Abstract
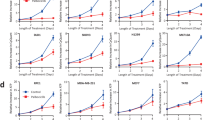
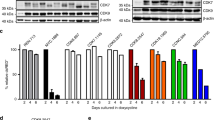
Dasatinib is an ATP-competitive, multi-targeted SRC and ABL kinase inhibitor that can bind BCR-ABL in both the active and inactive conformations. From a clinical standpoint, dasatinib is particularly attractive because it has been shown to induce hematologic and cytogenetic responses in imatinib-resistant chronic myeloid leukemia patients. The fact because the combination of imatinib and dasatinib shows the additive/synergistic growth inhibition on wild-type p210 BCR-ABL-expressing cells, we reasoned that these ABL kinase inhibitors might induce the different molecular pathways. To address this question, we used DNA microarrays to identify genes whose transcription was altered by imatinib and dasatinib. K562 cells were cultured with imatinib or dasatinib for 16 h, and gene expression data were obtained from three independent microarray hybridizations. Almost all of the imatinib- and dasatinib-responsive genes appeared to be similarly increased or decreased in K562 cells; however, small subsets of genes were identified as selectively altered expression by either imatinib or dasatinib. The distinct genes that are selectively modulated by dasatinib are cyclin-dependent kinase 2 (CDK2) and CDK8, which had a maximal reduction of <5-fold in microarray screen. To assess the functional importance of dasatinib regulated genes, we used RNA interference to determine whether reduction of CDK2 and CDK8 affected the growth inhibition. K562 and TF-1BCR-ABL cells, pretreated with CDK2 or CDK8 small interfering RNA, showed additive growth inhibition with imatinib, but not with dasatinib. These findings demonstrate that the additive/synergistic growth inhibition by imatinib and dasatinib may be mediated in part by CDK2 and CDK8.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Burgess MR, Skaggs BJ, Shah NP, Lee FY, Sawyers CL . (2005). Comparative analysis of two clinically active BCR-ABL kinase inhibitors reveals the role of conformation-specific binding in resistance. Proc Natl Acad Sci USA 102: 3395–3400.
Dai Y, Rahmani M, Corey SJ, Dent P, Grant S . (2004). A BCR/ABL-independent, Lyn-dependent form of imatinib mesylate (STI-571) resistance is associated with altered expression of Bcl-2. J Biol Chem 279: 34227–34239.
Donato NJ, Wu JY, Stapley J, Gallick G, Lin H, Arlinghaus R et al. (2003). BCR-ABL independent and LYN kinase overexpression in chronic myelogenous leukemia cells selected for resistance to STI571. Blood 101: 690–698.
Gambacorti-Passerini C, Gasser M, Ahmed S, Assouline S, Scapozza L . (2005). Abl inhibitor BMS354825 binding mode in Abelson kinase revealed by molecular docking studies. Leukemia 19: 1267–1269.
Gesbert F, Sellers WR, Signoretti S, Loda M, Griffin GD . (2000). BCR/ABL regulates expression of the cyclin-dependent kinase inhibitor p27Kip1 through the phosphatidylinositol 3-kinase/AKT pathway. J Biol Chem 275: 39223–39230.
Hofmann WK, de Vos S, Elashoff D, Gschaidmeier H, Hoelzer D, Koeffler HP et al. (2002). Relation between resistance of Philadelphia-chromosome-positive acute lymphoblastic leukemia to the tyrosine kinase inhibitor STI571 and gene-expression profiles: a gene expression study. Lancet 359: 481–486.
Jiang Y, Zhao RC, Verfaillie CM . (2000). Abnormal integrin-mediated regulation of chronic myelogenous leukemia CD34+ cell proliferation: BCR/ABL up-regulates the cyclin-dependent kinase inhibitor, p27Kip, which is relocated to the cell cytoplasm and incapable of regulating cdk2 activity. Proc Natl Acad Sci USA 97: 10538–10543.
Komatsu N, Watanabe T, Uchida M, Mori M, Kirito K, Kikuchi S et al. (2003). A member of Forkhead transcription factor FKHRL1 is a downstream effector of STI571-induced cell cycle arrest in BCR-ABL-expressing cells. J Biol Chem 278: 6411–6419.
Lionberger JM, Wilson MB, Smithgall TE . (2000). Transformation of myeloid leukemia cells to cytokine independence by BCR-ABL is suppressed by kinase-defective Hck. J Biol Chem 275: 18581–18585.
Nakajima A, Tauchi T, Sumi M, Bishop WR, Ohyashiki K . (2003). Efficacy of SCH66336, a farnesyl transferase inhibitor, in conjunction with imatinib against BCR-ABL-positive cells. Mol Cancer Ther 2: 219–224.
O'Hare T, Walters DK, Stoffregen EP, Jia T, Manley PW, Mestan J et al. (2005). In vitro activity of BCR-ABL inhibitors AMN107 and BMS-354825 against clinically relevant imatinib-resistant Abl kinase domain mutants. Cancer Res 65: 4500–4505.
Ohyashiki JH, Takaku T, Ojima T, Abe K, Yamamoto K, Zhang Y et al. (2005). Transcriptional profiling of human herpesvirus type B (HHV-6B) in an adult T cell leukemia cell line as in vitro model for persistent infection. Biochem Biophys Res Commun 329: 11–17.
Parada Y, Banerji L, Glassford J, Lea NC, Collada M, Rivas C et al. (2001). BCR-ABL and interleukin 3 promote haematopoietic cell proliferation and survival through modulation of cyclin D2 and p27Kip1 expression. J Biol Chem 276: 23572–23580.
Prathapam T, Tegen S, Oskarsson T, Trumpp A, Martin GS . (2006). Activated Src abrogates the Myc requirement for the G0/G1 transition but not for the G1/S transition. Proc Natl Acad Sci USA 103: 2695–2700.
Ptasznik A, Nakata Y, Kalota A, Emerson SG, Gewirtz AM . (2004). Short interfering RNA (siRNA) targeting the Lyn kinase induces apoptosis in primary, and drug-resistant, BCR-ABL1(+) leukemia cells. Nat. Med 10: 1187–1189.
Riley D, Carragher NO, Frame MC, Wyke JA . (2001). The mechanism of cell cycle regulation by v-Src. Oncogene 20: 5941–5950.
Samanta AK, Lin H, Sun T, Kantarjian H, Arlinghaus RB . (2006). Janus kinase 2: a critical target in chronic myelogenous leukemia. Cancer Res 66: 6468–6472.
Shah NP, Tran C, Lee FY, Chen P, Norris D, Sawyers CL . (2004). Overriding imatinib resistance with a novel ABL kinase inhibitor. Science 305: 399–401.
Sherr CJ, Roberts JM . (1999). CDK inhibitors: positive and negative regulators of G1-phase progression. Genes Dev 13: 1501–1512.
Stanglmaier M, Warmuth M, Kleinlein I, Hallek M . (2003). The interaction of the BCR-ABL tyrosine kinase Hck is mediated by multiple binding domains. Leukemia 17: 286–289.
Takaku T, Ohyashiki JH, Zhang Y, Ohyashiki K . (2005). Estimating immunoregulatory gene networks in human herpesvirus type 6-infected T cells. Biochem Biophys Res Commu 336: 469–477.
Talpaz M, Shah NP, Kantarjian H, Donato N, Nicoll J, Paquette R et al. (2006). Dasatinib in imatinib-resistant Philadelphia chromosome-positive leukemias. N Engl J Med 354: 2531–2541.
Tauchi T, Feng GS, Shen R, Song HY, Donner D, Pawson T et al. (1994). SH2-containing phosphotyrosine phosphatase Syp is a target of p210bcr-abl tyrosine kinase. J Biol Chem 269: 15381–15387.
Tetsu O, McCormick F . (2003). Proliferation of cancer cells despite CDK2 inhibition. Cancer Cell 3: 233–245.
Thomas SM, Brugge JS . (1997). Cellular functions regulated by Src family kinases. Annu Rev Cell Dev Biol 13: 513–609.
Warmuth M, Bergmann M, Priess A, Hauslmann K, Emmerich B, Hallek M . (1997). The Src family kinase Hck interacts with BCR-ABL by a kinase-independent mechanism and phosphorylates the Grb2-binding site of BCR. J Biol Chem 272: 33260–33270.
Zhang Y, Ohyashiki JH, Takaku T, Shimizu N, Ohyashiki K . (2006). Transcriptional profiling of Epstein–Barr virus (EBV) genes and host cellular genes in nasal NK/T-cell lymphoma and chronic active EBV infection. Br J Cancer 94: 599–608.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Supplementary information
Rights and permissions
About this article
Cite this article
Nunoda, K., Tauchi, T., Takaku, T. et al. Identification and functional signature of genes regulated by structurally different ABL kinase inhibitors. Oncogene 26, 4179–4188 (2007). https://doi.org/10.1038/sj.onc.1210179
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210179
Keywords
This article is cited by
-
Comparative expression analysis of dasatinib and ponatinib-regulated lncRNAs in chronic myeloid leukemia and their network analysis
Medical Oncology (2022)
-
Deciphering the mechanism of Indirubin and its derivatives in the inhibition of Imatinib resistance using a “drug target prediction-gene microarray analysis-protein network construction” strategy
BMC Complementary and Alternative Medicine (2019)
-
Dasatinib synergises with irinotecan to suppress hepatocellular carcinoma via inhibiting the protein synthesis of PLK1
British Journal of Cancer (2017)
-
Combined effects of novel heat shock protein 90 inhibitor NVP-AUY922 and nilotinib in a random mutagenesis screen
Oncogene (2011)
-
Dasatinib impairs long-term expansion of leukemic progenitors in a subset of acute myeloid leukemia cases
Annals of Hematology (2010)