Abstract
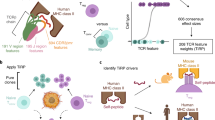
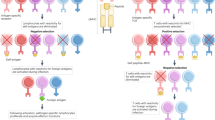
THE most exciting aspect of immunology at the moment is the way in which the major histocompatibility complex regulates certain functions of the immune system. Understanding the basis of this will revolutionise our thinking about T cell responsiveness and perhaps explain the puzzling degree of genetic polymorphism of the major histocompatibility complex. Most rewarding also will be the clinical applications, since susceptibility to many diseases is linked to HL-A (refs 1 and 2). Genes which control the response of thymus-derived (T) lymphocytes to certain antigens map in the major histocompatibility complex1; 1–10% of T lymphocytes respond in an allogeneic mixed lymphocyte culture, or graft-versus-host reaction, against non-self major histocompatibility antigens3–5; cytotoxic T cells derived from mixed lymphocyte cultures seem to be directed against the classical serologically defined antigens coded in this region6,7. The work described here derives from another recently reported link between murine T cell function and the major histocompatibility complex. It has now been demonstrated that T-cell mediated lysis of virus infected target cells is restricted by H-2 (refs 8–10). Lysis of TNP modified targets is similarly restricted in another system11. I have now shown that cytotoxic T cells derived from mice immunised with cells from an allogeneic strain which carries the same H-2 region will lyse only targets which share the same H-2 as the immunising strain.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
McDevitt, H. O., and Landy, M. M., (eds) Genetic Control of Immune Responsiveness: Relationship to Disease Susceptibility, (Academic, New York, 1972).
Morris, P. J., Contemporary Topics in Immunobiology (edit. by Cooper, M. D., and Warner N. L.), 3, 141–169 (Plenum, New York, 1974).
Wilson, D. B., Blyth, J. L., and Nowell, P. C., J. exp. Med., 128, 1157–1181 (1968).
Atkins, R. C., and Ford, W. L., J. exp. Med., 141, 664–680 (1975).
Ford, W. L., Simmonds, S. J., and Atkins, R. C., ibid. 681–696.
Nabholz, M., Vives, J., Young, H. M., Meo, T., Miggiano, V., Rinjbeck, A., and Shreffler, D. C., Eur. J. Immun., 4, 378–387 (1974).
Bevan, M. J., J. Immun., 114, 316–319 (1975).
Zinkernagel, R. M., and Doherty, P. C., Nature, 248, 701–702 (1974).
Doherty, P. C., and Zinkernagel, R. M., J. exp. Med., 141, 502–507 (1975).
Blanden, R. V., in Progress in Immunology, 2 (edit. by Brent, L., and Holborow, J.) 4, 117–125 (1974).
Shearer, G. M., Eur. J. Immunol., 4, 527–533 (1974).
Klein, J., Transplantation, 15, 137–153 (1973).
Bevan, M. J., and Cohn, M., J. Immun., 114, 559–565 (1975).
Ortiz de Landazuri, M., and Herberman, R. B., Nature new Biol., 238, 18–19, (1972).
Zinkernagel, R. M., and Doherty, P. C., Nature, 251, 547–548 (1974).
Gardner, I. D., Bowern, N. A., and Blanden, R. V., Eur. J. Immunol., 5, 122–127 (1975).
Cerottini, J. C., and Brunner, K. T., Adv. Immun., 18, 67–132 (1974).
Graff, R. J., and Bailey, D. W., Transplant. Rev., 15, 26–49 (1973).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
BEVAN, M. Interaction antigens detected by cytotoxic T cells with the major histocompatibility complex as modifier. Nature 256, 419–421 (1975). https://doi.org/10.1038/256419a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/256419a0
This article is cited by
-
Cross-priming
Nature Immunology (2006)
-
A possible association between basic amino acids of position 13 of DRB1 chains and autoimmune hepatitis
Immunogenetics (1992)
-
Identification of classical minor histocompatibility antigen as cell-derived peptide
Nature (1990)
-
Cellular peptide composition governed by major histocompatibility complex class I molecules
Nature (1990)
-
T cell-recognized antigenic peptides derived from the cellular genome are not protein degradation products but can be generated directly by transcription and translation of short subgenic regions. A hypothesis
Immunogenetics (1989)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.