Abstract
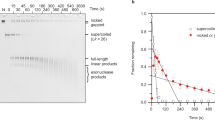
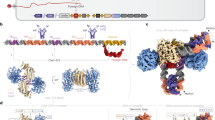
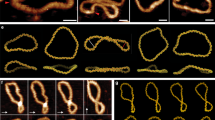
The topology and physical chemistry of closed circular DNA molecules are well understood, but the significance for events within living cells is less well appreciated. It has been demonstrated recently1–3 that the torsional constraint which arises from negative supercoiling (that is, reduction of linkage) can induce localized novel secondary structure in isolated plasmid and phage DNA. Inverted repeats adopt hairpin-loop structures not found in relaxed DNA. This structural perturbation might be expected to have functional significance within the living cell, but clearly this requires that the torsional free energy be available for unhindered partition between alterations of twist and writhe. Microheterogeneity in DNA structure has recently attracted considerable interest, especially with regard to left-handed sections of duplex4–6. The inverted repeats identified as sites of hairpin formation are relatively small, with stems of 13 base pairs (bp) or less. Whilst these hairpins could result in a relaxation of ∼10% of the plasmid supercoiling energy, it was of considerable interest to try to construct stem–loop features about 10 times larger so as to study the topological consequences. In the cloning experiment described here, designed to produce direct or inverted 130-bp repeats depending on insertional orientation, no inverse species could be discovered, and deletion events were frequent. It is concluded that the inverted repeat deprives Escherichia coli of its antibiotic resistance. Cruciform adoption by the inverted species can totally relax the torsional constraint in the plasmid. These experiments highlight the importance of topological considerations in the genetics of closed circular DNA, and confirm the availability of torsional constraint in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Lilley, D. M. J. Proc. natn. Acad. Sci. U.S.A. 77, 6468–6472 (1980).
Panayotatos, N. & Wells, R. D. Nature 289, 466–470 (1981).
Lilley, D. M. J. Nucleic Acids Res. 9, 1271–1289 (1981).
Wang, H-K. et al. Nature 282, 680–686 (1979).
Drew, H., Takano, T., Tanaka, S., Itakura, K. & Dickerson, R. E. Nature 286, 567–573 (1980).
Arnott, S., Chandrasekaran, R., Birdsall, D. L., Leslie, A. G. W. & Ratliff, R. L. Nature 283, 743–745 (1980).
Twigg, A. J. & Sherratt, D. Nature 283, 216–218 (1980).
Birnboim, H. C. & Doly, J. Nucleic Acids Res. 7, 1513–1523 (1979).
Wang, J. C. Proc. natn. Acad. Sci. U.S.A. 76, 200–203 (1979).
Vinograd, J. & Lebowitz, J. J. gen. Physiol. 49, 103–125 (1966).
Fuller, F. B. Proc. natn. Acad. Sci. U.S.A. 68, 815–819 (1971).
Shishido, K. FEES Lett. 111, 333–336 (1980).
Bolivar, F. et al. Proc. natn. Acad. Sci. U.S.A. 74, 5265–5269 (1977).
Sadler, J. R. et al. Gene 3, 211–232 (1978).
Gellert, M., Mizuuchi, K., O'Dea, M. H., Ohmori, H. & Tomizawa, J. Cold Spring Harb. Symp. quant. Biol. 43, 35–40 (1978).
Behnke, K., Malke, H., Hartmann, M. & Walter, F. Plasmid 2, 605–616 (1979).
Wang, J. C. J. molec. Biol. 87, 797–816 (1974).
Richardson, J. P. Biochemistry 13, 3164–3169 (1974).
Ullrich, A. et al. Science 196, 1313–1319 (1977).
Maniatis, T., Jeffrey, A. & Van de Sande, H. Biochemistry 14, 3787–3794 (1975).
Maxam, A. M. & Gilbert, W. Proc. natn. Acad. Sci. U.S.A. 74, 560–564 (1977).
Boyer, H. W. & Roullard-Dussoix, D. J. molec. Biol 41, 459–472 (1969).
Cohen, S. N., Chang, A. C. Y. & Hsu, L. Proc. natn. Acad. Sci. U.S.A. 69, 2110–2114 (1972).
Sutcliffe, J. G. Cold Spring Harb. Symp. quant. Biol. 43, 77–90 (1979).
Katz, L., Kingsbury, D. K. & Helinski, D. R. J. Bact. 114, 557–591 (1973).
Keller, W. Proc. natn. Acad. Sci. U.S.A. 72, 4876–4880 (1975).
Shure, M. & Vinograd, J. Cell 8, 215–226 (1976).
Sharp, P. A., Sugden, B. & Sambrook, J. Biochemistry 12, 3055–3063 (1973).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Lilley, D. In vivo consequences of plasmid topology. Nature 292, 380–382 (1981). https://doi.org/10.1038/292380a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/292380a0
This article is cited by
-
Termini and telomeres in T-DNA transformation
Plant Molecular Biology (1994)
-
A simple method to make better probes from short DNA fragments
Molecular Biotechnology (1994)
-
Formation of deletion in Escherichia coli between direct repeats located in the long inverted repeats of a cellular slime mold plasmid: Participation of DNA gyrase
Molecular and General Genetics MGG (1988)
-
Heterogeneity in the level of ampicillin resistance conferred by pBR322 derivatives with different DNA supercoiling
Molecular and General Genetics MGG (1987)
-
Bacterial chromatin: A new twist to an old story
Nature (1986)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.