Abstract
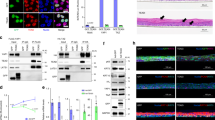
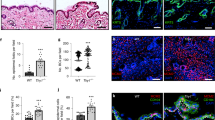
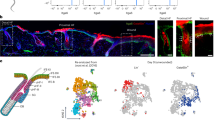
Activin is a member of the transforming growth factor β (TGF-β) family, which plays a crucial role in skin morphogenesis and wound healing. To gain insight into the underlying mechanisms of action, we searched for activin-regulated genes in cultured keratinocytes. One of the identified target genes encodes Id1, a negative regulator of helix–loop–helix transcription factors. We show that Id1, Id2, and Id3 are strongly downregulated by activin in keratinocytes in vitro and in vivo. To determine the role of Id1 in keratinocyte biology, we generated stable HaCaT keratinocyte cell lines overexpressing this protein. Our results revealed that enhanced levels of Id1 do not affect proliferation of keratinocytes in monoculture under exponential culture conditions or in response to activin or TGF-β1. However, in three-dimensional organotypic cultures, Id1-overexpressing HaCaT cells formed a hyperthickened and disorganized epithelium that was characterized by enhanced keratinocyte proliferation, abnormal differentiation, and an increased rate of apoptosis. These results identify an important function of Id1 in the regulation of epidermal homeostasis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bamberger C, Schärer A, Antsiferova M, Tychsen B, Pankow S, Müller M et al. (2005). Am J Pathol 167: 733–747.
Benezra R, Davis RL, Lockshon D, Turner DL, Weintraub H . (1990). Cell 61: 49–59.
Bjorntorp E, Parsa R, Thornemo M, Wennberg AM, Lindahl A . (2003). Acta Derm Venerol 83: 403–409.
Boukamp P, Petrussevska RT, Breitkreutz D, Hornung J, Markham A, Fusenig NE . (1988). J Cell Biol 106: 761–771.
Chambers RC, Leoni P, Kaminski N, Laurent GJ, Heller RA . (2003). Am J Pathol 162: 533–546.
Chaturvedi V, Bonish B, Bacon P, Qin JZ, Denning MF, Foreman K et al. (2003). Exp Dermatol 12: 255–260.
Chomczynski P, Sacchi N . (1987). Anal Biochem 162: 156–159.
Cruise BA, Xu P, Hall AK . (2004). Dev Biol 271: 1–10.
Deed RW, Hirose T, Mitchell EL, Santibanez-Koref MF, Norton JD . (1994). Gene 151: 309–314.
Frank S, Madlener M, Werner S . (1996). J Biol Chem 271: 10188–10193.
Hara E, Yamaguchi T, Nojima H, Ide T, Campisi J, Okayama H et al. (1994). J Biol Chem 269: 2139–2145.
Hübner G, Hu Q, Smola H, Werner S . (1996). Dev Biol 173: 490–498.
Kang Y, Chen CR, Massague J . (2003). Mol Cell 11: 915–926.
Korchynskyi O, ten Dijke P . (2002). J Biol Chem 277: 4883–4891.
Kowanetz M, Valcourt U, Bergstrom R, Heldin CH, Moustakas A . (2004). Mol Cell Biol 24: 4241–4254.
Kretschmer A, Moepert K, Dames S, Sternberger M, Kaufmann J, Klippel A . (2003). Oncogene 22: 6748–6763.
Kurisaki K, Kurisaki A, Valcourt U, Terentiev AA, Pardali K, Ten Dijke P et al. (2003). Mol Cell Biol 23: 4494–4510.
Langlands K, Down GA, Kealey T . (2000). Cancer Res 60: 5929–5933.
Lasorella A, Uo T, Iavarone A . (2001). Oncogene 20: 8326–8333.
Lin CQ, Parrinello S, Campisi J, Desprez PY . (1999). Endocr Relat Cancer 6: 49–50.
Ling MT, Wang X, Tsao SW, Wong YC . (2002). Biochim Biophys Acta 1570: 145–152.
Maas-Szabowski N, Starker A, Fusenig NE . (2003). J Cell Sci 116: 2937–2948.
Martin P . (1997). Science 276: 75–81.
Massague J . (1990). Annu Rev Cell Biol 6: 597–641.
Mathews LS, Vale WW . (1993). Receptor 3: 173–181.
Munz B, Smola H, Engelhardt F, Bleuel K, Brauchle M, Lein I et al. (1999). EMBO J 18: 5205–5215.
Nakamura T, Takio K, Eto Y, Shibai H, Titani K, Sugino H . (1990). Science 247: 836–838.
Norton JD, Deed RW, Craggs G, Sablitzky F . (1998). Trends Cell Biol 8: 58–65.
Ohga E, Matsuse T, Teramoto S, Katayama H, Nagase T, Fukuchi Y et al. (1996). Biochem Biophys Res Commun 228: 391–396.
Ruzinova MB, Benezra R . (2003). Trends Cell Biol 13: 410–418.
Ryle CM, Breitkreutz D, Stark HJ, Leigh IM, Steinert PM, Roop D et al. (1989). Differentiation 40: 42–54.
Schaefer BM, Koch J, Wirzbach A, Kramer MD . (2001). Exp Cell Res 266: 250–259.
Schoop VM, Mirancea N, Fusenig NE . (1999). J Invest Dermatol 112: 343–353.
Seishima M, Nojiri M, Esaki C, Yoneda K, Eto Y, Kitajima Y . (1999). J Invest Dermatol 112: 432–436.
Shimizu A, Kato M, Nakao A, Imamura T, ten Dijke P, Heldin CH et al. (1998). Genes Cells 3: 125–134.
Shipley GD, Pittelkow MR, Wille Jr JJ, Scott RE, Moses HL . (1986). Cancer Res 46: 2068–2071.
Siegel PM, Shu W, Massague J . (2003). J Biol Chem 278: 35444–35450.
Sikder HA, Devlin MK, Dunlap S, Ryu B, Alani RM . (2003). Cancer Cell 3: 525–530.
Singer AJ, Clark RA . (1999). N Engl J Med 341: 738–746.
Smola H, Stark HJ, Thiekotter G, Mirancea N, Krieg T, Fusenig NE . (1998). Exp Cell Res 239: 399–410.
Stark HJ, Baur M, Breitkreutz D, Mirancea N, Fusenig NE . (1999). J Invest Dermatol 112: 681–691.
Tsuchida K, Nakatani M, Yamakawa N, Hashimoto O, Hasegawa Y, Sugino H . (2004). Mol Cell Endocrinol 220: 59–65.
Wankell M, Kaesler S, Zhang YQ, Florence C, Werner S, Duan R . (2001a). J Endocrinol 171: 385–395.
Wankell M, Munz B, Hubner G, Hans W, Wolf E, Goppelt A et al. (2001b). EMBO J 20: 5361–5372.
Watt FM . (2002). EMBO J 21: 3919–3926.
Werner S, Beer HD, Mauch C, Luscher B, Werner S . (2001). Oncogene 20: 7494–7504.
Werner S, Smola H, Liao X, Longaker MT, Krieg T, Hofschneider PH et al. (1994). Science 266: 819–822.
Werner S, Weinberg W, Liao X, Peters KG, Blessing M, Yuspa SH et al. (1993). EMBO J 12: 2635–2643.
Yamashita S, Maeshima A, Kojima I, Nojima Y . (2004). J Am Soc Nephrol 15: 91–101.
Zhang L, Deng M, Parthasarathy R, Wang L, Mongan M, Molkentin JD et al. (2005). Mol Cell Biol 25: 60–65.
Acknowledgements
We thank Christiane Born-Berclaz and Iris Nord for excellent technical assistance and Dr Takako Sasaki, Max-Planck-Institute of Biochemistry, Martinsried, Germany, for kindly providing the laminin-5 antibody. This work was supported by a grant from the Swiss National Science Foundation (31-61358.00) to SW and a grant from the EC/Swiss Ministry for Education and Research (LSHG-CT-2003-503447, WOUND) to SW and PB.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rotzer, D., Krampert, M., Sulyok, S. et al. Id proteins: Novel targets of activin action, which regulate epidermal homeostasis. Oncogene 25, 2070–2081 (2006). https://doi.org/10.1038/sj.onc.1209230
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209230
Keywords
This article is cited by
-
Milk lactoperoxidase decreases ID1 and ID3 expression in human oral squamous cell carcinoma cell lines
Scientific Reports (2020)
-
The Id-protein family in developmental and cancer-associated pathways
Cell Communication and Signaling (2017)
-
Effects and mechanisms of a microcurrent dressing on skin wound healing: a review
Military Medical Research (2014)
-
Activin A balance regulates epithelial invasiveness and tumorigenesis
Laboratory Investigation (2014)
-
Molecular and cellular mechanisms of bone morphogenetic proteins and activins in the skin: potential benefits for wound healing
Archives of Dermatological Research (2013)