Abstract
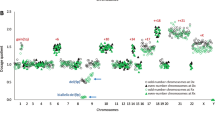
Secondary chromosomal aberrations in follicle center cell derived lymphomas (FCDL) usually involve gains and losses of genetic material and may be an important prognostic value. In the present study, we aimed to determine the power of comparative genomic hybridization (CGH) as compared to standard chromosome analysis (CA) to detect such secondary aberrations. The same lymph node cell suspensions prepared from 30 patients with FCDL were analyzed in parallel by CGH and CA based on R banding. In all, 73 discrepancies were found. Sixty-two imbalances were detected only by CA and 11 only by CGH. In cases with completely resolved karyotypes (n = 17), the median number of discrepancies between CGH and CA was one. However, when the karyotype was partially resolved (n = 12), the median was four (P < 0.01). discrepant results were further studied by fluorescence in situ hybridization using locus-specific probes. These data confirm, that not only for the detection of balanced aberrations, but also for the detection of unbalanced aberrations in FCDL, standard chromosome analysis is still the ‘gold standard’. In contrast, CGH is useful to detect chromosomal imbalances when no metaphases are found or no fresh material is available.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Bentz M, Dohner H, Cabot G, Lichter P . Fluorescence in situ hybridization in leukemias: ‘the FISH are spawning!’ Leukemia 1994 8: 1447–1452
Siebert R, Weber-Matthiesen K . Fluorescence in situ hybridization as a diagnostic tool in malignant lymphomas Histochem Cell Biol 1997 8: 391–402
Mitelman F . Catalog of Chromosome Aberrations in Cancer (sixth edn) Wiley-Liss: New York 1998
Cigudosa JC, Parsa NZ, Louie DC, Filippa DA, Jhanwar SC, Johansson B, Mitelman F, Chaganti RS . Cytogenetic analysis of 363 consecutively ascertained diffuse large B-cell lymphomas Genes Chromosomes Cancer 1999 25: 123–133
Poetsch M, Weber-Matthiesen K, Plendl HJ, Grote W, Schlegelberger B . Detection of the t(14;18) chromosomal translocation by interphase cytogenetics with yeast-artificial-chromosome probes in follicular lymphoma and nonneoplastic lymphoproliferation J Clin Oncol 1996 14: 963–969
Vaandrager JW, Schuuring E, Raap T, Philippo K, Kleiverda K, Kluin P . Interphase FISH detection of BCL2 rearrangement in follicular lymphoma using breakpoint-flanking probes Genes Chromosomes Cancer 2000 27: 85–94
Tilly H, Rossi A, Stamatoullas A, Lenormand B, Bigorgne C, Kunlin A, Monconduit M, Bastard C . Prognostic value of chromosomal abnormalities in follicular lymphoma Blood 1994 84: 1043–1049
Zhang Y, Matthiesen P, Harder S, Siebert R, Castoldi G, Calasanz MJ, Wong KF, Rosenwald A, Ott G, Atkin NB, Schlegelberger B . A 3-cM commonly deleted region in 6q21 in leukemias and lymphomas delineated by fluorescence in situ hybridization Genes Chromosomes Cancer 2000 27: 52–58
Kallioniemi A, Kallioniemi OP, Sudar D, Rutovitz D, Gray JW, Waldman F, Pinkel D . Comparative genomic hybridization for molecular cytogenetic analysis of solid tumors Science 1992 258: 818–821
Forozan F, Karhu R, Kononen J, Kallioniemi A, Kallioniemi OP . Genome screening by comparative genomic hybridization Trends Genet 1997 13: 405–409
Bentz M, Werner CA, Dohner H, Joos S, Barth TF, Siebert R, Schroder M, Stilgenbauer S, Fischer K, Moller P, Lichter P . High incidence of chromosomal imbalances and gene amplifications in the classical follicular variant of follicle center lymphoma Blood 1996 88: 1437–1444
Avet-Loiseau H, Vigier M, Moreau A, Mellerin MP, Gaillard F, Harousseau JL, Bataille R, Milpied N . Comparative genomic hybridization detects genomic abnormalities in 80% of follicular lymphomas Br J Haematol 1997 97: 119–122
Dierlamm J, Rosenberg C, Stul M, Pittaluga S, Wlodarska I, Michaux L, Dehaen M, Verhoef G, Thomas J, de Kelver W, Bakker-Schut T, Cassiman JJ, Raap AK, De Wolf-Peeters C, Van den Berghe H, Hagemeijer A . Characteristic pattern of chromosomal gains and losses in marginal zone B cell lymphoma detected by comparative genomic hybridization Leukemia 1997 11: 747–758
Nacheva EP, Grace CD, Bittner M, Ledbetter DH, Jenkins RB, Green AR . Comparative genomic hybridization: a comparison with molecular and cytogenetic analysis Cancer Genet Cytogenet 1998 100: 93–105
Larramendy ML, Huhta T, Vettenranta K, El-Rifai W, Lundin J, Pakkala S, Saarinen-Pihkala UM, Knuutila S . Comparative genomic hybridization in childhood acute lymphoblastic leukemia Leukemia 1998 12: 1638–1644
Cigudosa JC, Rao PH, Calasanz MJ, Odero MD, Michaeli J, Jhanwar SC, Chaganti RS . Characterization of nonrandom chromosomal gains and losses in multiple myeloma by comparative genomic hybridization Blood 1998 91: 3007–3010
Wilkens L, Tchinda J, Burkhardt D, Nolte M, Werner M, Georgii A . Analysis of hematologic diseases using conventional karyotyping, fluorescence in situ hybridization (FISH), and comparative genomic hybridization (CGH) Hum Pathol 1998 29: 833–839
Schlegelberger B, Zwingers T, Harder S, Nowotny H, Siebert R, Vesely M et al. Clinicopathogenetic significance of chromosomal abnormalities in patients with blastic peripheral B-cell lymphoma Blood 1999 94: 1–8
ISCN . An International System for Human Genetic Nomenclature. Mitelman F (ed) S Karger: Basel 1995
Sambrook J, Frisch EF, Maniatis T . Molecular Cloning: A Laboratory Manual Cold Spring Harbor Laboratory Press: Cold Spring Harbor 1989
Lichter P, Bentz M, du Manoir S, Joss S . Comparative genomic hybridization. In: Verna RS, Babu A (eds) Human Chromosomes McGraw Hill: New York 1995 pp 191–210
Bentz M, Plesch A, Bullinger L, Stilgenbauer S, Ott G, Müller-Hermelink HK, Baudis M, Barth TF, Moller P, Lichter P, Döhner H . t(11;14)-positive mantle cell lymphomas exhibit complex karyotypes and share similarities with B-cell chronic lymphocytic leukemia Genes Chromosomes Cancer 2000 27: 285–294
Schlegelberger B, Metzke S, Harder S, Zulke-Jenisch R, Zhang Y, Siebert R . Classical and molecular cytogenetics of tumor cells. In: Wegener R (ed) Cytogenetics Springer-Verlag: New York 1999 pp 151–185
Johansson B, Mertens F, Mitelman F . Cytogenetic evolution patterns in non-Hodgkin's lymphoma Blood 1995 86: 3905–3914
Wong N, Chen SJ, Cao Q, Su XY, Niu C, Wu QW, Leung TW, Wickham N, Johnson PJ, Chen Z . Detection of chromosome over- and underrepresentations in hyperdiploid acute lymphoblastic leukemia by comparative genomic hybridization Cancer Genet Cytogenet 1998 103: 20–24
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft (Grant Be1454/5–2), the Deutsche Krebshilfe (Grant M 47/95 Be I and Grant 10–1556 Schl4), the Interdisciplinary Center for Clinical Cancer Research of the University of Kiel and the Hermann and Lilly Schilling-Foundation. We would like to thank Maria Jose Calasanz for her critical review of the manuscript and helpful advice and A Borowski, A Schneider, F Kahlert for excellent technical assistance.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Viardot, A., Martin-Subero, J., Siebert, R. et al. Detection of secondary genetic aberrations in follicle center cell derived lymphomas: assessment of the reliability of comparative genomic hybridization and standard chromosome analysis. Leukemia 15, 177–183 (2001). https://doi.org/10.1038/sj.leu.2401969
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2401969