Abstract
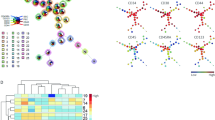
Natural killer (NK) cell-type lymphoproliferative disease of granular lymphocytes (LDGL) is characterized by the outgrowth of CD3−CD16/56+ NK cells, and can be further subdivided into two distinct categories: aggressive NK cell leukemia (ANKL) and chronic NK lymphocytosis (CNKL). To gain insights into the pathophysiology of NK cell-type LDGL, we here purified CD3−CD56+ fractions from healthy individuals (n=9) and those with CNKL (n=9) or ANKL (n=1), and compared the expression profiles of >12 000 genes. A total of 15 ‘LDGL-associated genes’ were identified, and a correspondence analysis on such genes could clearly indicate that LDGL samples share a ‘molecular signature’ distinct from that of normal NK cells. With a newly invented class prediction algorithm, ‘weighted distance method’, all 19 samples received a clinically matched diagnosis, and, furthermore, a detailed cross-validation trial for the prediction of normal or CNKL status could achieve a high accuracy (77.8%). By applying another statistical approach, we could extract other sets of genes, expression of which was specific to either normal or LDGL NK cells. Together with sophisticated statistical methods, gene expression profiling of a background-matched NK cell fraction thus provides us a wealth of information for the LDGL condition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Loughran Jr TP . Clonal diseases of large granular lymphocytes. Blood 1993; 82: 1–14.
Semenzato G, Zambello R, Starkebaum G, Oshimi K, Loughran Jr TP . The lymphoproliferative disease of granular lymphocytes: updated criteria for diagnosis. Blood 1997; 89: 256–260.
Oshimi K . Granular lymphocyte proliferative disorders: report of 12 cases and review of the literature. Leukemia 1988; 2: 617–627.
Loughran Jr TP, Starkebaum G . Large granular lymphocyte leukemia. Report of 38 cases and review of the literature. Medicine (Baltimore) 1987; 66: 397–405.
Jaffe ES, Harris NL, Stein H, Vardiman JW (eds). Pathology and Genetics of Tumours of Haematopoietic and Lymphoid Tissues. Lyon: IARC Press, 2001.
Gelb AB, van de Rijn M, Regula Jr DP, Cornbleet JP, Kamel OW, Horoupian DS et al. Epstein–Barr virus-associated natural killer-large granular lymphocyte leukemia. Hum Pathol 1994; 25: 953–960.
Tefferi A . Chronic natural killer cell lymphocytosis. Leuk Lymphoma 1996; 20: 245–248.
Rabbani GR, Phyliky RL, Tefferi A . A long-term study of patients with chronic natural killer cell lymphocytosis. Br J Haematol 1999; 106: 960–966.
Nash R, McSweeney P, Zambello R, Semenzato G, Loughran Jr TP . Clonal studies of CD3-lymphoproliferative disease of granular lymphocytes. Blood 1993; 81: 2363–2368.
Garcia-Suarez J, Prieto A, Reyes E, Arribalzaga K, Perez-Machado MA, Lopez-Rubio M et al. Persistent lymphocytosis of natural killer cells in autoimmune thrombocytopenic purpura (ATP) patients after splenectomy. Br J Haematol 1995; 89: 653–655.
Ohno Y, Amakawa R, Fukuhara S, Huang CR, Kamesaki H, Amano H et al. Acute transformation of chronic large granular lymphocyte leukemia associated with additional chromosome abnormality. Cancer 1989; 64: 63–67.
Tefferi A, Greipp PR, Leibson PJ, Thibodeau SN . Demonstration of clonality, by X-linked DNA analysis, in chronic natural killer cell lymphocytosis and successful therapy with oral cyclophosphamide. Leukemia 1992; 6: 477–480.
Morice WG, Kurtin PJ, Leibson PJ, Tefferi A, Hanson CA . Demonstration of aberrant T-cell and natural killer-cell antigen expression in all cases of granular lymphocytic leukaemia. Br J Haematol 2003; 120: 1026–1036.
Cheung VG, Morley M, Aguilar F, Massimi A, Kucherlapati R, Childs G . Making and reading microarrays. Nat Genet 1999; 21: 15–19.
Duggan DJ, Bittner M, Chen Y, Meltzer P, Trent JM . Expression profiling using cDNA microarrays. Nat Genet 1999; 21: 10–14.
Dhanasekaran SM, Barrette TR, Ghosh D, Shah R, Varambally S, Kurachi K et al. Delineation of prognostic biomarkers in prostate cancer. Nature 2001; 412: 822–826.
Miyazato A, Ueno S, Ohmine K, Ueda M, Yoshida K, Yamashita Y et al. Identification of myelodysplastic syndrome-specific genes by DNA microarray analysis with purified hematopoietic stem cell fraction. Blood 2001; 98: 422–427.
Ohmine K, Ota J, Ueda M, Ueno S-i, Yoshida K, Yamashita Y et al. Characterization of stage progression in chronic myeloid leukemia by DNA microarray with purified hematopoietic stem cells. Oncogene 2001; 20: 8249–8257.
Makishima H, Ishida F, Ito T, Kitano K, Ueno S, Ohmine K et al. DNA microarray analysis of T cell-type lymphoproliferative disease of granular lymphocytes. Br J Haematol 2002; 118: 462–469.
Van Gelder RN, von Zastrow ME, Yool A, Dement WC, Barchas JD, Eberwine JH . Amplified RNA synthesized from limited quantities of heterogeneous cDNA. Proc Natl Acad Sci USA 1990; 87: 1663–1667.
Fellenberg K, Hauser NC, Brors B, Neutzner A, Hoheisel JD, Vingron M . Correspondence analysis applied to microarray data. Proc Natl Acad Sci USA 2001; 98: 10781–10786.
Alon U, Barkai N, Notterman DA, Gish K, Ybarra S, Mack D et al. Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proc Natl Acad Sci USA 1999; 96: 6745–6750.
Ramaswamy S, Ross KN, Lander ES, Golub TR . A molecular signature of metastasis in primary solid tumors. Nat Genet 2003; 33: 49–54.
Alkema MJ, Wiegant J, Raap AK, Berns A, van Lohuizen M . Characterization and chromosomal localization of the human proto-oncogene BMI-1. Hum Mol Genet 1993; 2: 1597–1603.
Park IK, Qian D, Kiel M, Becker MW, Pihalja M, Weissman IL et al. Bmi-1 is required for maintenance of adult self-renewing haematopoietic stem cells. Nature 2003; 423: 302–305.
Lessard J, Sauvageau G . Bmi-1 determines the proliferative capacity of normal and leukaemic stem cells. Nature 2003; 423: 255–260.
Meagher MJ, Braun RE . Requirement for the murine zinc finger protein ZFR in perigastrulation growth and survival. Mol Cell Biol 2001; 21: 2880–2890.
Thornton S, Kuhn KA, Finkelman FD, Hirsch R . NK cells secrete high levels of IFN-gamma in response to in vivo administration of IL-2. Eur J Immunol 2001; 31: 3355–3360.
Ross ME, Caligiuri MA . Cytokine-induced apoptosis of human natural killer cells identifies a novel mechanism to regulate the innate immune response. Blood 1997; 89: 910–918.
Carson WE, Giri JG, Lindemann MJ, Linett ML, Ahdieh M, Paxton R et al. Interleukin (IL) 15 is a novel cytokine that activates human natural killer cells via components of the IL-2 receptor. J Exp Med 1994; 180: 1395–1403.
Mizuno S, Akashi K, Ohshima K, Iwasaki H, Miyamoto T, Uchida N et al. Interferon-gamma prevents apoptosis in Epstein–Barr virus-infected natural killer cell leukemia in an autocrine fashion. Blood 1999; 93: 3494–3504.
El-Deiry WS, Tokino T, Velculescu VE, Levy DB, Parsons R, Trent JM et al. WAF1, a potential mediator of p53 tumor suppression. Cell 1993; 75: 817–825.
Muda M, Boschert U, Dickinson R, Martinou JC, Martinou I, Camps M et al. MKP-3, a novel cytosolic proteintyrosine phosphatase that exemplifies a new class of mitogen-activated protein kinase phosphatase. J Biol Chem 1996; 271: 4319–4326.
Liston P, Roy N, Tamai K, Lefebvre C, Baird S, Cherton-Horvat G et al. Suppression of apoptosis in mammalian cells by NAIP and a related family of IAP genes. Nature 1996; 379: 349–353.
Iida S, Rao PH, Butler M, Corradini P, Boccadoro M, Klein B et al. Deregulation of MUM1/IRF4 by chromosomal translocation in multiple myeloma. Nat Genet 1997; 17: 226–230.
Woodage T, Basrai MA, Baxevanis AD, Hieter P, Collins FS . Characterization of the CHD family of proteins. Proc Natl Acad Sci USA 1997; 94: 11472–11477.
Acknowledgements
We thank all patients, healthy volunteers, and physicians who participated in the collection of NK cell depository. We are also grateful to Dr T Miwa for his helpful suggestions.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was supported in part by a grant-in-aid for research on the second-term comprehensive 10-year strategy for cancer control from the Ministry of Health, Labor, and Welfare of Japan, by a grant from Mitsubishi Pharma Research Foundation, by a grant from Takeda Science Foundation, and by a grant from Sankyo Foundation of Life Science.
b >Supplementary Information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu).
Rights and permissions
About this article
Cite this article
Choi, Y., Makishima, H., Ohashi, J. et al. DNA microarray analysis of natural killer cell-type lymphoproliferative disease of granular lymphocytes with purified CD3−CD56+ fractions. Leukemia 18, 556–565 (2004). https://doi.org/10.1038/sj.leu.2403261
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2403261
Keywords
This article is cited by
-
A genomic analysis of adult T-cell leukemia
Oncogene (2007)
-
Significance of chemokine receptor expression in aggressive NK cell leukemia
Leukemia (2005)