Abstract
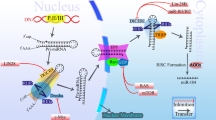
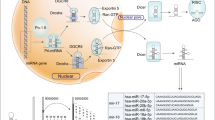
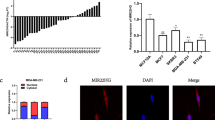
MicroRNAs (miRNAs) are ∼22 nucleotide non-coding RNA molecules that regulate gene expression post-transcriptionally. Although aberrant expression of miRNAs in various human cancers suggests a role for miRNAs in tumorigenesis, it remains largely unclear as to whether knockdown of a specific miRNA affects tumor growth. In this study, we profiled miRNA expression in matched normal breast tissue and breast tumor tissues by TaqMan real-time polymerase chain reaction miRNA array methods. Consistent with previous findings, we found that miR-21 was highly overexpressed in breast tumors compared to the matched normal breast tissues among 157 human miRNAs analysed. To better evaluate the role of miR-21 in tumorigenesis, we transfected breast cancer MCF-7 cells with anti-miR-21 oligonucleotides and found that anti-miR-21 suppressed both cell growth in vitro and tumor growth in the xenograft mouse model. Furthermore, this anti-miR-21-mediated cell growth inhibition was associated with increased apoptosis and decreased cell proliferation, which could be in part owing to downregulation of the antiapoptotic Bcl-2 in anti-miR-21-treated tumor cells. Together, these results suggest that miR-21 functions as an oncogene and modulates tumorigenesis through regulation of genes such as bcl-2 and thus, it may serve as a novel therapeutic target.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bartel DP . (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116: 281–297.
Berezikov E, Guryev V, van de Belt J, Wienholds E, Plasterk RH, Cuppen E . (2005). Phylogenetic shadowing and computational identification of human microRNA genes. Cell 120: 21–24.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E et al. (2002). Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 99: 15524–15529.
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S et al. (2004). Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA 101: 2999–3004.
Chan JA, Krichevsky AM, Kosik KS . (2005). MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 65: 6029–6033.
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT et al. (2005). Real-time quantification of microRNAs by stem-loop RT–PCR. Nucleic Acids Res 33: e179.
Chen CZ, Li L, Lodish HF, Bartel DP . (2004). MicroRNAs modulate hematopoietic lineage differentiation. Science 303: 83–86.
Cheng AM, Byrom MW, Shelton J, Ford LP . (2005). Antisense inhibition of human miRNAs and indications for an involvement of miRNA in cell growth and apoptosis. Nucleic Acids Res 33: 1290–1297.
Croce CM, Calin GA . (2005). miRNAs, cancer, and stem cell division. Cell 122: 6–7.
Eis PS, Tam W, Sun L, Chadburn A, Li Z, Gomez MF et al. (2005). Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc Natl Acad Sci USA 102: 3627–3632.
Fitzgerald K . (2005). RNAi versus small molecules: different mechanisms and specificities can lead to different outcomes. Curr Opin Drug Discov Dev 8: 557–566.
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S et al. (2005). MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65: 7065–7070.
Kim VN . (2005). MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol 6: 376–385.
Krutzfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, Manoharan M et al. (2005). Silencing of microRNAs in vivo with ‘antagomirs’. Nature 438: 685–689.
Lao K, Xu NL, Yeung V, Chen C, Livak KJ, Straus NA . (2006). Multiplexing RT–PCR for the detection of multiple miRNA species in small samples. Biochem Biophys Res Commun 343: 85–89.
Lee RC, Feinbaum RL, Ambros V . (1993). The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75: 843–854.
Metzler M, Wilda M, Busch K, Viehmann S, Borkhardt A . (2004). High expression of precursor microRNA-155/BIC RNA in children with Burkitt lymphoma. Genes Chromosomes Cancer 39: 167–169.
Michael MZ, SM OC, van Holst Pellekaan NG, Young GP, James RJ . (2003). Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 1: 882–891.
Mo YY, Yu Y, Ee PL, Beck WT . (2004). Overexpression of a dominant-negative mutant Ubc9 is associated with increased sensitivity to anticancer drugs. Cancer Res 64: 2793–2798.
Pillai RS . (2005). MicroRNA function: multiple mechanisms for a tiny RNA? RNA 11: 1753–1761.
Tang F, Hajkova P, Barton SC, Lao K, Surani MA . (2006). MicroRNA expression profiling of single whole embryonic stem cells. Nucleic Acids Res 34: e9.
Tanizawa A, Fujimori A, Fujimori Y, Pommier Y . (1994). Comparison of topoisomerase I inhibition, DNA damage, and cytotoxicity of camptothecin derivatives presently in clinical trials. J Natl Cancer Inst 86: 836–842.
Wightman B, Ha I, Ruvkun G . (1993). Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75: 855–862.
Zamore PD, Haley B . (2005). Ribo-gnome: the big world of small RNAs. Science 309: 1519–1524.
Zhang W, Ran S, Sambade M, Huang X, Thorpe PE . (2002). A monoclonal antibody that blocks VEGF binding to VEGFR2 (KDR/Flk-1) inhibits vascular expression of Flk-1 and tumor growth in an orthotopic human breast cancer model. Angiogenesis 5: 35–44.
Acknowledgements
We are grateful to CHTN and SIU tumor bank for providing patient specimens. We thank Rupinder Grewal and Heather Mizeur for cutting frozen tumor samples. This study was supported in part by Grants CA102630 from NCI and BC045418 from DOD.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Rights and permissions
About this article
Cite this article
Si, ML., Zhu, S., Wu, H. et al. miR-21-mediated tumor growth. Oncogene 26, 2799–2803 (2007). https://doi.org/10.1038/sj.onc.1210083
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210083
Keywords
This article is cited by
-
Blood-based microRNA profiling unveils complex molecular dynamics in breast cancer
Journal of Applied Genetics (2024)
-
RETRACTED ARTICLE: Catalytic Au/PEDOT/Pt micromotors for cancer biomarker detection and potential breast cancer treatment
Applied Nanoscience (2023)
-
The diagnostic value of serum miR-21 in patients with ovarian cancer: a systematic review and meta-analysis
Journal of Ovarian Research (2022)
-
Enzyme-free nucleic acid dual-amplification strategy combined with mimic enzyme catalytic precipitation reaction for the photoelectrochemical detection of microRNA-21
Microchimica Acta (2022)
-
Target-Responsive Template Structure Switching-Mediated Exponential Rolling Circle Amplification for the Direct and Sensitive Detection of MicroRNA
BioChip Journal (2022)