Abstract
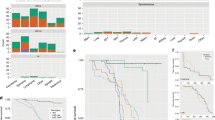
Although radiation can directly induce DNA damage and is a known human and animal carcinogen, the number of genetic changes in radiation-induced tumors, and the pathways responsible for generating them, are unknown. We have used high-density BAC arrays covering >95% of the mouse genome for analysis of genomic patterns of aberrations in spontaneous and radiation-induced mouse lymphomas. The majority of radiation-induced tumors exhibit one of three ‘signatures’ based on gene copy number changes. Some exhibit extensive scrambling of the genome, with very high numbers of recurrent gains and losses. Two other signatures are characterized by excess gains but relatively few losses, or vice versa. Changes in spontaneous tumors often involve whole chromosomes, whereas radiation-induced tumors exhibit a high frequency of localized deletion/amplification events. The number of copy number abnormalities does not correlate with the latency or pathology of the tumors. We propose that specific early events following radiation exposure induce changes in ‘caretaker’ genes that control specific downstream pathways involved in DNA damage repair. The nature of these early events may determine the overall genomic signature observed in the resulting tumor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Albertson DG . (2003). Breast Cancer Res. Treat., 78, 289–298.
Albertson DG, Collins C, McCormick F and Gray JW . (2003). Nat. Genet., 34, 369–376.
Alitalo K, Koskinen P, Makela TP, Saksela K, Sistonen L and Winqvist R . (1987). Biochim. Biophys. Acta, 907, 1–32.
Azzam EI, de Toledo SM and Little JB . (2003). Oncogene, 22, 7050–7057.
Bouffler SD, Kemp CJ, Balmain A and Cox R . (1995). Cancer Res., 55, 3883–3889.
Brenner DJ, Doll R, Goodhead DT, Hall EJ, Land CE, Little JB, Lubin JH, Preston DL, Preston RJ, Puskin JS, Ron E, Sachs RK, Samet JM, Setlow RB and Zaider M . (2003). Proc. Natl. Acad. Sci. USA, 100, 13761–13766.
Cai WW, Mao JH, Chow CW, Damani S, Balmain A and Bradley A . (2002). Nat. Biotechnol., 20, 393–396.
Ferguson DO and Alt FW . (2001). Oncogene, 20, 5572–5579.
Fukasawa K, Choi T, Kuriyama R, Rulong S and Vande Woude GF . (1996). Science, 271, 1744–1747.
Gregory SG, Sekhon M, Schein J, Zhao S, Osoegawa K, Scott CE, Evans RS, Burridge PW, Cox TV, Fox CA, Hutton RD, Mullenger IR, Phillips KJ, Smith J, Stalker J, Threadgold GJ, Birney E, Wylie K, Chinwalla A, Wallis J, Hillier L, Carter J, Gaige T, Jaeger S, Kremitzki C, Layman D, Maas J, McGrane R, Mead K, Walker R, Jones S, Smith M, Asano J, Bosdet I, Chan S, Chittaranjan S, Chiu R, Fjell C, Fuhrmann D, Girn N, Gray C, Guin R, Hsiao L, Krzywinski M, Kutsche R, Lee SS, Mathewson C, McLeavy C, Messervier S, Ness S, Pandoh P, Prabhu AL, Saeedi P, Smailus D, Spence L, Stott J, Taylor S, Terpstra W, Tsai M, Vardy J, Wye N, Yang G, Shatsman S, Ayodeji B, Geer K, Tsegaye G, Shvartsbeyn A, Gebregeorgis E, Krol M, Russell D, Overton L, Malek JA, Holmes M, Heaney M, Shetty J, Feldblyum T, Nierman WC, Catanese JJ, Hubbard T, Waterson RH, Rogers J, de Jong PJ, Fraser CM, Marra M, McPherson JD and Bentley DR . (2002). Nature, 418, 743–750.
Hande MP, Azizova TV, Geard CR, Burak LE, Mitchell CR, Khokhryakov VF, Vasilenko EK and Brenner DJ . (2003). Am. J. Hum. Genet., 72, 1162–1170.
Hodgson G, Hager JH, Volik S, Hariono S, Wernick M, Moore D, Nowak N, Albertson DG, Pinkel D, Collins C, Hanahan D and Gray JW . (2001). Nat. Genet., 29, 459–464.
Ionov Y, Peinado MA, Malkhosyan S, Shibata D and Perucho M . (1993). Nature, 363, 558–561.
Jallepalli PV and Lengauer C . (2001). Nat. Rev. Cancer, 1, 109–117.
Jones PA and Baylin SB . (2002). Nat. Rev. Genet., 3, 415–428.
Kadhim MA, Macdonald DA, Goodhead DT, Lorimore SA, Marsden SJ and Wright EG . (1992). Nature, 355, 738–740.
Kemp CJ, Wheldon T and Balmain A . (1994). Nat. Genet., 8, 66–69.
Kennedy AR, Cairns J and Little JB . (1984). Nature, 307, 85–86.
Kennedy AR and Little JB . (1984). Radiat. Res., 99, 228–248.
Khanna KK and Jackson SP . (2001). Nat. Genet., 27, 247–254.
Kinzler KW and Vogelstein B . (1997). Nature, 386, 761–763.
Liyanage M, Weaver Z, Barlow C, Coleman A, Pankratz DG, Anderson S, Wynshaw-Boris A and Ried T . (2000). Blood, 96, 1940–1946.
Loeb LA, Springgate CF and Battula N . (1974). Cancer Res., 34, 2311–2321.
Mao JH, Wu D, Perez-Losada J, Nagase H, DelRosario R and Balmain A . (2003). Oncogene, 22, 8379–8385.
Mao JH, Perez-Losada J, Wu D, Delrosario R, Tsunematsu R, Nakayama KI, Brown K, Bryson S and Balmain A . (2004). Nature, 432, 775–779.
Marx J . (2002). Science, 297, 544–546.
Mills KD, Ferguson DO and Alt FW . (2003). Immunol. Rev., 194, 77–95.
Mitelman Database of Chromosome Aberrations in Cancer. (2005). Mitelman F, Johansson B and Mertens F (eds), http://cgap.nci.nih.gov/Chromosomes/Mitelman.
Okano H, Saito Y, Miyazawa T, Shinbo T, Chou D, Kosugi S, Takahashi Y, Odani S, Niwa O and Kominami R . (1999). Oncogene, 18, 6677–6683.
Peterson LE . (2002). Comput. Meth. Prog. Biomed., 69, 179–188.
Rajagopalan H, Nowak MA, Vogelstein B and Lengauer C . (2003). Nat. Rev. Cancer, 3, 695–701.
Saito Y, Ochiai Y, Kodama Y, Tamura Y, Togashi T, Kosugi-Okano H, Miyazawa T, Wakabayashi Y, Hatakeyama K, Wakana S, Niwa O and Kominami R . (2001). Oncogene, 20, 5243–5247.
Sieber OM, Heinimann K and Tomlinson IP . (2003). Nat. Rev. Cancer, 3, 701–708.
Sigurdson AJ and Jones IM . (2003). Oncogene, 22, 7018–7027.
Thibodeau SN, Bren G and Schaid D . (1993). Science, 260, 816–819.
Uno M, Wirschubsky Z, Babonits M, Wiener F, Sumegi J and Klein G . (1987). Int. J. Cancer, 40, 540–549.
Volik S, Zhao S, Chin K, Brebner JH, Herndon DR, Tao Q, Kowbel D, Huang G, Lapuk A, Kuo WL, Magrane G, De Jong P, Gray JW and Collins C . (2003). Proc. Natl. Acad. Sci. USA, 100, 7696–7701.
Watson GE, Lorimore SA, Clutton SM, Kadhim MA and Wright EG . (1997). Int. J. Radiat. Biol., 71, 497–503.
Acknowledgements
These studies were initially supported by the Commission of the European Communities and the Cancer Research Campaign (UK), and subsequently by NCI Grant U01 CA84244 and a Grant DE-FG02-03ER63630 from the DOE to AB. Special thanks to the CRC Beatson Institute animal house staff for help with animal husbandry. Dr Jian-Hua Mao is the recipient of a Leukemia & Lymphoma Society Fellowship. Dr Jesus Perez-Losada has a Fellowship from the ‘Ministerio de Educacion y Ciencia of Spain’.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Mao, JH., Li, J., Jiang, T. et al. Genomic instability in radiation-induced mouse lymphoma from p53 heterozygous mice. Oncogene 24, 7924–7934 (2005). https://doi.org/10.1038/sj.onc.1208926
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1208926
Keywords
This article is cited by
-
FATS is an E2-independent ubiquitin ligase that stabilizes p53 and promotes its activation in response to DNA damage
Oncogene (2014)
-
New Biological Insights on the Link Between Radiation Exposure and Breast Cancer Risk
Journal of Mammary Gland Biology and Neoplasia (2013)
-
An HDAC1-binding domain within FATS bridges p21 turnover to radiation-induced tumorigenesis
Oncogene (2010)