Abstract
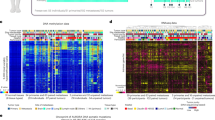
Metastases have been widely thought to arise from rare, selected, mutation-bearing cells in the primary tumor. Recently, however, it has been proposed that breast tumors are imprinted ab initio with metastatic ability. Thus, there is a debate over whether ‘phenotypic’ disease progression is really associated with ‘molecular’ progression. We profiled 26 matched primary breast tumors and lymph node metastases and identified 270 probesets that could discriminate between the two categories. We then used an independent cohort of breast tumors (81 samples) and unmatched distant metastases (32 samples) to validate and refine this list down to a 126-probeset list. A representative subset of these genes was subjected to analysis by in situ hybridization, on a third independent cohort (57 primary breast tumors and matched lymph node metastases). This not only confirmed the expression profile data, but also allowed us to establish the cellular origin of the signals. One-third of the analysed representative genes (4 of 11) were expressed by the epithelial component. The four epithelial genes alone were able to discriminate primary breast tumors from their metastases. Finally, engineered alterations in the expression of two of the epithelial genes (SERPINB5 and LTF) modified cell motility in vitro, in accordance with a possible causal role in metastasis. Our results show that breast cancer metastases are molecularly distinct from their primary tumors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Allinen M, Beroukhim R, Cai L, Brennan C, Lahti-Domenici J, Huang H et al. (2004). Molecular characterization of the tumor microenvironment in breast cancer. Cancer Cell 6: 17–32.
Beltran A, Parikh S, Liu Y, Cuevas BD, Johnson GL, Futscher BW et al. (2007). Re-activation of a dormant tumor suppressor gene maspin by designed transcription factors. Oncogene 26: 2791–2798.
Bezault J, Bhimani R, Wiprovnick J, Furmanski P . (1994). Human lactoferrin inhibits growth of solid tumors and development of experimental metastases in mice. Cancer Res 54: 2310–2312.
Capra M, Nuciforo PG, Confalonieri S, Quarto M, Bianchi M, Nebuloni M et al. (2006). Frequent alterations in the expression of serine/threonine kinases in human cancers. Cancer Res 66: 8147–8154.
Dave SS, Wright G, Tan B, Rosenwald A, Gascoyne RD, Chan WC et al. (2004). Prediction of survival in follicular lymphoma based on molecular features of tumor-infiltrating immune cells. N Engl J Med 351: 2159–2169.
Eberwine J, Yeh H, Miyashiro K, Cao Y, Nair S, Finnell R et al. (1992). Analysis of gene expression in single live neurons. Proc Natl Acad Sci USA 89: 3010–3014.
Feng Y, Sun B, Li X, Zhang L, Niu Y, Xiao C et al. (2007). Differentially expressed genes between primary cancer and paired lymph node metastases predict clinical outcome of node-positive breast cancer patients. Breast Cancer Res Treat 103: 319–329.
Fidler IJ, Kripke ML . (1977). Metastasis results from preexisting variant cells within a malignant tumor. Science 197: 893–895.
Gangnus R, Langer S, Breit E, Pantel K, Speicher MR . (2004). Genomic profiling of viable and proliferative micrometastatic cells from early-stage breast cancer patients. Clin Cancer Res 10: 3457–3464.
Gupta GP, Massague J . (2006). Cancer metastasis: building a framework. Cell 127: 679–695.
Gupta GP, Nguyen DX, Chiang AC, Bos PD, Kim JY, Nadal C et al. (2007). Mediators of vascular remodelling co-opted for sequential steps in lung metastasis. Nature 446: 765–770.
Hao X, Sun B, Hu L, Lahdesmaki H, Dunmire V, Feng Y et al. (2004). Differential gene and protein expression in primary breast malignancies and their lymph node metastases as revealed by combined cDNA microarray and tissue microarray analysis. Cancer 100: 1110–1122.
Hiratsuka S, Nakamura K, Iwai S, Murakami M, Itoh T, Kijima H et al. (2002). MMP9 induction by vascular endothelial growth factor receptor-1 is involved in lung-specific metastasis. Cancer Cell 2: 289–300.
Ivshina AV, George J, Senko O, Mow B, Putti TC, Smeds J et al. (2006). Genetic reclassification of histologic grade delineates new clinical subtypes of breast cancer. Cancer Res 66: 10292–10301.
Kang Y, Siegel PM, Shu W, Drobnjak M, Kakonen SM, Cordon-Cardo C et al. (2003). A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 3: 537–549.
Kononen J, Bubendorf L, Kallioniemi A, Barlund M, Schraml P, Leighton S et al. (1998). Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat Med 4: 844–847.
Kuperwasser C, Chavarria T, Wu M, Magrane G, Gray JW, Carey L et al. (2004). Reconstruction of functionally normal and malignant human breast tissues in mice. Proc Natl Acad Sci USA 101: 4966–4971.
Lamelas ML, Vazquez J, Enguita MI, Rodriguez JC, Gonzalez LO, Merino AM et al. (2000). Apolipoprotein D expression in metastasic lymph nodes of breast cancer. Int J Surg Invest 2: 285–293.
Luo JL, Tan W, Ricono JM, Korchynskyi O, Zhang M, Gonias SL et al. (2007). Nuclear cytokine-activated IKKalpha controls prostate cancer metastasis by repressing Maspin. Nature 446: 690–694.
Minn AJ, Gupta GP, Padua D, Bos P, Nguyen DX, Nuyten D et al. (2007). Lung metastasis genes couple breast tumor size and metastatic spread. Proc Natl Acad Sci USA 104: 6740–6745.
Minn AJ, Gupta GP, Siegel PM, Bos PD, Shu W, Giri DD et al. (2005a). Genes that mediate breast cancer metastasis to lung. Nature 436: 518–524.
Minn AJ, Kang Y, Serganova I, Gupta GP, Giri DD, Doubrovin M et al. (2005b). Distinct organ-specific metastatic potential of individual breast cancer cells and primary tumors. J Clin Invest 115: 44–55.
Muller A, Homey B, Soto H, Ge N, Catron D, Buchanan ME et al. (2001). Involvement of chemokine receptors in breast cancer metastasis. Nature 410: 50–56.
Nguyen DX, Massague J . (2007). Genetic determinants of cancer metastasis. Nat Rev Genet 8: 341–352.
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA et al. (2000). Molecular portraits of human breast tumours. Nature 406: 747–752.
Ramaswamy S, Ross KN, Lander ES, Golub TR . (2003). A molecular signature of metastasis in primary solid tumors. Nat Genet 33: 49–54.
Reddy KB, McGowen R, Schuger L, Visscher D, Sheng S . (2001). Maspin expression inversely correlates with breast tumor progression in MMTV/TGF-alpha transgenic mouse model. Oncogene 20: 6538–6543.
Rolli M, Fransvea E, Pilch J, Saven A, Felding-Habermann B . (2003). Activated integrin alphavbeta3 cooperates with metalloproteinase MMP-9 in regulating migration of metastatic breast cancer cells. Proc Natl Acad Sci USA 100: 9482–9487.
Schedin P, Elias A . (2004). Multistep tumorigenesis and the microenvironment. Breast Cancer Res 6: 93–101.
Schmidt-Kittler O, Ragg T, Daskalakis A, Granzow M, Ahr A, Blankenstein TJ et al. (2003). From latent disseminated cells to overt metastasis: genetic analysis of systemic breast cancer progression. Proc Natl Acad Sci USA 100: 7737–7742.
Shi HY, Zhang W, Liang R, Abraham S, Kittrell FS, Medina D et al. (2001). Blocking tumor growth, invasion, and metastasis by maspin in a syngeneic breast cancer model. Cancer Res 61: 6945–6951.
Suzuki M, Tarin D . (2007). Gene expression profiling of human lymph node metastases and matched primary breast carcinomas: clinical implications. Mol Oncol 1: 172–180.
Ushida Y, Sekine K, Kuhara T, Takasuka N, Iigo M, Tsuda H . (1998). Inhibitory effects of bovine lactoferrin on intestinal polyposis in the Apc(Min) mouse. Cancer Lett 134: 141–145.
van de Vijver MJ, He YD, van’t Veer LJ, Dai H, Hart AA, Voskuil DW et al. (2002). A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 347: 1999–2009.
van’t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M et al. (2002). Gene expression profiling predicts clinical outcome of breast cancer. Nature 415: 530–536.
Wang Y, Klijn JG, Zhang Y, Sieuwerts AM, Look MP, Yang F et al. (2005). Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 365: 671–679.
Weigelt B, Glas AM, Wessels LF, Witteveen AT, Peterse JL, van’t Veer LJ . (2003). Gene expression profiles of primary breast tumors maintained in distant metastases. Proc Natl Acad Sci USA 100: 15901–15905.
Weigelt B, Wessels LF, Bosma AJ, Glas AM, Nuyten DS, He YD et al. (2005). No common denominator for breast cancer lymph node metastasis. Br J Cancer 93: 924–932.
Yamashita K, Upadhyay S, Osada M, Hoque MO, Xiao Y, Mori M et al. (2002). Pharmacologic unmasking of epigenetically silenced tumor suppressor genes in esophageal squamous cell carcinoma. Cancer Cell 2: 485–495.
Yang J, Mani SA, Donaher JL, Ramaswamy S, Itzykson RA, Come C et al. (2004). Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell 117: 927–939.
Zou Z, Anisowicz A, Hendrix MJ, Thor A, Neveu M, Sheng S et al. (1994). Maspin, a serpin with tumor-suppressing activity in human mammary epithelial cells. Science 263: 526–529.
Acknowledgements
We thank the microarray, DNA sequencing and Q-PCR and molecular pathology facilities at the IFOM-IEO-Campus; Silvia Veneroni for metastatic sample selection; Joan Massaguè and Don Nguyen for the 4175 cell line and for helpful discussions; Giorgio Scita and Pascale Romano for helpful discussions and for critically reading the manuscript. This work was supported by grants from the Associazione Italiana per la Ricerca sul Cancro (AIRC) to PPDF, MAP and SP; CNR-MIUR to PPDF and MAP, Italian Ministry of Health to PPDF and SP; FIRB-MIUR to PPDF and MAP; the European Community (FP6) to MAP and PPDF; The Cariplo, Ferrari and Monzino Foundations to PPDF.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Supplementary information
Rights and permissions
About this article
Cite this article
Vecchi, M., Confalonieri, S., Nuciforo, P. et al. Breast cancer metastases are molecularly distinct from their primary tumors. Oncogene 27, 2148–2158 (2008). https://doi.org/10.1038/sj.onc.1210858
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210858