Abstract
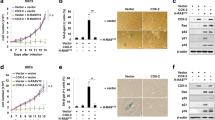
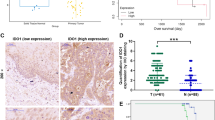
Overexpression of cyclooxygenase-2 (Cox-2) is thought to exert antiapoptotic effects in cancer. Here we show that the tumor suppressor p53 upregulated Cox-2 in esophageal and colon cancer cell lines by inducing the binding of nuclear factor-kappaB (NF-kappaB) to its response element in the COX-2 promoter. Inhibition of NF-kappaB prevented p53 induction of Cox-2 expression. Cooperation between p53 and NF-kappaB was required for activation of COX-2 promoter in response to daunomycin, a DNA-damaging agent. Pharmacological inhibition of Cox-2 enhanced apoptosis in response to daunomycin, in particular in cells containing active p53. In esophageal cancer, there was a correlation between Cox-2 expression and wild-type TP53 in Barrett's esophagus (BE) and in adenocarcinoma, but not in squamous cell carcinoma (P<0.01). These results suggest that p53 and NF-kappaB cooperate in upregulating Cox-2 expression, promoting cell survival in inflammatory precursor lesions such as BE.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Barnas C, Martel-Planche G, Furukawa Y, Hollstein M, Montesano R, Hainaut P . (1997). Int J Cancer 71: 79–87.
Beltrami E, Plescia J, Wilkinson JC, Duckett CS, Altieri DC . (2004). J Biol Chem 279: 2077–2084.
Benoit V, Hellin AC, Huygen S, Gielen J, Bours V, Merville MP . (2000). Oncogene 19: 4787–4794.
Bentires-Alj M, Dejardin E, Viatour P, van Lint C, Froesch B, Reed JC et al. (2001). Oncogene 20: 2805–2813.
Bentires-Alj M, Hellin AC, Ameyar M, Chouaib S, Merville MP, Bours V . (1999a). Cancer Res 59: 811–815.
Bentires-Alj M, Merville MP, Bours V . (1999b). Drug Resist Updat 2: 274–276.
Birkenkamp-Demtroder K, Olesen SH, Sorensen FB, Laurberg S, Laiho P, Aaltonen LA et al. (2005). Gut 54: 374–384.
Bohuslav J, Chen LF, Kwon H, Mu Y, Greene WC . (2004). J Biol Chem 279: 26115–26125.
Buskens CJ, Sivula A, van Rees BP, Haglund C, Offerhaus GJ, van Lanschot JJ et al. (2003). Gut 52: 1678–1683.
Castellone MD, Teramoto H, Williams BO, Durey KM, Gutkind JS . (2005). Science 310: 1504–1509.
Corcoran CA, He Q, Huang Y, Sheikh MS . (2005). Oncogene 24: 1634–1640.
Das KC, White CW . (1997). J Biol Chem 272: 14914–14920.
Devary Y, Rosette C, DiDonato JA, Karin M . (1993). Science 261: 1442–1445.
Eberhart CE, Coffey RJ, Radhika A, Giardiello FM, Ferrenbach S, DuBois RN . (1994). Gastroenterology 107: 1183–1188.
Gilroy DW, Saunders MA, Sansores-Garcia L, Matijevic-Aleksic N, Wu KK . (2001). FASEB J 15: 288–290.
Han JA, Kim JI, Ongusaha PP, Hwang DH, Ballou LR, Mahale A et al. (2002). EMBO J 21: 5635–5644.
Han JH, Roh MS, Park CH, Park KC, Cho KH, Kim KH et al. (2004). Mech Ageing Dev 125: 359–366.
Harris RE, Alshafie GA, Abou-Issa H, Seibert K . (2000). Cancer Res 60: 2101–2103.
Hla T, Bishop-Bailey D, Liu CH, Schaefers HJ, Trifan OC . (1999). Int J Biochem Cell Biol 31: 551–557.
Jeong SJ, Pise-Masison CA, Radonovich MF, Park HU, Brady JN . (2005). J Biol Chem 280: 10326–10332.
Jiang XH, Lam SK, Lin MC, Jiang SH, Kung HF, Slosberg ED et al. (2002). Oncogene 21: 6113–6122.
Jobin C, Morteau O, Han DS, Balfour SR . (1998). Immunology 95: 537–543.
Kaur BS, Khamnehei N, Iravani M, Namburu SS, Lin O, Triadafilopoulos G . (2002). Gastroenterology 123: 60–67.
Kaur BS, Triadafilopoulos G . (2002). Am J Physiol Gastrointest Liver Physiol 283: G327–G334.
Konturek PC, Nikiforuk A, Kania J, Raithel M, Hahn EG, Muhldorfer S . (2004). Dig Dis Sci 49: 1075–1083.
Krysan K, Dalwadi H, Sharma S, Pold M, Dubinett S . (2004a). Cancer Res 64: 6359–6362.
Krysan K, Merchant FH, Zhu L, Dohadwala M, Luo J, Lin Y et al. (2004b). FASEB J 18: 206–208.
Kuo KT, Chow KC, Wu YC, Lin CS, Wang HW, Li WY et al. (2003). Ann Thorac. Surg 76: 909–914.
Lagorce C, Paraf F, Vidaud D, Couvelard A, Wendum D, Martin A et al. (2003). Histopathology 42: 457–465.
Li N, Karin M . (1998). Proc Natl Acad Sci USA 95: 13012–13017.
Meyer M, Schreck R, Baeuerle PA . (1993). EMBO J 12: 2005–2015.
North S, Pluquet O, Maurici D, El Ghissassi F, Hainaut P . (2002). Mol Carcinog 33: 181–188.
Oshima M, Murai N, Kargman S, Arguello M, Luk P, Kwong E et al. (2001). Cancer Res 61: 1733–1740.
Rockwell P, Martinez J, Papa L, Gomes E . (2004). Cell Signal 16: 343–353.
Ryan KM, Ernst MK, Rice NR, Vousden KH . (2000). Nature 404: 892–897.
Sawaoka H, Kawano S, Tsuji S, Tsujii M, Gunawan ES, Takei Y et al. (1998). Am J Physiol 274: G1061–G1067.
Sengupta S, Harris CC . (2005). Nat Rev Mol Cell Biol 6: 44–55.
Sepehr A, Taniere P, Martel-Planche G, Zia'ee AA, Rastgar-Jazii F, Yazdanbod M et al. (2001). Oncogene 20: 7368–7374.
Shamma A, Yamamoto H, Doki Y, Okami J, Kondo M, Fujiwara Y et al. (2000). Clin Cancer Res 6: 1229–1238.
Shirvani VN, Ouatu-Lascar R, Kaur BS, Omary MB, Triadafilopoulos G . (2000). Gastroenterology 118: 487–496.
Siewert JR, Stein HJ . (1998). Br J Surg 85: 1457–1459.
Skehan P, Storeng R, Scudiero D, Monks A, McMahon J, Vistica D et al. (1990). J Natl Cancer Inst 82: 1107–1112.
Smith GV, Feakins R, Farthing MJ, Ballinger A . (2005). Am J Clin Pathol 123: 415–420.
Steinbach G, Lynch PM, Phillips RK, Wallace MH, Hawk E, Gordon GB et al. (2000). N Engl J Med 342: 1946–1952.
Strano S, Fontemaggi G, Costanzo A, Rizzo MG, Monti O, Baccarini A et al. (2002). J Biol Chem 277: 18817–18826.
Subbaramaiah K, Altorki N, Chung WJ, Mestre JR, Sampat A, Dannenberg AJ . (1999). J Biol Chem 274: 10911–10915.
Subbaramaiah K, Dannenberg AJ . (2003). Trends Pharmacol Sci 24: 96–102.
Swamy MV, Herzog CR, Rao CV . (2003). Cancer Res 63: 5239–5242.
Tanabe T, Tohnai N . (2002). Prostaglandins Other Lipid Mediat 68–69: 95–114.
Tsuji S, Tsujii M, Kawano S, Hori M . (2001). J Exp Clin Cancer Res 20: 117–129.
Webster GA, Perkins ND . (1999). Mol Cell Biol 19: 3485–3495.
Wu KK, Liou JY, Cieslik K . (2005). Arterioscler Thromb Vasc Biol 25 (4): 679–685.
Yang D, Welm A, Bishop JM . (2004). Proc Natl Acad Sci USA 101: 15100–15105.
Yu HP, Xu SQ, Liu L, Shi LY, Cai XK, Lu WH et al. (2003). Cancer Lett 198: 193–201.
Zhu Y, Saunders MA, Yeh H, Deng WG, Wu KK . (2002). J Biol Chem 277: 6923–6928.
Zimmermann KC, Sarbia M, Weber AA, Borchard F, Gabbert HE, Schror K . (1999). Cancer Res 59: 198–204.
Acknowledgements
We thank Mrs G Martel-Planche and Mrs N Lyandrat for technical assistance with TP53 mutation analysis and immunohistochemistry, respectively. We are also grateful to Dr X de Leval (Laboratory of Pharmaceutical Chemistry, University of Liège, Belgium) for providing Cox-2 inhibitors. E de Moraes was supported by a PhD fellowship from CAPES-COFECUB. V Benoit, M-P Merville and A Chariot, are respectively, senior research assistant and research associates at the National Fund for Scientific Research (Belgium). NA Dar is supported by a Special Training Award from IARC. This research was supported by ‘subvention libre’ from ARC (French Association Against Cancer) and by grants from Leon Fredericq Foundation, ‘Centre Anticancéreux de l'Université de Liège, Télévie, Belgian Federation against cancer and National Fund for Scientific Research (Belgium).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Supplementary information
Rights and permissions
About this article
Cite this article
Benoit, V., de Moraes, E., Dar, N. et al. Transcriptional activation of cyclooxygenase-2 by tumor suppressor p53 requires nuclear factor-kappaB. Oncogene 25, 5708–5718 (2006). https://doi.org/10.1038/sj.onc.1209579
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209579
Keywords
This article is cited by
-
Inflammation and insulin resistance exert dual effects on adipose tissue tumor protein 53 expression
International Journal of Obesity (2014)
-
Non-steroidal anti-inflammatory drugs (NSAIDs) and breast cancer risk: differences by molecular subtype
Cancer Causes & Control (2011)
-
Regulation of MCP-1 chemokine transcription by p53
Molecular Cancer (2010)
-
A crucial role for adipose tissue p53 in the regulation of insulin resistance
Nature Medicine (2009)
-
The dark side of a tumor suppressor: anti-apoptotic p53
Cell Death & Differentiation (2008)