Abstract
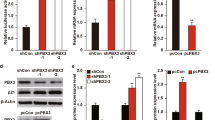
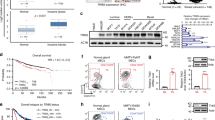
Brn-3b transcription factor enhances proliferation of neuroblastoma (NB) and breast cancer cell lines in vitro and increases the rate and size of in vivo tumour growth, whereas reducing Brn-3b slows growth, both in vitro and in vivo. Brn-3b is elevated in >65% of breast cancer biopsies, and here we demonstrate that Brn-3b is also elevated in NB tumours. We show a significant correlation between Brn-3b and cyclin D1 (CD1) in breast cancers and NB tumours and cell lines. Brn-3b directly transactivates the CD1 promoter in co-transfection experiments, whereas electrophoretic mobility shift assay and chromatin immunoprecipitation assays demonstrate that Brn-3b protein binds to an octamer sequence located in the proximal CD1 promoter. Site-directed mutagenesis of this sequence resulted in loss of transactivation of the CD1 promoter by Brn-3b. Thus, Brn-3b may act to alter growth properties of breast cancer and NB cells by enhancing CD1 expression in these cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
09 February 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41388-023-02614-9
Abbreviations
- CD1:
-
cyclin D1
- ChIP:
-
chromatin immunoprecipitation
- EMSA:
-
electrophoretic mobility shift assay
- LTR:
-
long terminal repeat
- NB:
-
neuroblastoma
- POU:
-
Pit-Oct-Unc
- RA:
-
retinoic acid
References
Arnold A, Motokura T, Bloom T, Kronenberg H, Ruderman J, Juppner H et al. (1991). The putative oncogene PRAD1 encodes a novel cyclin. Cold Spring Harb Symp Quant Biol 56: 93–97.
Barnes DM, Gillett CE . (1998). Cyclin D1 in breast cancer. Breast Cancer Res Treat 52: 1–15.
Bienvenu F, Gascan H, Coqueret O . (2001). Cyclin D1 represses STAT3 activation through a Cdk4-independent mechanism. J Biol Chem 276: 16840–16847.
Brodeur GM . (2003). Neuroblastoma: biological insights into a clinical enigma. Nat Rev Cancer 3: 203–216.
Budhram-Mahadeo V, Lillycrop KA, Latchman DS . (1995). The levels of the antagonistic POU family transcription factors Brn-3a and Brn-3b in neuronal cells are regulated in opposite directions by serum growth factors. Neurosci Lett 185: 48–51.
Budhram-Mahadeo V, Morris PJ, Ward T, Weber B, Sassone-Corsi P, Latchman DS . (2001). The closely related POU family transcription factors Brn-3a and Brn-3b are expressed in distinct cell types in the testis. Int J Biochem Cell Biol 33: 1027–1039.
Budhram-Mahadeo V, Ndisang D, Ward T, Weber BL, Latchman DS . (1999). The Brn-3b POU family transcription factor represses expression of the BRCA-1 anti-oncogene in breast cancer cells. Oncogene 18: 6684–6691.
Budhram-Mahadeo V, Theil T, Morris PJ, Lillycrop KA, Moroy T, Latchman DS . (1994). The DNA target site for the Brn-3 POU family transcription factors can confer responsiveness to cyclic AMP and removal of serum in neuronal cells. Nucleuc Acid Res 22: 3092–3098.
Budhram-Mahadeo VS, Latchman DS . (2006). Targeting Brn-3b in breast cancer therapy. Expert Opin Ther Targets 10: 15–25.
Dennis JH, Budhram-Mahadeo V, Latchman DS . (2001). The Brn-3b POU family transcription factor regulates the cellular growth, proliferation, and anchorage dependence of MCF7 human breast cancer cells. Oncogene 20: 4961–4971.
Erkman L, McEvilly RJ, Luo L, Ryan AK, Hooshmand F, O'Connell SM et al. (1996). Role of transcription factors Brn-3.1 and Brn-3.2 in auditory and visual system development. Nature 381: 603–606.
Fantl V, Stamp G, Andrews A, Rosewell I, Dickson C . (1995). Mice lacking cyclin D1 are small and show defects in eye and mammary gland development. Genes Dev 9: 2364–2372.
Gan L, Xiang M, Zhou L, Wagner DS, Klein WH, Nathans J . (1996). POU domain factor Brn-3b is required for the development of a large set of retinal ganglion cells. Proc Natl Acad Sci USA 93: 3920–3925.
Guo Y, Yang K, Harwalkar J, Nye JM, Mason DR, Garrett MD et al. (2005). Phosphorylation of cyclin D1 at Thr 286 during S phase leads to its proteasomal degradation and allows efficient DNA synthesis. Oncogene 24: 2599–2612.
Irshad S, Pedley RB, Anderson J, Latchman DS, Budhram-Mahadeo V . (2004). The Brn-3b transcription factor regulates the growth, behavior, and invasiveness of human neuroblastoma cells in vitro and in vivo. J Biol Chem 279: 21617–21627.
Israels ED, Israels LG . (2000). The cell cycle. Oncologist 5: 510–513.
Knudsen KE, Cavenee WK, Arden KC . (1999). D-type cyclins complex with the androgen receptor and inhibit its transcriptional transactivation ability. Cancer Res 59: 2297–2301.
Latchman DS . (1998a). Regulation of neuronal differentiation and apoptosis by Brn-3 POU family transcription factors: potential use in gene therapy. Gene Ther Mol Biol 2: 95–101.
Latchman DS . (1998b). The Brn-3a transcription factor. Int J Biochem Cell Biol 30: 1153–1157.
Lee SA, Ndisang D, Patel C, Dennis JH, Faulkes DJ, D'Arrigo C et al. (2005). Expression of the Brn-3b transcription factor correlates with expression of HSP-27 in breast cancer biopsies and is required for maximal activation of the HSP-27 promoter. Cancer Res 65: 3072–3080.
Lillycrop KA, Budhram-Mahadeo V, Lakin ND, Terrenghi G, Wood JN, Polak JM et al. (1992). A novel POU family transcription factor is closely related to Brn-3 but has a distinct expression pattern in neuronal cells. Nucleic Acids Res 20: 5093–5096.
Molenaar JJ, van Sluis P, Boon K, Versteeg R, Caron HN . (2003). Rearrangements and increased expression of cyclin D1 (CCND1) in neuroblastoma. Genes Chromosomes Cancer 36: 242–249.
Motokura T, Bloom T, Kim HG, Juppner H, Ruderman JV, Kronenberg HM et al. (1991). A novel cyclin encoded by a bcl1-linked candidate oncogene. Nature 350: 512–515.
Mu X, Zhao S, Pershad R, Hsieh TF, Scarpa A, Wang SW et al. (2001). Gene expression in the developing mouse retina by EST sequencing and microarray analysis. Nucleic Acids Res 29: 4983–4993.
Redeuilh G, Attia A, Mester J, Sabbah M . (2002). Transcriptional activation by the oestrogen receptor alpha is modulated through inhibition of cyclin-dependent kinases. Oncogene 21: 5773–5782.
Sabbah M, Courilleau D, Mester J, Redeuilh G . (1999). Estrogen induction of the cyclin D1 promoter: involvement of a cAMP response-like element. Proc Natl Acad Sci USA 96: 11217–11222.
Samady L, Dennis J, Budhram-Mahadeo V, Latchman DS . (2004). Activation of CDK4 gene expression in human breast cancer cells by the Brn-3b POU family transcription factor. Cancer Biol Ther 3: 317–323.
Sicinski P, Donaher JL, Parker SB, Li T, Fazeli A, Gardner H et al. (1995). Cyclin D1 provides a link between development and oncogenesis in the retina and breast. Cell 82: 621–630.
Smith MD, Latchman DS . (1996). The functionally antagonistic POU family transcription factors Brn-3a and Brn-3b show opposite changes in expression during the growth arrest and differentiation of human neuroblastoma cells. Int J Cancer 67: 653–660.
Turner EE, Jenne KJ, Rosenfeld MG . (1994). Brn-3.2: a Brn-3-related transcription factor with distinctive central nervous system expression and regulation by retinoic acid. Neuron 12: 205–218.
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A et al. (2002). Accurate normalization of real-time quantitative RT–PCR data by geometric averaging of multiple internal control genes. Genome Biol 3 (7): RESEARCH0034; 1–12.
Wang TC, Cardiff RD, Zukerberg L, Lees E, Arnold A, Schmidt EV . (1994). Mammary hyperplasia and carcinoma in MMTV-cyclin D1 transgenic mice. Nature 369: 669–671.
Weinberg RA . (1995). The retinoblastoma protein and cell cycle control. Cell 81: 323–330.
Xiang M . (1998). Requirement for Brn-3b in early differentiation of postmitotic retinal ganglion cell precursors. Dev Biol 197: 155–169.
Xiang M, Zhou L, Nathans J . (1996). Similarities and differences among inner retinal neurons revealed by the expression of reporter transgenes controlled by Brn-3a, Brn-3b, and Brn-3c promotor sequences. Vis Neurosci 13: 955–962.
Xiang M, Zhou L, Peng YW, Eddy RL, Shows TB, Nathans J . (1993). Brn-3b: a POU domain gene expressed in a subset of retinal ganglion cells. Neuron 11: 689–701.
Zwijsen RM, Klompmaker R, Wientjens EB, Kristel PM, van der BB, Michalides RJ . (1996). Cyclin D1 triggers autonomous growth of breast cancer cells by governing cell cycle exit. Mol Cell Biol 16: 2554–2560.
Zwijsen RM, Wientjens E, Klompmaker R, van der SJ, Bernards R, Michalides RJ . (1997). CDK-independent activation of estrogen receptor by cyclin D1. Cell 88: 405–415.
Acknowledgements
We thank Dr Gerard Redeuilh (Paris, France) for cyclin D1 reporter constructs; Dr J Anderson and Dr P Brock (Great Ormond Street Hospital (GOSH), London) for support and discussions; Italian Neuroblastoma Society; Pathology Department (GOSH, London) and UKCCSG, UK for NB RNA or biopsies; Dr D' Arrigo (Guy's Hospital, London) for breast cancer biopsies and Candis Tissue Bank (Liverpool, UK) for RNA from breast cancer. This work was supported by Child Health Research Action Trust (CHRAT); Breast Cancer Campaign (BCC) UK; Association for International Cancer Research (AICR), UK.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Budhram-Mahadeo, V., Irshad, S., Bowen, S. et al. Proliferation-associated Brn-3b transcription factor can activate cyclin D1 expression in neuroblastoma and breast cancer cells. Oncogene 27, 145–154 (2008). https://doi.org/10.1038/sj.onc.1210621
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210621
Keywords
This article is cited by
-
Vascular dysfunction caused by loss of Brn-3b/POU4F2 transcription factor in aortic vascular smooth muscle cells is linked to deregulation of calcium signalling pathways
Cell Death & Disease (2023)
-
Linking metabolic dysfunction with cardiovascular diseases: Brn-3b/POU4F2 transcription factor in cardiometabolic tissues in health and disease
Cell Death & Disease (2021)
-
The POU4F2/Brn-3b transcription factor is required for the hypertrophic response to angiotensin II in the heart
Cell Death & Disease (2019)
-
Essential but partially redundant roles for POU4F1/Brn-3a and POU4F2/Brn-3b transcription factors in the developing heart
Cell Death & Disease (2017)
-
Co-expression of POU4F2/Brn-3b with p53 may be important for controlling expression of pro-apoptotic genes in cardiomyocytes following ischaemic/hypoxic insults
Cell Death & Disease (2014)