Abstract
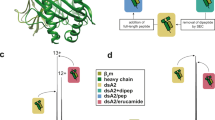
The major histocompatibility complex (MHC)—HLA in man and H–2 in mouse—encodes two classes of cell-surface antigens involved in the immune response. The amino acid sequences have been determined for a number of these molecules1–11. Class I antigens, typified by the HLA–ABC antigens, are composed of a 43,000-molecular weight (MW) glycosylated transmembrane polypeptide with three external domains (α1, α2 and α3), of which the one nearest the membrane (α3) is associated with a 12,000-MW nonglycosylated poly peptide, β2-microglobulin. The HLA-D-region or class II antigens, DR, DC and SB, are composed of two glycosylated transmembrane polypeptides, of MWs 34,000 (α-chain) and 28,000 (β-chain). Both chains have two external domains which presumably associate with each other, α2, β2 being membrane proximal and α1,β1 N-terminal and membrane distal. All four membrane-proximal domains (class I α3, β2-microglobulin, class II α2 and β2) have amino acid sequences that show significant similarities with immunoglobulin constant-region domains3,6,9,12,13. This, together with the similarly placed internal disulphide bonds, suggests they might have an immunoglobulin-like structure (Fig. 1). We have now used computer graphics techniques to predict a detailed three-dimensional structure for the membrane-proximal domains of the class II antigens (α2 and β2) based on the known coordinates of immunoglobulin constant domains (Fig. 2). The transmembrane regions of class II antigens have been modelled as two α-helices packed together. The proposed structure accounts for conservation of amino acids and leads to evolutionary predictions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Ploegh, H. L., Orr, H. T. & Strominger, J. L. Cell 24, 287–299 (1981).
Shackleford, D. A., Kaufman, J. F., Korman, A. J. & Strominger, J. L. Immun. Rev. 66, 133–187 (1982).
Smithies, O. & Poulik, M. D. Science 175, 187–189 (1972).
Coligan, J. E. et al. Proc. natn. Acad. Sci. U.S.A. 75, 3390–3394 (1978).
Kratzin, J. H. et al. Hoppe-Seyler's Z. physiol. Chem. 362, 1665–1669 (1981).
Larhammar, D. et al. Proc. natn. Acad. Sci. U.S.A. 79, 3687–3691 (1982).
Long, E. O., Wake, C. T., Gorski, J. & Mach, B. EMBO J. 2, 389–394 (1983).
Malissen, M., Matissen, B. & Jordan, B. R. Proc. natn. Acad. Sci. U.S.A. 79, 893–897 (1982).
Lee, J. S. et al. Nature 299, 750–752 (1982).
Auffray, C., Korman, A. J., Roux-Dosseto, M., Bono, R. & Strominger, J. L. Proc. natn. Acad. Sci. U.S.A. 79, 6337–6341 (1982).
Benoist, C. O., Mathis, D. J., Kanter, M. R., Williams, V. E. & McDevitt, M. O. Proc. natn. Acad. Sci. U.S.A. 80, 534–538 (1983).
Orr, M. T., Lopez De Castro, J. A., Lancet, D. & Strominger, J. L. Biochemistry 18, 5711–5719 (1979).
Larhammar, D. et al. Scand. J. Immun. 14, 617–622 (1981).
Poljak, R. J. et al. Proc. natn. Acad. Sci. U.S.A. 75, 6002–6006 (1973).
Chou, P. Y. & Fasman, G. D. Adv. Enzym. 47, 45–148 (1978).
Garnier, J., Osguthorpe, D. J. & Robson, B. J. molec. Biol. 120, 97–120 (1978).
Jones, T. A. J. appl. Crystallogr. 11, 268–272 (1978).
Edmundson, A. B., Ely, K. R., Abola, E. E., Schiffer, M. & Panagiotopoulos, N. Biochemistry 18, 3953–3961 (1975).
Tanford, C. Science 200, 1012–1018 (1978).
Rogers, J. et al. Cell 26, 19–27 (1981).
Crick, F. H. C. Acta crystallogr. 6, 689–697 (1953).
Chothia, C., Levitt, M. & Richardson, D. J. molec. Biol. 145, 215–262 (1981).
Gething, M. J., Bye, J., Skehel, J. & Waterfield, M. D. Nature 287, 301–306 (1981).
Gething, M. J., White, J. M. & Waterfield, M. D. Proc. natn. Acad. Sci. U.S.A. 75, 2737–2740 (1978).
Furthmayer, M., Galardy, R. E., Domita, M. & Marchesi, V. T. Archs Biochem. Biophys. 185, 21–29(1978).
Hildemann, W. H. in Comprehensive Immunogenetics (eds Hildemann, W. H., Clark, E. A. & Raison, R. L.) 302–346 (Blackwell, Oxford, 1981).
Humphreys, T. Nature 228, 685–686 (1970).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Travers, P., Blundell, T., Sternberg, M. et al. Structural and evolutionary analysis of HLA-D-region products. Nature 310, 235–238 (1984). https://doi.org/10.1038/310235a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/310235a0
This article is cited by
-
Ancient features of the MHC class II presentation pathway, and a model for the possible origin of MHC molecules
Immunogenetics (2019)
-
Solid-State NMR Investigations of the MHC II Transmembrane Domains: Topological Equilibria and Lipid Interactions
The Journal of Membrane Biology (2019)
-
MHC class II interaction with CD4 mediated by a region analogous to the MHC class I binding site for CD8
Nature (1992)
-
Interisotypic α- and β-chain assembly as a source of HLA class-II diversity
Immunologic Research (1990)
-
A hypothetical model of the foreign antigen binding site of Class II histocompatibility molecules
Nature (1988)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.