Abstract
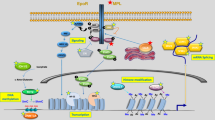
Some myeloproliferative disorders (MPD) result from a reciprocal translocation that involves the FGFR1 gene and a partner gene. The event creates a chimeric gene that encodes a fusion protein with constitutive FGFR1 tyrosine kinase activity. FGFR1-MPD is a rare disease, but its study may provide interesting clues on different processes such as cell signalling, oncogenesis and stem cell renewal. Some partners of FGFR1 are centrosomal proteins. The corresponding oncogenic fusion kinases are targeted to the centrosome. Constitutive phosphorylation at this site may perturbate centrosome function and the cell cycle. Direct attack at this small organelle may be an efficient way for oncogenes to alter regulation of signalling for proliferation and survival and get rid of checkpoints in cell cycle progression. The same effect might be triggered by other fusion kinases in other MPD and non-MPD malignancies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Passegué E, Jamieson CH, Ailles LE, Weissman IL . Normal and leukemic hematopoiesis: are leukemias a stem cell disorder or a reacquisition of stem cell characteristics? Proc Natl Acad Sci USA 2003; 100 (Suppl. 1): 11842–11849.
Gilliland DG . Molecular genetics of human leukemias: new insights into therapy. Semin Hematol 2002; 39: 6–11.
Murati A, Arnoulet C, Lafage-Pochitaloff M, Adélaïde J, Derré M, Slama B et al. Dual lympho-myeloproliferative disorder in a patient with t(8;22) with BCR-FGFR1 gene fusion. Int J Oncol 2005; 26: 1485–1492.
Cross NC, Reiter A . Tyrosine kinase fusion genes in chronic myeloproliferative diseases. Leukemia 2002; 16: 1207–1212.
Chaffanet M, Popovici C, Leroux D, Jacrot M, Adélaïde J, Dastugue N et al. t(6;8), t(8;9) and t(8;13) translocations associated with stem cell myeloproliferative disorders have close or identical breakpoints in chromosome region 8p11–12. Oncogene 1998; 16: 945–949.
Guasch G, Mack GJ, Popovici C, Dastugue N, Birnbaum D, Rattner JB et al. FGFR1 is fused to the centrosome-associated protein CEP110 in the 8p12 stem cell myeloproliferative disorder with t(8;9)(p12;q33). Blood 2000; 95: 1788–1796.
Popovici C, Zhang B, Grégoire MJ, Jonveaux P, Lafage-Pochitlaoof M, Birnbaum D et al. The t(6;8)(q27;p11) translocation in a stem cell myeloproliferative disorder fuses a novel gene, FOP, to fibroblast growth factor receptor 1. Blood 1999; 93: 1381–1389.
Popovici C, Adélaide J, Ollendorff V, Chaffanet M, Guasch G, Jacrot M et al. Fibroblast growth factor receptor 1 is fused to FIM in stem-cell myeloproliferative disorder with t(8;13). Proc Natl Acad Sci USA 1998; 95: 5712–5717.
Xiao S, Nalabolu SR, Aster JC, Ma J, Abruzzo L, Jaffe ES et al. FGFR1 is fused with a novel zinc-finger gene, ZNF198, in the t(8;13) leukaemia/lymphoma syndrome. Nat Genet 1998; 18: 84–87.
Demiroglu A, Steer EJ, Heath C, Taylor K, Bentley M, Allen SL et al. The t(8;22) in chronic myeloid leukemia fuses BCR to FGFR1: transforming activity and specific inhibition of FGFR1 fusion proteins. Blood 2001; 98: 3778–3783.
Fioretos T, Panagopoulos I, Lassen C, Swedin A, Billstrom R, Isaksson M et al. Fusion of the BCR and the fibroblast growth factor receptor-1 (FGFR1) genes as a result of t(8;22)(p11;q11) in a myeloproliferative disorder: the first fusion gene involving BCR but not ABL. Genes Chromosomes Cancer 2001; 32: 302–310.
Ollendorff V, Guasch G, Isnardon D, Galindo R, Birnbaum D, Pébusque MJ . Characterization of FIM-FGFR1, the fusion product of the myeloproliferative disorder-associated t(8;13) translocation. J Biol Chem 1999; 274: 26922–26930.
Xiao S, McCarthy JG, Aster JC, Fletcher J . ZNF198-FGFR1 transforming activity depends on a novel proline-rich ZNF198 oligomerization domain. Blood 2000; 96: 699–704.
Baumann H, Kunapuli P, Tracy E, Cowell JK . The oncogenic fusion protein-tyrosine kinase ZNF198/fibroblast growth factor receptor-1 has signaling function comparable with interleukin-6 cytokine receptors. J Biol Chem 2003; 278: 16198–16208.
Guasch G, Ollendorff V, Borg JP, Birnbaum D, Pébusque MJ . 8p12 stem cell myeloproliferative disorder: the FOP-fibroblast growth factor receptor 1 fusion protein of the t(6;8) translocation induces cell survival mediated by mitogen-activated protein kinase and phosphatidylinositol 3-kinase/Akt/mTOR pathways. Mol Cell Biol 2001; 21: 8129–8142.
Heath C, Cross NC . Critical role of STAT5 activation in transformation mediated by ZNF198-FGFR1. J Biol Chem 2004; 279: 6666–6673.
Chen J, Deangelo DJ, Kutok JL, Williams IR, Lee BH, Wadleigh M et al. PKC412 inhibits the zinc finger 198-fibroblast growth factor receptor 1 fusion tyrosine kinase and is active in treatment of stem cell myeloproliferative disorder. Proc Natl Acad Sci USA 2004; 101: 14479–14484.
Guasch G, Delaval B, Arnoulet C, Xie MJ, Xerri L, Sainty D et al. FOP-FGFR1 tyrosine kinase, the product of a t(6;8) translocation, induces a fatal myeloproliferative disease in mice. Blood 2004; 103: 309–312.
Roumiantsev S, Krause DS, Neumann CA, Dimitri CA, Asiedu F, Cross NC et al. Distinct stem cell myeloproliferative/T lymphoma syndromes induced by ZNF198-FGFR1 and BCR-FGFR1 fusion genes from 8p11 translocations. Cancer Cell 2004; 5: 287–298.
Ou YY, Mack GJ, Zhang M, Rattner JB . CEP110 and ninein are located in a specific domain of the centrosome associated with centrosome maturation. J Cell Sci 2002; 115: 1825–1835.
Andersen JS, Wilkinson CJ, Mayor T, Mortensen P, Nigg EA, Mann M . Proteomic characterization of the human centrosome by protein correlation profiling. Nature 2003; 426: 570–574.
Delaval B, Létard S, Lelièvre H, Chevrier V, Daviet L, Dubreuil P et al. oncogenic kinase of malignant hemopathy targets the centrosome. Cancer Res 2005; 65 (in press).
Emes RD, Ponting CP . A new sequence motif linking lissencephaly, Treacher–Collins and oral-facial-digital type 1 syndromes, microtubule dynamics and cell migration. Hum Mol Genet 2001; 10: 2813–2820.
Tanaka T, Serneo FF, Higgins C, Gambello MJ, Wynshaw-Boris A, Gleeson JG . Lis1 and doublecortin function with dynein to mediate coupling of the nucleus to the centrosome in neuronal migration. J Cell Biol 2004; 165: 709–721.
Kim MH, Cooper DR, Oleksy A, Devedjiev Y, Derewenda U, Reiner O et al. The structure of the N-terminal domain of the product of the lissencephaly gene Lis1 and its functional implications. Structure (Cambridge) 2004; 12: 987–998.
Walz C, Chase A, Schoch C, Weisser A, Schlegel F, Hochhaus A et al. The t(8;17)(p11;q23) in the 8p11 myeloproliferative syndrome fuses MYO18A to FGFR1. Leukemia 2005; 19: 1005–1009.
Belloni E, Trubia M, Gasparini P, Mecucci C, Tapinassi C, Confalioneri S et al. 8p11 myeloproliferative syndrome with a novel t(7;8) translocation leading to fusion of the FGFR1 and TIF1 genes. Genes Chromosomes Cancer 2005; 42: 320–325.
Vizmanos JL, Novo FJ, Roman JP, Baxter EJ, Lahortiga I, Odero MD et al. NIN, a gene encoding a CEP110-like centrosomal protein, is fused to PDGFRB in a patient with a t(5;14)(q33;q24) and an imatinib-responsive myeloproliferative disorder. Cancer Res 2004; 64: 2673–2676.
Abe A, Emi N, Tanimoto M, Terasaki H, Marunouchi T, Saito H . Fusion of the platelet-derived growth factor receptor β to a novel gene CEV14 in acute myelogenous leukemia after clonal evolution. Blood 1997; 90: 4271–4277.
Kulkarni S, Heath C, Parker S, Chase A, Iqbal S, Pocock CF et al. Fusion of H4/D10S170 to the platelet-derived growth factor receptor beta in BCR-ABL-negative myeloproliferative disorders with a t(5;10)(q33;q21). Cancer Res 2000; 60: 3592–3598.
Magnusson MK, Meade KE, Brown KE, Arthur DC, Krueger LA, Barrett AJ et al. Rabaptin-5 is a novel fusion partner to platelet-derived growth factor beta receptor in chronic myelomonocytic leukemia. Blood 2001; 98: 2518–2525.
Morerio C, Acquila M, Rosanda C, Rapella A, Dufour C, Locatelli F et al. HCMOGT-1 is a novel fusion partner to PDGFRB in juvenile myelomonocytic leukemia with t(5;17)(q33;p11.2). Cancer Res 2004; 64: 2649–2651.
Wilkinson K, Velloso ER, Lopes LF, Lee C, Aster JC, Shipp MA et al. Cloning of the t(1;5)(q23;q33) in a myeloproliferative disorder associated with eosinophilia: involvement of PDGFRB and response to imatinib. Blood 2003; 102: 4187–4190.
Levine RL, Wadleigh M, Sternberg DW, Wlodarska I, Galinsky I, Stone RM et al. KIAA1509 is a novel PDGFRB fusion partner in imatinib-responsive myeloproliferative disease associated with a t(5;14)(q33;q32). Leukemia 2005; 19: 27–30.
Grand FH, Burgstaller S, Kuhr T, Baxter EJ, Webersinke G, Thaler J et al. P53-binding protein 1 is fused to the platelet-derived growth factor receptor beta in a patient with a t(5;15)(q33;q22) and an imatinib-responsive eosinophilic myeloproliferative disorder. Cancer Res 2004; 64: 7216–7219.
Gotlib J, Cools J, Malone III JM, Schrier SL, Gilliland DG, Coutre SE et al. The FIP1L1-PDGFRalpha fusion tyrosine kinase in hypereosinophilic syndrome and chronic eosinophilic leukemia: implications for diagnosis, classification, and management. Blood 2004; 103: 2879–2891.
Vandenberghe P, Wlodarska I, Michaux L, Zachee P, Boogaerts M, Vanstraelen D et al. Clinical and molecular features of FIP1L1-PDFGRA (+) chronic eosinophilic leukemias. Leukemia 2004; 18: 734–742.
Infante C, Ramos-Morales F, Fedriani C, Bornens M, Rios RM . GMAP-210, a cis-Golgi network-associated protein, is a minus end microtubule-binding protein. J Cell Biol 1999; 145: 83–98.
Verde I, Pahlke G, Salanova M, Zhang G, Wang S, Coletti D et al. Myomegalin is a novel protein of the Golgi/centrosome that interacts with a cyclic nucleotide phosphodiesterase. J Biol Chem 2001; 276: 11189–11198.
Alberti L, Carniti C, Miranda C, Roccato E, Pierotti MA . RET and NTRK1 proto-oncogenes in human diseases. J Cell Physiol 2003; 195: 168–186.
Wang B, Matsuoka S, Carpenter PB, Elledge SJ . 53BP1, a mediator of the DNA damage checkpoint. Science 2002; 298: 1435–1438.
Reiter A, Walz C, Watmore A, Schoch C, Blau I, Schlegelberger B et al. The t(8;9)(p22;p24) is a recurrent abnormality in chronic and acute leukemia that fuses PCM1 to JAK2. Cancer Res 2005; 65: 2662–2667.
Bousquet M, Quelen C, De Mas V, Duchayne E, Roquefeuil B, Delsol G et al. The t(8;9)(p22;p24) translocation in atypical chronic myeloproliferative disorders yields a new PCM1-JAK2 fusion gene. Oncogene (in press).
Murati A, Gelsi-Boyer V, Adélaïde J, Perot C, Talmant P, Giraudier S et al. PCM1-JAK2 fusion in myeloproliferative disorders and acute erythtroid leukemia with t(8;9) translocation. Leukemia 2005, 21 July [E-pub ahead of print].
Faruki S, Geahlen RL, Asai DJ . Syk-dependent phosphorylation of microtubules in activated B-lymphocytes. J Cell Sci 2000; 113: 2557–2565.
Takahashi S, Inatome R, Hotta A, Qin Q, Hackenmiller R, Simon MC et al. Role for Fes/Fps tyrosine kinase in microtubule nucleation through is Fes/CIP4 homology domain. J Biol Chem 2003; 278: 49129–49133.
Steindler C, Li Z, Algarte M, Alcover A, Libri V, Ragimbeau J et al. Jamip1 (marlin-1) defines a family of proteins interacting with janus kinases and microtubules. J Biol Chem 2004; 279: 43168–43177.
Kuno Y, Abe A, Emi N, Iida M, Yokozawa T, Towatari M et al. Constitutive kinase activation of the TEL-Syk fusion gene in myelodysplastic syndrome with t(9;12)(q22;p12). Blood 2001; 97: 1050–1055.
Baxter EJ, Scott LM, Campbell PJ, East C, Fourouclas N, Swanton S et al. Acquired mutation of the tyrosine kinase JAK2 in human myeloproliferative disorders. Lancet 2005; 365: 1054–1061.
Kralovics R, Passamonti F, Buser AS, Teo SS, Tiedt R, Passweg JR et al. A gain-of-function mutation of JAK2 in myeloproliferative disorders. N Engl J Med 2005; 352: 1779–1790.
Levine RL, Wadleigh M, Cools J, Ebert BL, Wernig G, Huntly BJ et al. Activating mutation in the tyrosine kinase JAK2 in polycythemia vera, essential thrombocythemia, and myeloid metaplasia with myelofibrosis. Cancer Cell 2005; 7: 387–397.
James C, Ugo V, Le Couedic JP, Staerk J, Delhommeau F, Lacout C et al. A unique clonal JAK2 mutation leading to constitutive signalling causes polycythaemia vera. Nature 2005; 434: 1144–1148.
Tuveson DA, Fletcher JA . Signal transduction pathways in sarcoma as targets for therapeutic intervention. Curr Opin Oncol 2001; 13: 249–255.
Kutok JL, Aster JC . Molecular biology of anaplastic lymphoma kinase-positive anaplastic large-cell lymphoma. J Clin Oncol 2002; 20: 3691–3702.
Ventura RA, Martin-Subero JI, Knippschild U, Gascoyne RD, Delsol G, Mason DY et al. Centrosome abnormalities in ALK-positive anaplastic large-cell lymphoma. Leukemia 2004; 18: 1910–1911.
Falini B, Mecucci C, Tiacci E, Alcalay M, Rosati R, Pasqualucci L et al. Cytoplasmic nucleophosmin in acute myelogenous leukemia with a normal karyotype. N Engl J Med 2005; 352: 254–266.
Xu ZX, Zou WX, Lin P, Chang KS . A role for PML3 in centrosome duplication and genome stability. Mol Cell 2005; 17: 721–732.
Corvi R, Berger N, Balczon R, Romeo G . RET/PCM-1: a novel fusion gene in papillary thyroid carcinoma. Oncogene 2000; 19: 4236–4242.
Tognon C, Knezevich SR, Huntsman D, Roskelley CD, Melnyk M, Mathers JA et al. Expression of the ETV6-NTRK3 gene fusion as a primary event in human secretory breast carcinoma. Cancer Cell 2002; 2: 367–376.
Kramer A, Neben K, Ho AD . Centrosomal aberrations in hematological malignancies. Cell Biol Int 2005; 29: 376–384.
Nigg EA . Centrosome aberrations: cause or consequence of cancer progression? Nat Rev Cancer 2002; 2: 815–825.
Storchova Z, Pellman D . From polyploidy to aneuploidy, genome instability and cancer. Nat Rev Mol Cell Biol 2004; 5: 45–54.
Brinkley BR . Managing the centrosome numbers game: from chaos to stability in cancer cell division. Trends Cell Biol 2001; 11: 18–21.
Neben K, Tews B, Wrobel G, Hahn M, Kokocinski F, Giesecke C et al. Gene expression patterns in acute myeloid leukemia correlate with centrosome aberrations and numerical chromosome changes. Oncogene 2004; 23: 2379–2384.
Kramer A, Schweizer S, Neben K, Giesecke C, Kalla J, Katzenberger T et al. Centrosome aberrations as a possible mechanism for chromosomal instability in non-Hodgkin's lymphoma. Leukemia 2003; 17: 2207–2213.
Joos S, Kupper M, Ohl S, von Bonin F, Mechtersheimer C, Bentz M et al. Genomic imbalances including amplification of the tyrosine kinase gene JAK2 in CD30+ Hodgkin cells. Cancer Res 2000; 60: 549–552.
Giehl M, Fabarius A, Frank O, Hochhaus A, Hafner M, Hehlmann R et al. Centrosome aberrations in chronic myeloid leukemia correlate with stage of disease and chromosomal instability. Leukemia 2005; 19: 1192–1197.
Hinchcliffe EH, Miller FJ, Cham M, Khodjakov A, Sluder G . Requirement of a centrosomal activity for cell cycle progression through G1 into S phase. Science 2001; 291: 1547–1550.
Khodjakov A, Rieder CL . Centrosomes enhance the fidelity of cytokinesis in vertebrates and are required for cell cycle progression. J Cell Biol 2001; 153: 237–242.
Rieder CL, Faruki S, Khodjakov A . The centrosome in vertebrates: more than a microtubule-organizing center. Trends Cell Biol 2001; 11: 413–419.
Griffin CS . Aneuploidy, centrosome activity and chromosome instability in cells deficient in homologous recombination repair. Mutat Res 2002; 504: 149–155.
Wang Q, Hirohashi Y, Furuuchi K, Zhao H, Liu Q, Zhang H et al. The centrosome in normal and transformed cells. DNA Cell Biol 2004; 23: 475–489.
D’Assoro AB, Lingle WL, Salisbury JL . Centrosome amplification and the development of cancer. Oncogene 2002; 21: 6146–6153.
Matsumoto Y, Maller JL . A centrosomal localization signal in cyclin E required for Cdk2-independent S phase entry. Science 2004; 306: 885–888.
Massague J . G1 cell-cycle control and cancer. Nature 2004; 432: 298–306.
Chang F, Lee JT, Navolanic PM, Steelman LS, Shelton JG, Blalock WL et al. Involvement of PI3K/Akt pathway in cell cycle progression, apoptosis, and neoplastic transformation: a target for cancer chemotherapy. Leukemia 2003; 17: 590–603.
Wakefield JG, Stephens DJ, Tavare JM . A role for glycogen synthase kinase-3 in mitotic spindle dynamics and chromosome alignment. J Cell Sci 2003; 116: 637–646.
Koziczak M, Holbro T, Hynes NE . Blocking of FGFR signaling inhibits breast cancer cell proliferation through downregulation of D-type cyclins. Oncogene 2004; 23: 3501–3508.
Münger K, Duensing S . Human papillomavirus infection and centrosome anomalies in cervical cancer. In: Nigg E (ed). Centrosomes in Development and Disease. New York: Wiley-VCH, 2004, pp 353–370.
Paling NR, Wheadon H, Bone HK, Welham MJ . Regulation of embryonic stem cell self-renewal by phosphoinositide 3-kinase-dependent signaling. J Biol Chem 2004; 279: 48063–48070.
Acknowledgements
Work in our laboratory on MPD is supported by Inserm, Institut Paoli-Calmettes, Association Laurette Fugain and Ministries of Health and Research (Cancéropôle). BD has been successively supported by Ministry of Research, Ligue Nationale Contre le Cancer and Société Française d’Hématologie.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Delaval, B., Lelièvre, H. & Birnbaum, D. Myeloproliferative disorders: the centrosome connection. Leukemia 19, 1739–1744 (2005). https://doi.org/10.1038/sj.leu.2403926
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2403926
Keywords
This article is cited by
-
DLGAP1 directs megakaryocytic growth and differentiation in an MPL dependent manner in hematopoietic cells
Biomarker Research (2019)
-
Centrosomal targeting of tyrosine kinase activity does not enhance oncogenicity in chronic myeloproliferative disorders
Leukemia (2012)
-
Myeloproliferative disorder FOP-FGFR1 fusion kinase recruits phosphoinositide-3 kinase and phospholipase Cγ at the centrosome
Molecular Cancer (2008)
-
MassARRAY assay: a more accurate method for JAK2V617F mutation detection in Chinese patients with myeloproliferative disorders
Leukemia (2008)
-
Myeloproliferative disorders: premalignant, stem cell, G1 diseases?
Leukemia (2006)