Abstract
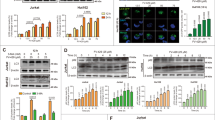
The tyrosine kinase inhibitor imatinib (Gleevec, Novartis Pharmaceuticals Corporation; Basel, Switzerland) is a powerful drug for treatment of chronic myelogenous leukemia (CML) and other malignancies. It selectively targets various tyrosine kinases, thereby leading to growth arrest of respective cancer cells. Given its wide application, it is of high importance to know all related underlying molecular mechanisms. We had previously found that imatinib increases the cellular clearance of intracellular protein aggregates by targeting the abl pathway and thereby upregulating lysosomal activity. Here, we describe that imatinib dose dependently activates the cellular autophagy machinery in mammalian cells, independently of tissue type, species origin or immortalization status of cells. Autophagy is an archetypical cellular degradation mechanism implicated in many physiological and pathophysiological conditions. Our data link for the first time the process of autophagy with the mode of action of imatinib. Induction of autophagy might represent an additional mechanism of imatinib to induce growth arrest, promote apoptosis in cancer cells and eventually even promote tumour regression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Druker BJ, Tamura S, Buchdunger E, Ohno S, Segal GM, Fanning S et al. Effects of a selective inhibitor of the Abl tyrosine kinase on the growth of Bcr-Abl positive cells. Nat Med 1996; 2: 561–566.
Buchdunger E, Zimmermann J, Mett H, Meyer T, Muller M, Druker BJ et al. Inhibition of the Abl protein-tyrosine kinase in vitro by a 2-phenylaminopyrimidine derivative. Cancer Res 1996; 56: 1000–1004.
Heinrich MC, Griffith DJ, Druker BJ, Wait CL, Ott KA, Zigler AJ . Inhibition of c-kit receptor tyrosine kinase activity by STI571, a selective tyrosine kinase inhibitor. Blood 2000; 96: 925–932.
Dewar AL, Zannettino AC, Hughes TP, Lyons AB . Inhibition of c-fms by imatinib: expanding the spectrum of treatment. Cell Cycle 2005; 4: 851–853.
Sawyers CL . Chronic myeloid leukaemia. N Engl J Med 1999; 340: 1330–1340.
Kurzrock R, Kantarjian HM, Drucker BJ, Talpaz M . Philadelphia chromosome-positive leukemias: from basic mechanisms to molecular therapeutics. Ann Int Med 2003; 138: 819–830.
Pendergast AM . The Abl family kinases: mechanisms of regulation and signalling. Adv Cancer Res 2002; 85: 51–100.
Kilic T, Alberta J, Zdunek PR, Acar M, Iannarelli P, O’Reilly T et al. Intracranial inhibition of platelet-derived growth factor-mediated glioblastoma cell growth by an orally active kinase inhibitor of the 2-phenylaminopyrimidine class. Cancer Res 2000; 60: 5143–5150.
Russell J, Brady K, Burgan W, Cerra MA, Oswald KA, Camphausen K et al. STI571-mediated inhibition of Rad51 expression and enhancement of tumor cell radiosensitivity. Cancer Res 2003; 63: 7377–7383.
Netzer WJ, Dou F, Cai D, Veach D, Jean S, Li Y et al. Gleevec inhibits beta-amyloid production but not Notch cleavage. Proc Natl Acad Sci USA 2003; 100: 12444–12449.
Ertmer A, Gilch S, Yun SW, Flechsig E, Klebl B, Stein-Gerlach M et al. The tyrosine kinase inhibitor STI571 induces cellular clearance of PrPSc in prion-infected cells. J Biol Chem 2004; 279: 41918–41927.
Kabeya Y, Mizushima N, Ueno T, Yamamotot A, Kirisako T, Noda T et al. LC3, a mammalian homologue of yeast Apg8p is localized in autophagosome membranes after processing. EMBO J 2000; 19: 5720–5728.
Ito S, Karnovsky MJ . Formaledhyde-glutaraldehyde fixatives containing trinitro compounds. J Cell Biol 1968; 39: 168a–169a.
Blommaart EF, Luiken JJ, Meijer AJ . Autophagic proteolysis: control and specificity. Histochem J 1997; 29: 365–385.
Seglen PO, Bohley P . Autophagy and other vacuolar protein degradation mechanisms. Experientia 1992; 48: 158–172.
Hoyvik H, Gordon PB, Seglen PO . Use of a hydrolysable probe, [14C]lactose, to distinguish between pre-lysosomal and lysosomal steps in the autophagic pathway. Exp Cell Res 1986; 166: 1–14.
Raught B, Gingras AC, Sonenberg N . The target of rapamycin (TOR) proteins. Proc Natl Acad Sci USA 2001; 98: 7037–7044.
Yu JC, Lokker NA, Hollenbach S, Apatira M, Li J, Betz A et al. Efficacy of the novel selective platelet derived growth factor receptor antagonist CT52923 on cellular proliferation, migration, and suppression of neointima following vascular injury. J Pharmacol Exp Ther 2001; 298: 1172–1178.
Sun X, Layton JF, Elefanty A, Lieschke GJ . Comparison of effects of the tyrosine kinase inhibitors AG957, AG490 and STI571 on BCR-ABL-expressing cells, demonstrating synergy between AG490 and STI571. Blood 2001; 97: 2008–2015.
Klionsky DJ, Ohsumi Y . Vacuolar import of proteins and organelles from the cytoplasm. Annu Rev Cell Dev Biol 1999; 15: 1–32.
Yoshimori T . Autophagy : a regulated bulk degradation process inside cells. Biochem Biophys Res Commun 2004; 313: 453–458.
Otsuka H, Moskowitz M . Differences in the rates of protein degradation in untransformed and transformed cell lines. Exp Cell Res 1978; 112: 127–135.
Schwarze PE, Seglen PO . Reduced autophagic activity, improved protein balance and enhanced in vitro survival of hepatocytes isolated from carcinogen-treated rats. Exp Cell Res 1985; 157: 15–28.
Qu X, Yu J, Bhagat G, Furuya N, Hibshoosh H, Troxel A et al. Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J Clin Invest 2003; 112: 1809–1820.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 1999; 402: 672–676.
Cuervo AM . Autophagy: in sickness and in health. Trends Cell biol 2004; 14: 70–77.
Kerr JF, Wyllie AH, Currie AR . Apoptosis: a baxic biological phenomenon with wide-ranging implications in tissue kinetics. Br J Cancer 1972; 26: 239–257.
Syntichaki P, Tavernakis N . Death by necrosis. Uncontrollable catastrophe, or is there order behind the chaos? EMBO Rep 2002; 3: 604–609.
Bursch W, Ellinger A, Kienzl H, Torok L, Pandey S, Sikorska M et al. Active cell death induced by the anti-estrogens tamoxifen and ICI 164 384 in human mammary carcinoma cells (MCF-7) in culture: the role of autophagy. Carcinogenesis 1996; 17: 1595–1607.
Kanzawa T, Germano IM, Komata T, Ito H, Kondo Y, Kondo S . Role of autophagy in temozolomide-inducedcytotoxicity for malignant glioma cells. Cell Death Differ 2004; 11: 448–457.
Paglin S, Hollister T, Delohery T, Hackett N, McMahill M, Sphicas E et al. A novel response of cancer cells to radiation involves autophagy and formation of acidic vesicles. Cancer Res 2001; 61: 439–444.
Ito H, Daido S, Kanzawa T, Kondo S, Kondo Y . Radiation-induced autophagy is associated with LC3 and its inhibition sensitizes malignant glioma cells. Int J Oncol 2005; 26: 1401–1410.
Yao KC, Komata T, Kondo Y, Kanzawa T, Kondo S, Germano IM . Molecular response of human glioblastoma multiforme cells to ionizing radiation: cell cycle arrest modulation of the expression of cyclin-dependent kinase inhibitors, and autophagy. J Neurosurg 2003; 98: 378–384.
Shao Y, Gao Z, Marks PA, Jiang X . Apoptotic and autophagic cell death induced by histone deacetylase inhibitors. Proc Natl Acad Sci USA 2004; 101: 18030–18035.
Komata T, Kanzawa T, Nashimoto T, Aoki H, Endo S, Nameta M et al. Mild heat shock induces autophagic growth arrest, but not apoptosis in U251-MG and U87-MG human malignant glioma cells. J Neurooncol 2004; 68: 101–111.
Kanzawa T, Kondo Y, Ito H, Kondo S, Germano I . Induction of autophagic cell death in malignant glioma cells by arsenic trioxide. Cancer Res 2003; 63: 2103–2108.
Kanzawa T, Zhang L, Xiao L, Germano IM, Kondo Y, Kondo S . Arsenic trioxide induces autophagic cell death in malignant glioma cells by upregulation of mitochondrial cell death protein BNIP3. Oncogene 2005; 24: 980–991.
Opipari Jr AW, Tan L, Boitani AE, Sorenson DR, Aurora A, Liu JR . Resveratrol-induced autophagocytosis in ovarian cancer cells. Cancer Res 2004; 64: 696–703.
Ellington AA, Berhow M, Singletary KW . Induction of macroautophagy in human colon cancer cells by soybean B-group triterpenoid saponins. Carcinogenesis 2005; 26: 159–167.
Takeuchi H, Kondo Y, Fujiwara K, Kanzawa T, Aoki H, Mills GB et al. Synergistic augmentation of rapamycin-induced autophagy in malignant glioma cells by phosphatidylinositol 3-phosphate kinase/protein kinase B inhibitors. Cancer Res 2005; 65: 3336–3346.
Ogier-Denis E, Codogno P . Autophagy: a barrier or an adaptive response to cancer. Biochim Biophys Acta 2003; 1603: 113–128.
Gozuacik D, Kimchi A . Autophagy as a cell death and tumor suppressor mechanism. Oncogene 2004; 23: 2891–2906.
Burchert A, Wang Y, Cai D, von Bubnoff N, Paschka P, Muller-Brusselbach S et al. Compensatory PI3-kinase/Akt/mTor activation regulates imatinib resistance development. Leukemia 2005; 19: 1774–1782.
Tsurutani J, West KA, Sayyah J, Gills JJ, Dennis PA . Inhibition of the phosphatidylinositol 3-kinase/Akt/mammalian target of rapamycin pathway but not the MEK/ERK pathway attenuates laminin-mediated small cell lung cancer cellular survival and resistance to imatinib mesylate or chemotherapy. Cancer Res 2005; 65: 8423–8432.
Acknowledgements
We thank Dr Max Nunziante for helpful discussions and critically reading of the manuscript. This work was supported by the SFB-596 (project A8), FORPRION grant number LMU 5* and the DFG (SCHA 594/3-4), and was performed within the framework of the EU FP6 Network of Excellence Neuroprion.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ertmer, A., Huber, V., Gilch, S. et al. The anticancer drug imatinib induces cellular autophagy. Leukemia 21, 936–942 (2007). https://doi.org/10.1038/sj.leu.2404606
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2404606
Keywords
This article is cited by
-
Extraction yield optimization of Oleaster (Olea europaea var. sylvestris) fruits using response surface methodology, LC/MS profiling and evaluation of its effects on antioxidant activity and autophagy in HFF cells
Journal of Food Measurement and Characterization (2021)
-
Autophagy and autophagy-related proteins in cancer
Molecular Cancer (2020)
-
Role of PARP1-mediated autophagy in EGFR-TKI resistance in non-small cell lung cancer
Scientific Reports (2020)
-
The role of autophagy in intestinal epithelial injury
Pediatric Surgery International (2019)
-
HAMdb: a database of human autophagy modulators with specific pathway and disease information
Journal of Cheminformatics (2018)