Abstract
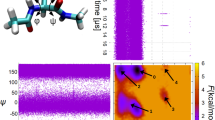
Comparisons between molecular dynamics simulations of proteins and experiment have been based on temperature factors1–5 derived from X-ray spectra and on the stability of hydrogen bonds6. Here we present a novel method of testing molecular dynamics simulations against nuclear magnetic resonance (NMR) relaxation measurements, based on the recently developed model-free approach7 to the interpretation of NMR data. As NMR relaxation in proteins is determined by dynamics on the picosecond–nanosecond time scale, a comparison of NMR experiments and molecular dynamics simulations that last for less than ∼100ps is more meaningful than previous ones3–6 involving much longer time scales. We make contact between a molecular dynamics simulation8 and a 13C-NMR relaxation study9 of pancreatic trypsin inhibitor (PTI) by comparing generalized order parameters (which are measures of the extent of angular motion of the bonds) extracted from the relaxation data9 with those calculated10 from a 96-ps trajectory8. We show that the theoretical order parameters indicate less motion than their experimental counterparts. The relative flexibility of the residues studied, however, is reasonably well described by the simulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Frauenfelder, H., Petsko, G. & Tsernoglu, A. Nature 280, 558–563 (1979).
Artymiuk, P. J. et al. Nature 280, 563–568 (1979).
Northrup, S. H., Pear, M. R., Morgan, J. D., McCammon, J. A. & Karplus, M. J. molec. Biol. 153 1087–1109 (1981).
Northrup, S. H., Pear, M. R., McCammon, J. A., Karplus, M. & Takano, T. Nature 287, 659–660 (1980).
Van Gunsteren, W. F. & Karplus, M. Nature 293, 677–678 (1981).
Levitt, M. Nature 294, 379–380 (1981).
Lipari, G. & Szabo, A. J. Am. chem. Soc. 104, 4546–4570 (1982).
Karplus, M. & McCammon, J. A. Nature 277, 578 (1979).
Richarz, R., Nagayama, K. & Wüthrich, K. Biochemistry 19, 5189–5196 (1980).
Levy, R. M., Karplus, M. & McCammon, J. A. J. Am. chem. Soc. 103, 994–996 (1981).
Lehman, M. S., Koetzle, T. F. & Hamilton, W. C. J. Am. chem. Soc. 94, 2657–2660 (1972).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Lipari, G., Szabo, A. & Levy, R. Protein dynamics and NMR relaxation: comparison of simulations with experiment. Nature 300, 197–198 (1982). https://doi.org/10.1038/300197a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/300197a0
This article is cited by
-
A method to construct the dynamic landscape of a bio-membrane with experiment and simulation
Nature Communications (2022)
-
Crankshaft motions of the polypeptide backbone in molecular dynamics simulations of human type-α transforming growth factor
Journal of Biomolecular NMR (1995)
-
Molecular dynamics simulations of cyclosporin A: The crystal structure and dynamic modelling of a structure in apolar solution based on NMR data
Journal of Computer-Aided Molecular Design (1987)
-
Effect of protein packing structure on side-chain methyl rotor conformations
Nature (1984)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.