediated gene silencing in vitro and in vivo. Nat Biotechnol 2002; 20: 1006–1010.
Fallaux FJ et al. Characterization of 911: a new helper cell line for the titration and propagation of early region 1-deleted adenoviral vectors. Hum Gene Ther 1996; 7: 215–222.
Liu X, Garriga P, Khorana HG . Structure and function in rhodopsin: correct folding and misfolding in two point mutants in the intradiscal domain of rhodopsin identified in retinitis pigmentosa. Proc Natl Acad Sci USA 1996; 93: 4554–4559.
Lewin AS et al. Ribozyme rescue of photoreceptor cells in a transgenic rat model of autosomal dominant retinitis pigmentosa. Nat Med 1998; 4: 967–971.
Illing ME, Rajan RS, Bence NF, Kopito RR . A rhodopsin mutant linked to autosomal dominant retinitis pigmentosa is prone to aggregate and interacts with the ubiquitin proteasome system. J Biol Chem 2002; 277: 34150–34160.
O’Neill B et al. Ribozyme-based therapeutic approaches for autosomal dominant retinitis pigmentosa. Invest Ophthalmol Vis Sci 2000; 41: 2863–2869.
Sullivan JM, Pietras KM, Shin BJ, Misasi JN . Hammerhead ribozymes designed to cleave all human rod opsin mRNAs which cause autosomal dominant retinitis pigmentosa. Mol Vis 2002; 8: 102–113.
Ke SH, Madison EL . Rapid and efficient site-directed mutagenesis by single-tube ‘megaprimer’ PCR method. Nucleic Acids Res 1997; 25: 3371–3372.
Acknowledgements
We thank Beverly Davidson (University of Iowa) for providing the plasmid pmCMVmpAGFPi, Robert Molday (University of British Columbia) for providing antibody 1D4 and Jeremy Nathans (Johns Hopkins) for providing the human rhodopsin cDNA. We would also like to thank Laura McCullough for technical assistance. This study was supported by grants to RK-S from the National Eye Institute (1R01EY013837 and 1R01EY014991), a career development award from Research to Prevent Blindness and generous funding from The Foundation Fighting Blindness (FFB) and the Knight's Templar Eye Foundation.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Cashman, S., Binkley, E. & Kumar-Singh, R. Towards mutation-independent silencing of genes involved in retinal degeneration by RNA interference. Gene Ther 12, 1223–1228 (2005). https://doi.org/10.1038/sj.gt.3302512
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.gt.3302512
Keywords
This article is cited by
-
Mutation-independent rescue of a novel mouse model of Retinitis Pigmentosa
Gene Therapy (2013)
-
AAV-mediated knockdown of Peripherin-2 in vivo using miRNA-based hairpins
Gene Therapy (2010)
-
Improved Retinal Function in a Mouse Model of Dominant Retinitis Pigmentosa Following AAV-delivered Gene Therapy
Molecular Therapy (2009)
-
AAV-mediated gene therapy for retinal disorders: from mouse to man
Gene Therapy (2008)