Abstract
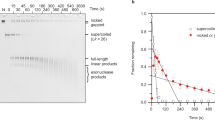
Cruciform structures1,2 in DNA are of considerable interest, both as extreme examples of sequence-dependent structural heterogeneity and as models for four-way junctions such as the Holliday junction3 of homologous genetic recombination. Cruciforms are of lower thermodynamic stability than regular duplex DNA, and have been observed only in negatively supercoiled molecules4–6, where the unfavourable free energy of formation is offset by the topological relaxation of the torsionally stressed molecule. From an experimental viewpoint this can be a disadvantage, as cruciform structures can be studied only in relatively large supercoiled DNA circles, and are destabilized when a break is introduced at any point. We therefore set out to construct a pseudo-cruciform junction—by generating hereroduplex formation between two inverted repeat sequences. Stereochemically, this should closely resemble a true cruciform but remain stable in a linear DNA fragment. We have now created such a junction and find that it has the expected sensitivities to endonucleases. These DNA fragments exhibit extremely anomalous gel electrophoretic mobility, the extent of which depends on the relative position of the pseudo-cruciform along the length of the molecule. Our results are very similar to those obtained by Wu and Crothers7 using kinetoplast DNA, and we conclude that the pseudo-cruciform junction introduces a bend in the linear DNA molecule.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Platt, J. R. Proc. natn. Acad. Sci. U.S.A. 41, 181–183 (1955).
Gierer, A. Nature 212, 1480–1481 (1966).
Holliday, R. Genet. Res. 5, 282–304 (1964).
Gellert, M., Mizuuchi, K., O'Dea, M. H., Ohmori, H. & Tomizawa, J. Cold Spring Harb. Symp. quant. Biol. 43, 35–40 (1979).
Lilley, D. M. J. Proc. natn. Acad. Sci. U.S.A. 77, 6468–6472 (1980).
Panayotatos, N. & Wells, R. D. Nature 289, 466–470 (1981).
Wu, H.-M. & Crothers, D. M. Nature 308, 509–513 (1984).
Mizuuchi, K., Kemper, B., Hays, J. & Weisberg, R. A. Cell 29, 357–365 (1982).
Lilley, D. M. J. & Kemper, B. Cell 36, 413–422 (1984).
Lafer, E. M., Moller, A., Nordheim, A., Stollar, B. D. & Rich, A. Proc. natn. Acad. Sci. U.S.A. 78, 3546–3550 (1981).
Potter, H. & Dressler, D. Proc. natn. Acad. Sci. U.S.A. 75, 3698–3702 (1978).
Bell, L. & Byers, B. Proc. natn. Acad. Sci. U.S.A. 76, 3445–3449 (1979).
Kallenbach, N. R., Ma, R-I. & Seeman, N. C. Nature 305, 829–831 (1983).
Lilley, D. M. J. & Markham, A. F. EMBO J. 2, 527–533 (1983).
Messing, J. & Vieira, J. Gene 19, 269–276 (1982).
Frederick, C. A. et al. Nature 309, 327–331 (1984).
Crick, F. H. C. & Klug, A. Nature 255, 530–533 (1975).
Sobell, H. M., Tsai, C., Gilbert, S. G., Jain, S. C. & Sakore, T. D. Proc. natn. Acad. Sci. U.S.A. 73, 3068–3072 (1976).
Levitt, M. Proc. natn. Acad. Sci. U.S.A. 75, 640–644 (1978).
Sussman, J. L. & Trifonov, E. N. Proc. natn. Acad. Sci. U.S.A. 75, 103–107 (1978).
Twigg, A. J. & Sherratt, D. Nature 283, 216–218 (1980).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Gough, G., Lilley, D. DNA bending induced by cruciform formation. Nature 313, 154–156 (1985). https://doi.org/10.1038/313154a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/313154a0
This article is cited by
-
Ionic interactions and the global conformations of the hammerhead ribozyme
Nature Structural Biology (1995)
-
The sequence-directed bent DNA detected in the replication origin of Chlamydomonas reinhardtii chloroplast DNA is important for the replication function
Molecular and General Genetics MGG (1991)
-
Fluorescence energy transfer shows that the four-way DNA junction is a right-handed cross of antiparallel molecules
Nature (1989)
-
Bent DNA is a structural feature of scaffold-attached regions in Drosophila melanogaster interphase nuclei
Chromosoma (1989)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.