Abstract
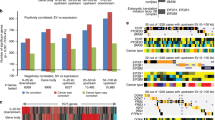
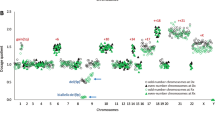
Seventeen ETV6/RUNX1-positive pediatric acute lymphoblastic leukemias were investigated by high-resolution array-based comparative genomic hybridization (array CGH), gene expression profiling and fluorescence in situ hybridization. Comparing the array CGH and gene expression patterns revealed that genomic imbalances conferred a great impact on the expression of genes in the affected regions. The array CGH analyses identified a high frequency of cytogenetically cryptic genetic changes, for example, del(9p) and del(12p). Interestingly, a duplication of Xq material, varying between 30 and 60 Mb in size, was found in 6 of 11 males (55%), but not in females. Genes on Xq were found to have a high expression level in cases with dup(Xq); a similar overexpression was confirmed in t(12;21)-positive cases in an external gene expression data set. By studying the expression profile and the proposed function of genes in the minimally gained region, several candidate target genes (SPANXB, HMGB3, FAM50A, HTATSF1 and RAP2C) were identified. Among them, the testis-specific SPANXB gene was the only one showing a high and uniform overexpression, irrespective of gender and presence of Xq duplication, suggesting that this gene plays an important pathogenetic role in t(12;21)-positive leukemia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Gustafsson G, Schmiegelow K, Forestier E, Clausen N, Glomstein A, Jonmundsson G et al. Improving outcome through two decades in childhood ALL in the Nordic countries: the impact of high-dose methotrexate in the reduction of CNS irradiation. Nordic Society of Pediatric Haematology and Oncology (NOPHO). Leukemia 2000; 14: 2267–2275.
Golub TR, Barker GF, Bohlander SK, Hiebert SW, Ward DC, Bray-Ward P et al. Fusion of the TEL gene on 12p13 to the AML1 gene on 21q22 in acute lymphoblastic leukemia. Proc Natl Acad Sci USA 1995; 92: 4917–4921.
Romana SP, Mauchauffé M, Le Coniat M, Chumakov I, Le Paslier D, Berger R et al. The t(12;21) of acute lymphoblastic leukemia results in a tel-AML1 gene fusion. Blood 1995; 85: 3662–3670.
Shurtleff SA, Buijs A, Behm FG, Rubnitz JE, Raimondi SC, Hancock ML et al. TEL/AML1 fusion resulting from a cryptic t(12;21) is the most common genetic lesion in pediatric ALL and defines a subgroup of patients with an excellent prognosis. Leukemia 1995; 9: 1985–1989.
Raynaud S, Cave H, Baens M, Bastard C, Cacheux V, Grosgeorge J et al. The 12;21 translocation involving TEL and deletion of the other TEL allele: two frequently associated alterations found in childhood acute lymphoblastic leukemia. Blood 1996; 87: 2891–2899.
Loh ML, Goldwasser MA, Silverman LB, Poon WM, Vattikuti S, Cardoso A et al. Prospective analysis of TEL/AML1-positive patients treated on Dana-Farber Cancer Institute Consortium Protocol 95-01. Blood 2006; 107: 4508–4513.
Harbott J, Viehmann S, Borkhardt A, Henze G, Lampert F . Incidence of TEL/AML1 fusion gene analyzed consecutively in children with acute lymphoblastic leukemia in relapse. Blood 1997; 90: 4933–4937.
Nakao M, Yokota S, Horiike S, Taniwaki M, Kashima K, Sonoda Y et al. Detection and quantification of TEL/AML1 fusion transcripts by polymerase chain reaction in childhood acute lymphoblastic leukemia. Leukemia 1996; 10: 1463–1470.
Wiemels JL, Cazzaniga G, Daniotti M, Eden OB, Addison GM, Masera G et al. Prenatal origin of acute lymphoblastic leukaemia in children. Lancet 1999; 354: 1499–1503.
Ford AM, Bennett CA, Price CM, Bruin MCA, Van Wering ER, Greaves M . Fetal origins of the TEL-AML1 fusion gene in identical twins with leukemia. Proc Natl Acad USA 1998; 95: 4584–4588.
Mori H, Colman SM, Xiao Z, Ford AM, Healy LE, Donaldson C et al. Chromosome translocations and covert leukemic clones are generated during normal fetal development. Proc Natl Acad USA 2002; 99: 8242–8247.
Fischer M, Schwieger M, Horn S, Niebuhr B, Ford A, Roscher S et al. Defining the oncogenic function of the TEL/AML1 (ETV6/RUNX1) fusion protein in a mouse model. Oncogene 2005; 24: 7579–7591.
Stams WAG, Beverloo HB, den Boer ML, de Menezes RX, Stigter RL, van Drunen E et al. Incidence of additional genetic changes in the TEL and AML1 genes in DCOG and COALL-treated t(12;21)-positive pediatric ALL, and their relation with drug sensitivity and clinical outcome. Leukemia 2006; 20: 410–416.
Forestier E, Andersen MK, Autio K, Blennow E, Borgström G, Golovleva I et al. Cytogenetic patterns in ETV6/RUNX1-positive pediatric B-cell precursor acute lymphoblastic leukemia: a Nordic series of 245 cases and review of the literature. Genes Chromosomes Cancer 2007; 46: 440–450.
Lu XY, Harris CP, Cooley L, Margolin J, Steuber PC, Sheldon M et al. The utility of spectral karyotyping in the cytogenetic analysis of newly diagnosed pediatric acute lymphoblastic leukemia. Leukemia 2002; 16: 2222–2227.
Martineau M, Jalali GR, Barber KE, Broadfield ZJ, Cheung KL, Lilleyman J et al. ETV6/RUNX1 fusion at diagnosis and relapse: some prognostic indications. Genes Chromosomes Cancer 2005; 43: 54–71.
Nordgren A, Heyman M, Sahlén S, Schoumans J, Söderhäll S, Nordenskjöld M et al. Spectral karyotyping and interphase FISH reveal abnormalities not detected by conventional G-banding. Implications for treatment stratification of childhood acute lymphoblastic leukaemia: detailed analysis of 70 cases. Eur J Haematol 2002; 68: 31–41.
Andersson A, Olofsson T, Lindgren D, Nilsson B, Ritz C, Edén P et al. Molecular signatures in childhood acute leukemia and their correlations to expression patterns in normal hematopoietic subpopulations. Proc Natl Acad Sci USA 2005; 102: 19069–19074.
Andreasson P, Höglund M, Békássy AN, Garwicz S, Heldrup J, Mitelman F et al. Cytogenetic and FISH studies of a single center consecutive series of 152 childhood acute lymphoblastic leukemias. Eur J Haematol 2000; 65: 40–51.
Paulsson K, Heidenblad M, Mörse H, Borg Å, Fioretos T, Johansson B . Identification of cryptic aberrations and characterization of translocation breakpoints using array CGH in high hyperdiploid childhood acute lymphoblastic leukemia. Leukemia 2006; 20: 2002–2007.
Heidenblad M, Hallor KH, Staaf J, Jönsson G, Borg Å, Höglund M et al. Genomic profiling of bone and soft tissue tumors with supernumerary ring chromosomes using tiling resolution bacterial artificial chromosome microarrays. Oncogene 2006; 25: 7106–7116.
Ross ME, Zhou X, Song G, Shurtleff SA, Girtman K, Williams WK et al. Classification of pediatric acute lymphoblastic leukemia by gene expression profiling. Blood 2003; 102: 2951–2959.
Andersson A, Ritz C, Lindgren D, Edén P, Lassen C, Heldrup J et al. Microarray-based classification of a consecutive series of 121 childhood acute leukemias: prediction of leukemic and genetic subtype as well as of minimal residual disease status. Leukemia 2007; 21: 1198–1203.
Schaffner C, Stilgenbauer S, Rappold GA, Döhner H, Lichter P . Somatic ATM mutations indicate a pathogenic role of ATM in B-cell chronic lymphocytic leukemia. Blood 1999; 94: 748–753.
Wang Z, Zhang J, Zhang Y, Lim SH . SPAN-Xb expression in myeloma cells is dependent on promoter hypomethylation and can be upregulated pharmacologically. Int J Cancer 2006; 118: 1436–1444.
Mitelman F, Johansson B, Mertens F . Mitelman database of chromosome aberrations in cancer. 2007. http://cgap.nci.nih.gov/Chromosomes/Mitelman.
Ma SK, Wan TSK, Cheuk ATC, Fung LF, Chan GCF, Chan SY et al. Characterization of additional genetic events in childhood acute lymphoblastic leukemia with TEL/AML1 gene fusion: a molecular cytogenetics study. Leukemia 2001; 15: 1442–1447.
Jarošová M, Holzerová M, Mihál V, Lakomá I, Divoký V, Blažek B et al. Complex karyotypes in childhood acute lymphoblastic leukemia: cytogenetic and molecular cytogenetic study of 21 cases. Cancer Genet Cytogenet 2003; 145: 161–168.
Kuchinskaya E, Heyman M, Grandér D, Linderholm M, Söderhäll S, Zaritskey A et al. Children and adults with acute lymphoblastic leukaemia have similar gene expression profiles. Eur J Haematol 2005; 74: 466–480.
Poppe B, Cauwelier B, Van Limbergen H, Yigit N, Philippé J, Verhasselt B et al. Novel cryptic chromosomal rearrangements in childhood acute lymphoblastic leukemia detected by multiple color fluorescent in situ hybridization. Haematologica 2005; 90: 1179–1185.
Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 2007; 446: 758–764.
Gartler SM, Goldman MA . Biology of the X chromosome. Curr Opin Pediatr 2001; 13: 340–345.
Richardson AL, Wang ZC, De Nicolo A, Lu X, Brown M, Miron A et al. X chromosomal abnormalities in basal-like human breast cancer. Cancer Cell 2006; 9: 121–132.
Wang Z, Zhang Y, Liu H, Salati E, Chiriva-Internati M, Lim SH . Gene expression and immunologic consequence of SPAN-Xb in myeloma and other hematologic malignancies. Blood 2003; 101: 955–960.
Kouprina N, Pavlicek A, Noskov VN, Solomon G, Otstot J, Isaacs W et al. Dynamic structure of the SPANX gene cluster mapped to the prostate cancer susceptibility locus HPCX at Xq27. Genome Res 2005; 15: 1477–1486.
Acknowledgements
This work has been supported by grants from the Swedish Children's Cancer Foundation, the Swedish Cancer Society, the Swedish Research Council and the Medical Faculty at Lund University. We thank the Swegene DNA Microarray Resource Center in Lund, supported by the Knut and Alice Wallenberg Foundation through the Swegene consortium, for providing BAC arrays.
The expression data discussed in this publication have been deposited in NCBI's Gene Expression Omnibus (GEO, http://www.ncbi.nlm.nih.gov/geo/) and are accessible through GEO Series accession number GSE7186.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu)
Supplementary information
Rights and permissions
About this article
Cite this article
Lilljebjörn, H., Heidenblad, M., Nilsson, B. et al. Combined high-resolution array-based comparative genomic hybridization and expression profiling of ETV6/RUNX1-positive acute lymphoblastic leukemias reveal a high incidence of cryptic Xq duplications and identify several putative target genes within the commonly gained region. Leukemia 21, 2137–2144 (2007). https://doi.org/10.1038/sj.leu.2404879
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2404879
Keywords
This article is cited by
-
Disruption to the FOXO-PRDM1 axis resulting from deletions of chromosome 6 in acute lymphoblastic leukaemia
Leukemia (2023)
-
High HSPA8 expression predicts adverse outcomes of acute myeloid leukemia
BMC Cancer (2021)
-
KDM3B suppresses APL progression by restricting chromatin accessibility and facilitating the ATRA-mediated degradation of PML/RARα
Cancer Cell International (2019)
-
Chromosomal Aberrations in ETV6/RUNX1-positive Childhood Acute Lymphoblastic Leukemia using 244K Oligonucleotide Array Comparative Genomic Hybridization
Molecular Cytogenetics (2012)
-
Genome-wide profiling of genetic alterations in acute lymphoblastic leukemia: recent insights and future directions
Leukemia (2009)