Abstract
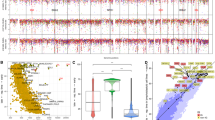
Retroviral vector-mediated overexpression of c-myc in embryonic bursal precursors induces multi-staged tumorigenesis beginning with preneoplastic-transformed follicles (TF) and progressing to clonal metastatic B-cell lymphomas. Using a 13K chicken cDNA microarray, specifically enriched for chicken immune system expressed sequence tagged (ESTs), we carried out array-based comparative genomic hybridization (array-CGH) and detected significant DNA copy number change at many loci on most or all chromosomes in both early TF and end-stage lymphomas. Formation of long palindromes, through breakage–fusion–bridge cycles, is thought to play an early role in gene amplification. Employing genome-wide analysis of palindrome formation (GAPF), we detected extensive palindrome formation in early TF and end-stage lymphomas. The population of loci showing amplification by array-CGH was enriched for palindromes detected by GAPF providing strong evidence for genetic instability early in Myc-induced tumorigenesis and further support for the role of palindromes in gene amplification. Comparing gene copy number change and RNA expression changes profiled on the same cDNA array, we detected very little consistent contribution of gene copy number change to RNA expression changes. Palindromic loci in TF and tumors, however, were expressed, many at high levels, suggesting an abundance of RNA species with long double-stranded segments generated during tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Accession codes
References
Boardman E, Sanz-Ezquerro J, Overton IM, Burt DW, Bosch E, Fong WT et al. (2002). Curr Biol 12: 1965–1969.
Brandvold KA, Neiman P, Ruddell A . (2000). Oncogene 19: 2780–2785.
Brown CY, Bowers SJ, Loring G, Heberden C, Lee R-m, Neiman PE . (2004). Dev Comp Immunol 28: 619–634.
Burnside J, Neiman P, Tang J, Basom R, Talbot R, Aronszajn M et al. (2005). BMC Genom 6: 13.
Butler DK, Yasuda LE, Yao MC . (1996). Cell 87: 1115–1122.
Chang H, Delany ME . (2004). Chromosome Res 12: 299–307.
Ellerman V, Bang O . (1908). ZBL Bakt Parisit Infekt Hygiene 46: 595–609.
Enrietto PJ, Payne LN, Hayman MJ . (1983). Cell 35: 369–379.
Eskola J, Toivanen P . (1974). Cell Immunol 13: 459–471.
Felsher DW, Bishop JM . (1999). Proc Natl Acad Sci USA 96: 3940–3944.
Feo S, Di Liegro C, Mangano R, Read M, Fried M . (1996). Oncogene 13: 1521–1529.
Ford M, Fried M . (1986). Cell 45: 425–430.
Grandori C, Cowley SM, James LP, Eisenman RN . (2000). Annu Rev Cell Dev Biol 16: 653–699.
Hillier LW, Miller W, Birney E, Warren W, Hardison RC, Ponting CP et al. (2004). Nature 432: 695–716.
Hughes S, Greenhouse JJ, Petropoulos CJ, Sutrave P . (1987). J Virol 61: 3004–3012.
Kittler R, Putz G, Pelletier L, Poser I, Heninger AK, Drechsel D et al. (2004). Nature 432: 1036–1040.
Lengauer C, Kinzler KW, Vogelstein B . (1998). Nature 396: 643–649.
Looney JE, Hamlin JL . (1987). Mol Cell Biol 7: 569–577.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al. (2005). Nature 435: 834–838.
Ma C, Martin S, Trask B, Hamlin JL . (1993). Genes Dev 7: 605–620.
Mai S, Fluri M, Siwarski D, Huppi K . (1996a). Chromosome Res 4: 365–371.
Mai S, Hanley-Hyde J, Fluri M . (1996b). Oncogene 12: 277–288.
Mangano R, Piddini E, Carramusa L, Duhig T, Feo S, Fried M . (1998). Oncogene 17: 2771–2777.
Miller AD, Garcia JV, vonSuhr N, Lynch CM, Wilson C, Eiden M . (1991). J Virol 65: 2220–2224.
Neiman P, Burnside J, Elsaesser K, Hwang H, Clurman BE, Kimmel R et al. (2006). In: Buerstedde J-M (ed). Reviews and Protocols in DT40 Research: Subcellular Biochemistry. Springer: Dordrecht, (in press).
Neiman P, Payne LN, Weiss RA . (1980). J Virol 34: 178–186.
Neiman P, Wolf C, Enrietto PJ, Cooper GM . (1985). Proc Natl Acad Sci USA 82: 222–236.
Neiman PE . (1994). Adv Immunol 56: 467–484.
Neiman PE, Grbic JJ, Polony TS, Kimmel R, Bowers SJ, Delrow J et al. (2003). Oncogene 22: 1073–1086.
Neiman PE, Purchase HG, Okazaki W . (1975). Cell 4: 311–319.
Neiman PE, Ruddell A, Jasoni C, Loring G, Thomas SJ, Brandvold KA et al. (2001). Proc Natl Acad Sci USA 98: 6378–6383.
Neiman PE, Thomas SJ, Loring G . (1991). Proc Natl Acad Sci USA 88: 5857–5861.
O'Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT . (2005). Nature 435: 839–843.
Pollack JR, Perou CM, Alizadeh AA, Eisen MB, Pergamenschikov A, Williams CF et al. (1999). Nat Genet 23: 41–46.
Tanaka H, Bergstrom DA, Yao MC, Tapscott SJ . (2005). Nat Genet 37: 320–327.
Tanaka H, Tapscott SJ, Trask BJ, Yao MC . (2002). Proc Natl Acad Sci USA 99: 8772–8777.
Thompson CB, Humphries EH, Carlson LM, Chen C-LH, Neiman PE . (1987). Cell 51: 371–381.
Tlsty TD . (1997). Curr Top Microbiol Immunol 221: 37–46.
Van Gelder RN, von Zastrow ME, Yool A, Dement WC, Barchas JD, Eberwine JH . (1990). Proc Natl Acad Sci USA 87: 1663–1667.
Yang YH, Dudoit S, Luu P, Lin DM, Peng V, Ngai J et al. (2002). Nucleic Acids Res 30: e15.
Yasuda LF, Yao MC . (1991). Cell 67: 505–516.
Zeller KI, Jegga AG, Aronow BJ, O'Donnell KA, Dang CV . (2003). Genome Biol 4: R69.
Acknowledgements
We thank Alana Ruddell, Brian Freie, Mark Groudine and Steve Tapscott for helpful comments. This work was supported by NIH Grants R01 CA-20068 and R01 CA-109365 to PEN.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Rights and permissions
About this article
Cite this article
Neiman, P., Kimmel, R., Icreverzi, A. et al. Genomic instability during Myc-induced lymphomagenesis in the bursa of Fabricius. Oncogene 25, 6325–6335 (2006). https://doi.org/10.1038/sj.onc.1209646
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209646
Keywords
This article is cited by
-
PARP inhibitors enhance replication stress and cause mitotic catastrophe in MYCN-dependent neuroblastoma
Oncogene (2017)
-
The MRN complex is transcriptionally regulated by MYCN during neural cell proliferation to control replication stress
Cell Death & Differentiation (2016)
-
Transformation, genomic instability and senescence mediated by platelet/megakaryocyte glycoprotein Ibα
Oncogene (2008)