Abstract
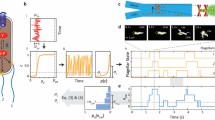
The bacterium Escherichia coli responds to changes in the concentrations of various chemicals in its environment1. A cell swims along a smooth trajectory (runs), moves erratically for a brief time (tumbles) and then runs again, choosing a new direction at random2. If a run happens to carry the cell up a gradient of an attractant (such as aspartate, serine and certain sugars), the occupancy of the appropriate chemoreceptor increases with time3,4 and a signal is sent to the flagellar motors that increases their counterclockwise bias5. On the average, this extends favourable runs and the cell moves up the gradient. The receptors for aspartate and serine6–8 are proteins found in the cytoplasmic membrane, known as methyl-accepting chemotaxis proteins9,10, and are the products of the tar and tsr genes11. A cell can adapt to sustained changes of receptor occupancy by carboxymethylating these proteins12; it is not known, however, how these proteins signal the flagellar motors or how the signal controls the direction of flagellar rotation. Products of several che genes involved in signalling and adaptation have been identified13,14, but with the exception of a methyltransferase15 (the cheR product) and a demethylase16 (the cheB product), their functions are largely unknown. In an attempt to learn more about the events that trigger a chemotactic response, we have now exposed cells to rapid changes in the concentration of attractants and repellents and measured the time required for flagellar reversal. In wild-type cells and in cells containing a cheR–cheB deletion, the response latency is ∼0.2 s. In cheZ mutants, it is much longer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Adler, J. Science 166, 1588–1597 (1969).
Berg, H. C. & Brown, D. A. Nature 239, 500–504 (1972).
Macnab, R. & Koshland, D. E. Jr Proc. natn. Acad. Sci. U.S.A. 69, 2509–2512 (1972).
Brown, D. A. & Berg, H. C. Proc. natn. Acad. Sci. U.S.A. 71, 1388–1392 (1974).
Larsen, S. H., Reader, R. W., Kort, E. N., Tso, W.-W. & Adler, J. Nature 249, 74–77 (1974).
Clarke, S. & Koshland, D. E. Jr J. biol. Chem. 254, 9695–9702 (1979).
Hedblom, M. L. & Adler, J. J. Bact. 144, 1048–1060 (1980).
Wang, E. A. & Koshland, D. E. Jr Proc. natn. Acad. Sci. U.S.A. 77, 7157–7161 (1980).
Kort, E. N., Goy, M. F., Larsen, S. H. & Adler, J. Proc. natn. Acad. Sci. U.S.A. 72, 3939–3943 (1975).
Springer, M. S., Goy, M. F. & Adler, J. Proc. natn. Acad. Sci. U.S.A. 74, 3312–3316 (1977).
Silverman, M. & Simon, M. Proc. natn. Acad. Sci. U.S.A. 74, 3317–3321 (1977).
Springer, M. S., Goy, M. F. & Adler, J. Nature 280, 279–284 (1979).
Parkinson, J. S. A. Rev. Genet. 11, 397–414 (1977).
Silverman, M. & Simon, M. I. A. Rev. Microbiol. 31, 397–419 (1977).
Springer, W. R. & Koshland, D. E. Jr Proc. natn. Acad. Sci. U.S.A. 74, 533–537 (1977).
Stock, J. B. & Koshland, D. E. Jr Proc. natn. Acad. Sci. U.S.A. 75, 3659–3663 (1978).
Silverman, M. & Simon, M. Nature 249, 73–74 (1974).
Kobayasi, S., Maeda, K. & Imae, Y. Rev. scient. Instrum. 48, 407–410 (1977).
Berg, H. C., Manson, M. D. & Conley, M. P. Symp. Soc. exp. Biol. 35, 1–31 (1981).
Berg, H. C. & Tedesco, P. M. Proc. natn. Acad. Sci. U.S.A. 72, 3235–3239 (1975).
Spudich, J. L. & Koshland, D. E. Jr Proc. natn. Acad. Sci. U.S.A. 72, 710–713 (1975).
Crank, J. The Mathematics of Diffusion, 2nd edn (Clarendon, Oxford, 1975).
Purves, R. D. J. Neurosci. Meth. 1, 165–178 (1979).
Muskavitch, M. A., Kort, E. N., Springer, M. S., Goy, M. F. & Adler, J. Science 201, 63–65 (1978).
Berg, H. C. Cold Spring Harb. Conf. Cell Proliferation 3, A47–A56 (1976).
Sherris, D. & Parkinson, J. S. Proc. natn. Acad. Sci. U.S.A. 78, 6051–6055 (1981).
Parkinson, J. S. J. Bact. 135, 45–53 (1978).
Parkinson, J. S. Symp. Soc. gen. Microbiol. 31, 265–290 (1981).
Spudich, J. L. & Koshland, D. E. Jr Nature 262, 467–471 (1976).
Parkinson, J. S. & Parker, S. R. Proc. natn. Acad. Sci. U.S.A. 76, 2390–2394 (1979).
Berg, H. C. & Purcell, E. M. Biophys. J. 20, 193–219 (1977).
Dreyer, F. & Peper, K. Pflügers Arch. ges. Physiol. 348, 263–272 (1974).
Reilley, C. N. & Vavoulis, A. Analyt. Chem. 31, 243–248 (1959).
Schanne, O. F., Lavalée, M., Laprade, R. & Gagné, S. Proc. IEEE 56, 1072–1082 (1968).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Segall, J., Manson, M. & Berg, H. Signal processing times in bacterial chemotaxis. Nature 296, 855–857 (1982). https://doi.org/10.1038/296855a0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/296855a0
This article is cited by
-
Deletion of the cheZ gene results in the loss of swimming ability and the decrease of adhesion ability to Caco-2 cells in Escherichia coli Nissle 1917
Folia Microbiologica (2023)
-
Run and Tumble Chemotaxis in a Shear Flow: The Effect of Temporal Comparisons, Persistence, Rotational Diffusion, and Cell Shape
Bulletin of Mathematical Biology (2009)
-
Phenotype prediction in regulated metabolic networks
BMC Systems Biology (2008)
-
Persistence of direction increases the drift velocity of run and tumble chemotaxis
Journal of Mathematical Biology (2007)
-
Molecular model of a lattice of signalling proteins involved in bacterial chemotaxis
Nature Cell Biology (2000)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.