Abstract
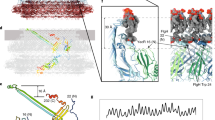
The bacterial chemotaxis receptors are transmembrane receptors with a simple signalling pathway which has elements relevant to the general understanding of signal recognition and transduction across membranes, how signals are relayed between molecules in a pathway, and how adaptation to a persistent signal is achieved1. In contrast to many mammalian receptors which signal by oligomerizing upon ligand binding2, the chemotaxis receptors are dimeric even in the absence of their ligands, and their signalling does not depend on a monomer–dimer equilibrium3. Bacterial chemotaxis receptors are composed of a ligand-binding domain, a transmembrane domain consisting of two helices TM1 and TM2, and a cytoplasmic domain. All known bacterial chemotaxis receptors have a highly conserved cytoplasmic domain, which unites signals from different ligand domains into a single signalling pathway to flagella motors. Here we report the crystal structure of the cytoplasmic domain of a serine chemotaxis receptor of Escherichia coli, which reveals a 200 å-long coiled-coil of two antiparallel helices connected by a ‘U-turn’. Two of these domains form a long, supercoiled, four-helical bundle in the cytoplasmic portion of the receptor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Falke, J. J., Bass, R. B., Butler, S. L., Chervitz, S. A. & Danielson, M. A. The two-component signaling pathway of bacterial chemotaxis: a molecular view of signal transduction by receptors, kinases, and adaptation enzymes. Annu. Rev. Cell Biol. Dev. Biol. 13, 457–512 (1997).
Ullrich, A. & Schlessinger, J. Signal transduction by receptors with tyrosine kinase activity. Cell 61, 203–212 (1990).
Chervitz, S. A. & Falke, J. J. Molecular mechanism of transmembrane signaling by the aspartate receptor: a model. Proc. Natl Acad. Sci. USA 93, 2545–2550 (1996).
Le Moual, H. & Koshland, D. E. J Molecular evolution of the C-terminal cytoplasmic domain of a superfamily of bacterial receptors involved in taxis. J. Mol. Biol. 261, 568–585 (1996).
Cochran, A. G. & Kim, P. S. Imitation of Escherichia coli aspartate receptor signaling in engineered dimers of the cytoplasmic domain. Science 271, 1113–1116 (1996).
Ames, P., Yu, Y. A. & Parkinson, J. S. Methylation segments are not required for chemotactic signalling by cytoplasmic fragments of Tsr, the methyl-accepting serine chemoreceptor of Escherichia coli. Mol. Microbiol. 19, 737–746 (1996).
Surette, M. G. & Stock, J. B. Role of α-helical coiled-coil interactions in receptor dimerization, signaling, and adaptation during bacterial chemotaxis. J. Biol. Chem. 271, 17966–17973 (1996).
Maddock, J. R. & Shapiro, L. Polar location of the chemoreceptor complex in the Escherichia coli cell. Science 259, 1717–1723 (1993).
Liu, Y., Levit, M., Lurz, R., Surette, M. G. & Stock, J. B. Receptor-mediated protein kinase activation and the mechanism of transmembrane signaling in bacterial chemotaxis. EMBO J. 16, 7231–7240 (1997).
Chi, Y. I., Yokota, H. & Kim, S.-H. Apo structure of the ligand-binding domain of aspartate receptor from Escherichia coli and its comparison with ligand-bound or pseudoligand-bound structures. FEBS Lett. 414, 327–332 (1997).
Lynch, B. A. & Koshland, D. E. J The fifth Datta Lecture. Structural similarities between the aspartate receptor of bacterial chemotaxis and the trp repressor of E. coli. Implications for transmembrane signaling. FEBS Lett. 307, 3–9 (1992).
Gardina, P. J. & Manson, M. D. Attractant signaling by an aspartate chemoreceptor dimer with a single cytoplasmic domain. Science 274, 425–426 (1996).
Tatsuno, I., Homma, M., Oosawa, K. & Kawagishi, I. Signaling by the Escherichia coli aspartate chemoreceptor Tar with a single cytoplasmic domain per dimer. Science 274, 423–425 (1996).
Hughson, A. G. & Hazelbauer, G. L. Detecting the conformational change of transmembrane signaling in a bacterial chemoreceptor by measuring effects on disulfide cross-linking in vivo. Proc. Natl Acad. Sci. USA 93, 11546–11551 (1996).
Milburn, M. V. et al. Three-dimensional structures of the ligand-binding domain of the bacterial aspartate receptor with and without a ligand. Science 254, 1342–1347 (1991).
Kim, S.-H. “Frozen” dynamic dimer model for transmembrane signaling in bacterial chemotaxis receptors. Protein Sci. 3, 159–165 (1994).
Maruyama, I. N., Mikawa, Y. G. & Maruyama, H. I. Amodel for transmembrane signaling by the aspartate receptor based on random-cassette mutagenesis and site-directed disulfide cross-linking. J. Mol. Biol. 253, 530–546 (1995).
Borkovich, K. A., Alex, L. A. & Simon, M. I. Attenuation of sensory receptor signaling by covalent modification. Proc. Natl Acad. Sci. USA 89, 6756–6760 (1992).
Ames, P. & Parkinson, J. S. Transmembrane signaling by bacterial chemoreceptors: E. coli transducers with locked signal output. Cell 55, 817–826 (1988).
Ames, P., Chen, J., Wolff, C. & Parkinson, J. S. Structure–function studies of bacterial chemosensors. Cold Spring Harb. Symp. Quant. Biol. 53, 59–65 (1988).
Danielson, M. A., Bass, R. B. & Falke, J. J. Cysteine and disulfide scanning reveals a regulatory α-helix in the cytoplasmic domain of the aspartate receptor. J. Biol. Chem. 272, 32878–32888 (1997).
Bass, R. B. & Falke, J. J. Detection of a conserved α-helix in the kinase docking region of the aspartate receptor by cysteine and disulfide scanning. J. Biol. Chem. 273, 25006–25014 (1998).
Kunkel, T. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Proc. Natl Acad. Sci. USA 82, 488 (1985).
Jancarik, J. & Kim, S.-H. Sparse matrix sampling: a screening method for crystallization of proteins. J. Appl. Crystallogr. 24, 409–411 (1991).
Otwinowski, Z. in Data Collection and Processing (eds Sawyer, L., Isaacs, N. & Bailey, S.) 56–62 (SERC Daresbury Laboratory, Warrington, UK, (1993).
Collaborative Computing Project No. 4. The CCP4 suite: Programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Jones, T. A., Zou, J.-Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for binding protein models inelectron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A. T. X-PLOR Version 3.1 (Yale Univ. Press, New Haven, CT, (1993).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thronton, J. M. Procheck: A program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Kraulis, P. I. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Acknowledgements
We thank R. Sweet, P. Kuhn and H. Bellamy for data collection; D. King for performing the electrospray mass spectrometry; C. Park for plasmid HB915; K. Kamata for help with sample preparation; Z. Zhang and E. Berry for preparing some of the figures; and J. Falke and S. Parkinson for discussion. This work was supported by grants from the Office of Biological and Environmental Research, Office of Science, DOE and the NIH.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kim, K., Yokota, H. & Kim, SH. Four-helical-bundle structure of the cytoplasmic domain of a serine chemotaxis receptor. Nature 400, 787–792 (1999). https://doi.org/10.1038/23512
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/23512
This article is cited by
-
Segmentation strategy of de novo designed four-helical bundles expands protein oligomerization modalities for cell regulation
Nature Communications (2023)
-
Molecular model of a sensor of two-component signaling system
Scientific Reports (2021)
-
Structure and dynamics of the E. coli chemotaxis core signaling complex by cryo-electron tomography and molecular simulations
Communications Biology (2020)
-
Inverted signaling by bacterial chemotaxis receptors
Nature Communications (2018)
-
Helicobacter pylori chemoreceptor TlpC mediates chemotaxis to lactate
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.