Abstract
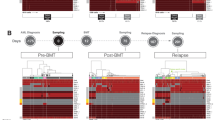
Increasing mixed chimerism (MC) after allogeneic stem cell transplantation (SCT) has been associated with a high risk of relapse in acute leukemia. We evaluated a new method for chimerism detection, based on the quantitative real-time PCR (qrt-PCR) amplification of null alleles or insertion/deletion polymorphisms (indels). All qrt-PCR assays with null alleles and indels attained a sensitivity of at least 10−4, as well as good intra- and interassay concordance, and a high accuracy in experiments with cell mixtures. Informativeness was found in 80.3% of the 61 donor/recipient pairs tested. Nonrelapsed patients showed a progressive decrease in peripheral blood chimerism to values below 0.01% (complete chimerism (CC)). Bone marrow chimerism failed to reach CC more than 4 years after SCT. Increasing MC was observed prior to relapse in 88.2% of patients. Compared with conventional PCR amplification of variable number of tandem repeats, qrt-PCR predicted a significantly higher number of relapses (88.2 vs 44.4%) with a median anticipation period of 58 days. In conclusion, chimerism determination by qrt-PCR amplification of null alleles and indels constitutes a useful tool for the follow-up of patients with acute leukemia after SCT, showing better results than those obtained with conventional PCR.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Kumar L . Leukemia: management of relapse after allogeneic bone marrow transplantation. J Clin Oncol 1994; 12: 1710–1717.
Boiron JM, Lerner D, Pigneux A, Fabères C, Bordessoule D, Turlure P et al. Allogeneic transplantation for patients with advanced acute leukemia: a single centre retrospective study of 92 patients. Leukemia Lymphoma 2001; 41: 285–296.
Bader P, Kreyenberg HW, Dueckers G, Handgretinger R, Lang P et al. Increasing mixed chimerism is an important prognostic factor for unfavorable outcome in children with acute lymphoblastic leukemia after allogeneic stem-cell transplantation: possible role for pre-emptive immunotherapy? J Clin Oncol 2004; 22: 1696–1706.
Keil F, Prinz E, Kalhs P, Lechner K, Moser K, Schwarzinger I et al. Treatment of leukemic relapse after allogeneic stem cell transplantation with cytoreductive chemotherapy and/or immunotherapy or second transplants. Leukemia 2001; 15: 355–361.
Bader P, Beck J, Frey A, Schlegel PG, Hebarth H, Handgretinger R et al. Serial and quantitative analysis of mixed hematopoietic chimerism by PCR in patients with acute leukemias allows the prediction of relapse after allogeneic BMT. Bone Marrow Transplant 1998; 21: 487–495.
Mattson J, Uzunel M, Tammik L, Aschan J, Ringdén O . Leukemia lineage-specific chimerism analysis is a sensitive predictor of relapse in patients with acute myeloid leukemia and myelodysplastic syndrome after allogeneic stem cell transplantation. Leukemia 2001; 15: 1976–1985.
Barrios M, Jiménez-Velasco A, Román-Gómez J, Madrigal E, Castillejo JA, Torres A et al. Chimerism status is a useful predictor of relapse alter allogeneic stem cell transplantation for acute leukemia. Haematologica 2003; 88: 801–810.
Bader P, Klingebiel T, Schaudt A, Theurer-Mainka U, Handgretinger R, Lang P et al. Prevention of relapse in pediatric patients with acute leukemias and MDS after allogeneic SCT by early immunotherapy initiated on the basis of increasing mixed chimerism: a single center experience of 12 children. Leukemia 1999; 13: 2079–2086.
Shimoni A, Nagler A . Non-myeloablative stem cell transplantation (NST): chimerism testing as guidance for immune-therapeutic manipulations. Leukemia 2001; 15: 1967–1975.
Schaap N, Schattenberg A, Mensink E, Preijers F, Hillegers M, Knops R et al. Long-term follow-up of persisting mixed chimerism after partially T cell-depleted allogeneic stem cell transplantation. Leukemia 2002; 16: 13–21.
Bertheas MF, Lafage M, Levy P, Blaise D, Stoppa AM, Viens P et al. Influence of mixed chimerism on the results of allogeneic bone marrow transplantation for leukemia. Blood 1991; 78: 3103–3106.
Wittwer C . Rapid cycle real-time PCR: methods and applications. In: Meuer S, Wittwer C, Nakagawara K (eds) Rapid Cycle Real-time PCR: Methods and Applications. Berlin: Springer-Verlag, 2001, pp. 1–8.
Oliver DH, Thompson RE, Griffin CA, Eshleman JR . Use of single nucleotide polymorphisms (SNP) and real-time polymerase chain reaction for bone marrow engraftment analysis. J Mol Diagn 2000; 2: 202–208.
Alizadeh M, Bernard M, Danic B, Dauriac C, Birebent B, Lapart C et al. Quantitative assessment of hematopoietic chimerism after bone marrow transplantation by real-time quantitative polymerase chain reaction. Blood 2002; 99: 4618–4625.
Najfeld V, Burnett W, Vlachos A, Scigliano E, Isola L, Fruchtman S . Interphase FISH analysis of sex-mismatched BMT using dual color XY probes. Bone Marrow Transplant 1997; 19: 829–834.
Peters SO, Bauermeister K, Simon JP, Branke B, Wagner T . Quantitative polymerase chain reaction-based assay with fluorogenic Y-chromosome specific probes to measure bone marrow chimerism in mice. J Immunol Meth 2002; 260: 109–116.
Costa J-M, Benachi A, Gautier E, Jouannic J-M, Ernault P, Dumez Y . First-trimester fetal sex determination in maternal serum using real-time PCR. Prenat Diagn 2001; 21: 1070–1074.
Maas F, Schaap N, Kolen S, Zoetbrood A, Buño I, Dolstra H et al. Quantification of donor and recipient hemopoietic cells by real-time PCR of single nucleotide polymorphisms. Leukemia 2003; 17: 621–629.
Weber JL, David D, Heil J, Fan Y, Zhao C, Marth G . Human diallelic insertion/deletion polymorphisms. Am J Hum Genet 2002; 71: 854–862.
Wilhelm J, Reuter H, Tews B, Pingoud A, Hahn M . Detection and quantification of insertion/deletion variations by allele-specific real-time PCR: application for genotyping and chimerism analysis. Biol Chem 2002; 383: 1423–1433.
Serrano J, Román J, Sánchez J, Jiménez A, Castillejo JA, Herrera C et al. Molecular analysis of lineage-specific chimerism and minimal residual disease by RT-PCR of p210BCR-ABL after allogeneic bone marrow transplantation for chronic myeloid leukemia: increasing mixed myeloid chimerism and p190BCR-ABL detection precede cytogenetic relapse. Blood 2000; 95: 2659–2665.
Ugozzoli L, Yam P, Petz LD, Ferrara GB, Champlin RE, Forman SJ et al. Amplification by the polymerase chain reaction of hypervariable regions of the human genome for evaluation of chimerism after bone marrow transplantation. Blood 1991; 77: 1607–1615.
Stuppia L, Calabrese G, Di Bartolomeo P, Peila R, Franchi PG, Morizio E et al. Retrospective investigation of hematopoietic chimerism after BMT by PCR amplification of hypervariable DNA regions. Cancer Genet Cytogenet 1995; 85: 124–128.
Ko Y, Koch B, Hart V, Sachinidis A, Thier R, Vetter H et al. Rapid analysis of GSTM1, GSTT1 and GSTP1 polymorphisms using real-time polymerase chain reaction. Pharmacogenetics 2000; 10: 271–274.
Pemble S, Schroeder KR, Spencer SR, Meyer DJ, Hallier E, Bolt HM et al. Human glutathione S-transferase theta (GSTT1): cDNA cloning and the characterization of a genetic polymorphism. Biochem J 1994; 300: 271–276.
Costa J-M, Giovangrandi Y, Ernault P, Lohmann L, Nataf V, El Halali N et al. Fetal RHD genotyping in maternal serum during the first trimester of pregnancy. Br J Haematol 2002; 119: 255–260.
Hamdy SI, Hiratsuka M, Narahara K, El-Enany M, Moursi N, Ahmed MS et al. Allele and genotype frequencies of polymorphic DCP1, CETP, ADRB2, and HTR2A in the Egyptian population. Eur J Clin Pharmacol 2002; 58: 29–36.
Di Castelnuovo A, D'Orazio A, Amore C, Falanga A, Donati MB, Iacoviello L . The decanucleotide insertion/deletion polymorphism in the promoter region of the coagulation factor VII gene and the risk of familial myocardial infarction. Thromb Res 2000; 98: 9–17.
Robledo R, Orru S, Sidoti A, Muresu R, Esposito D, Grimaldi MC et al. A 9.1-kb gap in the genome reference map is shown to be a stable deletion/insertion polymorphism of ancestral origin. Genomics 2002; 80: 585–592.
Moya CM, Varela V, Rivolta CM, Mendive FM, Targovnik HM . Identification and characterization of a novel large insertion/deletion polymorphism of 1464 base pair in the human thyroglobulin gene. Thyroid 2003; 13: 319–323.
Pfaffl MW . A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res 2001; 29: 2002–2007.
van Dongen JJ, Macintyre EA, Gabert JA, Delabesse E, Rossi V, Saglio G et al. Standardized RT-PCR analysis of fusion gene transcripts from chromosome aberrations in acute leukemia for detection of minimal residual disease. Report of the BIOMED-1 Concerted Action: investigation of minimal residual disease in acute leukemia. Leukemia 1999; 13: 1901–1928.
Wäsh R, Bertz H, Kunzmann R, Finke J . Incidence of mixed chimaerism and clinical outcome in 101 patients after myeloablative conditioning regimens and allogeneic stem cell transplantation. Br J Haematol 2000; 109: 743–750.
Wang DG, Fan JB, Siao CJ, Berno A, Young P, Sapolsky R et al. Large-scale identification, mapping and genotyping of single-nucleotide polymorphismsin the human genome. Science 1998; 280: 1077–1082.
Thiede C, Florek M, Bornhauser M, Ritter M, Mohr B, Brendel C et al. Rapid quantification of mixed chimerism using multiplex amplification of short tandem repeat marker and fluorescence detection. Bone Marrow Transplant 1999; 23: 1055–1060.
González M, López-Pérez R, García-Sanz R, Pérez-Simón JA, San Miguel JF . Debate round table: comments concerning chimerism studies. Leukemia 2001; 15: 1986–1988.
Lion T, Daxberger H, Dubovsky J, Filipcik P, Fritsch G, Printz D et al. Analysis of chimerism whithin specific leukocyte subsets for detection of residual or recurrent leukemia in pediatric patients after allogeneic stem cell transplantation. Leukemia 2001; 15: 307–310.
Roux E, Heig C, Dumont-Girard F, Chapuis B, Jeannet M, Roosnek E . Analysis of T-cell repopulation after allogeneic bone marrow transplantation: significant differences between recipients of T-cell depleted and unmanipulated grafts. Blood 1996; 87: 3984–3992.
Petit T, Raynal B, Soci G, Landman-Parker J, Bourhis J-H, Gluckman E et al. Highly sensitive polymerase chain reaction methods show the frequent survival of residual recipient multipotent progenitors after non-T-cell-depleted bone marrow transplantation. Blood. 1994; 84: 3575–3583.
Leewen van JE, van Tol MJ, Joosten AM, Wijnen JT, Khan PM, Voosen JM . Mixed T-lymphoid chimerism after allogeneic bone marow transplantation for hematologic malignancies of children is not correlated with relapse. Blood 1993; 82: 1921–1928.
Suttorp M, Schmitz N, Dreger P, Schaub J, Löffler H . Monitoring of chimerism after allogeneic bone marrow transplantation with unmanipulated marrow by use of DNA polymorphisms. Leukemia 1993; 5: 679–687.
Choi S-J, Lee K-H, Lee J-H, Kim S, Chung H-J, Lee J-S et al. Prognostic value of hematopoietic chimerism in patients with acute leukemia after allogeneic bone marrow transplantation: a prospective study. Bone Marrow Transplant 2000; 26: 327–332.
Acknowledgements
We would like to thank the nursing and clinical staff, the medical residents of the Hematology Department, and the Transplantation Unit for taking care of our patients and helping us to collect the present data. We are grateful to Beli Torroba for her useful technical assistance and to Ian Johnstone for help with the English editing of the manuscript. This work was supported by grants 02/1299 and 03/0141 from the Fondo de Investigaciones Sanitarias (FIS) and grants 143/03 and 144/03 from Junta de Andalucía. GN holds fellowship Cajamar-Fundación Hospital Carlos Haya and MB FIS 01/F018
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu).
Supplementary information
Rights and permissions
About this article
Cite this article
Jiménez-Velasco, A., Barrios, M., Román-Gómez, J. et al. Reliable quantification of hematopoietic chimerism after allogeneic transplantation for acute leukemia using amplification by real-time PCR of null alleles and insertion/deletion polymorphisms. Leukemia 19, 336–343 (2005). https://doi.org/10.1038/sj.leu.2403622
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2403622
Keywords
This article is cited by
-
Can novel methods replace the gold standard chimerism method after allogeneic hematopoietic stem cell transplantation?
Annals of Hematology (2023)
-
Beyond chimerism analysis: methods for tracking a new generation of cell-based medicines
Bone Marrow Transplantation (2020)
-
Quantitative chimerism in CD3-negative mononuclear cells predicts prognosis in acute myeloid leukemia patients after hematopoietic stem cell transplantation
Leukemia (2020)
-
Diagnostic value of highly-sensitive chimerism analysis after allogeneic stem cell transplantation
Bone Marrow Transplantation (2018)
-
Chimerism interpretation with a highly sensitive quantitative PCR method: 6 months median latency before chimerism drop below 0.1%
Bone Marrow Transplantation (2016)