Abstract
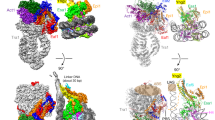
Gene transcription requires the release of inactive DNA from its packaging of histone proteins. Following the discovery of the first transcription-associated histone acetyltransferase, tetrahymena GCN51, it was shown that yeast GCN5 is recruited to the promoter and causes hyper-acetylation of histones and transcriptional activation of target genes2,3, establishing a direct connection between histone acetylation and transcriptional activation. Many other important transcription regulators have been found to have histone acetyltransferase activity, including TAFii230/250, p300/CBP and its associated factor PCAF4,5,6,7,8,9. Here we present the solution structure of the catalytic domain of tGCN5 (residues 47–210) in complex with coenzyme A. The structure contains two domains; the amino-terminal domain is similar to those of other GCN5-related N-acetyltransferases10,11 but the carboxy-terminal domain is not. Coenzyme A binds in a deep hydrophobic pocket between the two domains. Chemical shift changes upon titration with histone H3 peptides indicate a binding site at the domain boundary opposite to the coenzyme A site. The structural data indicate a single-step acetyl-transfer reaction mechanism catalysed by a hydrogen bond to the backbone amide group of leucine 126 and the side-chain carboxyl group of a conserved acidic residue.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Brownell, J. E. et al . Tetrahymena histone acetyltransferase A: a homolog to yeast Gcn5p linking histone acetylation to gene activation. Cell 84, 843–851 (1996).
Kuo, M. H., Zhou, J., Jambeck, P., Churchill, M. E. A. & Allis, C. D. Histone acetyltransferase activity of yeast Gcn5p is required for the activation of target genes in vivo. Genes Dev. 12, 627–639 (1998).
Wang, L. et al . Histone acetyltransferase activity is conserved between yeast and human GCN5 and is required for complementation of growth and transcriptional activation. Mol. Cell. Biol. 17, 519–527 (1997).
Bannister, A. J. & Kouzarides, T. The CBP co-activator is a histone acetyltransferase. Nature 384, 641–643 (1996).
Grant, P. A. et al . Asubset of TAF(II)s are integral components of the SAGA complex required for nucleosome acetylation and transcriptional stimulation. Cell 94, 45–53 (1998).
Mizzen, C. A. et al . The TAF(II)250 subunit of TFIID has histone acetyltransferase activity. Cell 87, 1261–1270 (1996).
Ogryzko, V. V., Schiltz, R. L., Russanova, V., Howard, B. H. & Nakatani, Y. The transcriptional coactivators p300 and CBP are histone acetyltransferases. Cell 87, 953–959 (1996).
Ogryzko, V. V. et al . Histone-like TAFs within the PCAF histone acetylase complex. Cell 94, 35–44 (1998).
Yang, X. J., Ogryzko, V. V., Nishikawa, J., Howard, B. H. & Nakatani, Y. Ap300/CBP-associated factor that competes with the adenoviral oncoprotein E1A. Nature 382, 319–324 (1996).
Wolf, E. et al . Crystal structure of a GCN5-related N-acetyltransferase: Serratia marcescens aminoglycoside 3-N-acetyltransferase. Cell 94, 439–449 (1998).
Dutnall, R. N., Tafrov, S. T., Sternglanz, R. & Ramakrishnan, V. Structure of the histone acetyltransferase Hat1: a paradigm for the GCN5-related N-acetyltransferase superfamily. Cell 94, 427–438 (1998).
Candau, R., Zhou, J. X., Allis, C. D. & Berger, S. L. Histone acetyltransferase activity and interaction with ADA2 are critical for GCN5 function in vivo. EMBO J. 16, 555–565 (1997).
Neuwald, A. F. & Landsman, D. GCN5-related histone N-acetyltransferases belong to a diverse superfamily that includes the yeast SPT10 protein. Trends Biochem. Sci. 22, 154–155 (1997).
Tanner, K. G. et al . Catalytic mechanism and function of invariant glutamic acid-173 from the histone acetyltransferase GCN5 transcriptional activator. J. Biol. Chem. 274, 18157–18160 (1999).
Bhatnagar, R. S. et al . Structure of N-myristoyltransferase with bound myristoyl-CoA and peptide substrate analogs. Nature Struct. Biol. 5, 1091–1097 (1998).
Parthun, M. R., Widom, J. & Gottschling, D. E. The major cytoplasmic histone acetyltransferase in yeast: links to chromatin replication and histone metabolism. Cell 87, 85–94 (1996).
Matsuo, H. et al . Structure of translation factor eIF4E bound to m7GDP and interaction with 4E-binding protein. Nature Struct. Biol. 4, 717–724 (1997).
Fletcher, C. M., Pestova, T. V., Hellen, C. U. T. & Wagner, G. Structure and interactions of the translation initiation factor eIF1. EMBO J. 18, 2631–2637 (1999).
Lin, Y. & Wagner, G. Efficient side chain and backbone assignments in large proteins. J. Am. Chem. Soc. (submitted).
Kraulis, P. J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
We thank R. Marmorstein for initiating this work and providing the histone peptides; G. Heffron for help with NMR experiments; V. Ramakrishnan for providing the coordinates of yHat1 before their release by the PDB; and J. Denu for providing a copy of his paper before publication. This project was supported by grants from NSF, NIH, the Harvard Center for Structural Biology and the Giovanni Armenise-Harvard Foundation for Advanced Scientific Research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lin, Y., Fletcher, C., Zhou, J. et al. Solution structure of the catalytic domain of GCN5 histone acetyltransferase bound to coenzyme A. Nature 400, 86–89 (1999). https://doi.org/10.1038/21922
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/21922
This article is cited by
-
KAT2A coupled with the α-KGDH complex acts as a histone H3 succinyltransferase
Nature (2017)
-
50 years of protein acetylation: from gene regulation to epigenetics, metabolism and beyond
Nature Reviews Molecular Cell Biology (2015)
-
Structure of an acetyl-CoA binding protein from Staphylococcus aureus representing a novel subfamily of GCN5-related N-acetyltransferase-like proteins
Journal of Structural and Functional Genomics (2008)
-
Chemistry of acetyl transfer by histone modifying enzymes: structure, mechanism and implications for effector design
Oncogene (2007)
-
Primers on chromatin
Nature Structural & Molecular Biology (2007)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.